Differentiation During Th1 and Th2 Il18r1 Remodeling of Transcription
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Role of IL-4 Receptor &Alpha
Role of IL-4 receptor a–positive CD41 T cells in chronic airway hyperresponsiveness Frank Kirstein, PhD, Natalie E. Nieuwenhuizen, PhD,* Jaisubash Jayakumar, PhD, William G. C. Horsnell, PhD, and Frank Brombacher, PhD Cape Town, South Africa, and Berlin, Germany Background: TH2 cells and their cytokines are associated with IL-17–producing T cells and, consequently, increased airway allergic asthma in human subjects and with mouse models of neutrophilia. allergic airway disease. IL-4 signaling through the IL-4 receptor Conclusion: IL-4–responsive T helper cells are dispensable for 1 a (IL-4Ra) chain on CD4 T cells leads to TH2 cell acute OVA-induced airway disease but crucial in maintaining differentiation in vitro, implying that IL-4Ra–responsive CD41 chronic asthmatic pathology. (J Allergy Clin Immunol T cells are critical for the induction of allergic asthma. However, 2016;137:1852-62.) mechanisms regulating acute and chronic allergen-specific T 2 H Key words: responses in vivo remain incompletely understood. TH2 cell, acute allergic airway disease, chronic asthma, Objective: This study defines the requirements for IL-4Ra– cytokine receptors, IL-4, IL-13, gene-deficient mice responsive CD41 T cells and the IL-4Ra ligands IL-4 and IL-13 in the development of allergen-specific TH2 responses during the Allergic asthma is a chronic inflammatory disease of the airways onset and chronic phase of experimental allergic airway disease. characterized by an inappropriate immune response to harmless Methods: Development of acute and chronic ovalbumin environmental antigens. T 2 cells regulate adaptive immune 1 H (OVA)–induced allergic asthma was assessed weekly in CD4 T responses to allergens, and their presence correlates with disease 2 cell–specific IL-4Ra–deficient BALB/c mice (LckcreIL-4Ra /lox) symptoms in human subjects and mice.1 IL-4 plays a crucial role and respective control mice in the presence or absence of IL-4 in the in vitro and in vivo differentiation of TH2 cells, suggesting or IL-13. -

Genetics of Interleukin 1 Receptor-Like 1 in Immune and Inflammatory Diseases
Current Genomics, 2010, 11, 591-606 591 Genetics of Interleukin 1 Receptor-Like 1 in Immune and Inflammatory Diseases Loubna Akhabir and Andrew Sandford* Department of Medicine, University of British Columbia, UBC James Hogg Research Centre, Providence Heart + Lung Institute, Room 166, St. Paul's Hospital, 1081 Burrard Street, Vancouver, BC V6Z 1Y6, Canada Abstract: Interleukin 1 receptor-like 1 (IL1RL1) is gaining in recognition due to its involvement in immune/inflamma- tory disorders. Well-designed animal studies have shown its critical role in experimental allergic inflammation and human in vitro studies have consistently demonstrated its up-regulation in several conditions such as asthma and rheumatoid ar- thritis. The ligand for IL1RL1 is IL33 which emerged as playing an important role in initiating eosinophilic inflammation and activating other immune cells resulting in an allergic phenotype. An IL1RL1 single nucleotide polymorphism (SNP) was among the most significant results of a genome-wide scan inves- tigating eosinophil counts; in the same study, this SNP associated with asthma in 10 populations. The IL1RL1 gene resides in a region of high linkage disequilibrium containing interleukin 1 receptor genes as well as in- terleukin 18 receptor and accessory genes. This poses a challenge to researchers interested in deciphering genetic associa- tion signals in the region as all of the genes represent interesting candidates for asthma and allergic disease. The IL1RL1 gene and its resulting soluble and receptor proteins have emerged as key regulators of the inflammatory proc- ess implicated in a large variety of human pathologies We review the function and expression of the IL1RL1 gene. -
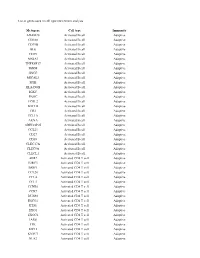
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

B Cell Checkpoints in Autoimmune Rheumatic Diseases
REVIEWS B cell checkpoints in autoimmune rheumatic diseases Samuel J. S. Rubin1,2,3, Michelle S. Bloom1,2,3 and William H. Robinson1,2,3* Abstract | B cells have important functions in the pathogenesis of autoimmune diseases, including autoimmune rheumatic diseases. In addition to producing autoantibodies, B cells contribute to autoimmunity by serving as professional antigen- presenting cells (APCs), producing cytokines, and through additional mechanisms. B cell activation and effector functions are regulated by immune checkpoints, including both activating and inhibitory checkpoint receptors that contribute to the regulation of B cell tolerance, activation, antigen presentation, T cell help, class switching, antibody production and cytokine production. The various activating checkpoint receptors include B cell activating receptors that engage with cognate receptors on T cells or other cells, as well as Toll-like receptors that can provide dual stimulation to B cells via co- engagement with the B cell receptor. Furthermore, various inhibitory checkpoint receptors, including B cell inhibitory receptors, have important functions in regulating B cell development, activation and effector functions. Therapeutically targeting B cell checkpoints represents a promising strategy for the treatment of a variety of autoimmune rheumatic diseases. Antibody- dependent B cells are multifunctional lymphocytes that contribute that serve as precursors to and thereby give rise to acti- cell- mediated cytotoxicity to the pathogenesis of autoimmune diseases -

Cd4/Cd8 Panel, Blood
Lab Dept: Flow and Immunology Test Name: CD4/CD8 PANEL, BLOOD General Information Lab Order Codes: C48P Synonyms: Helper/Suppressor Ratio; T- cells; T-cell subsets; T-cell phenotyping See also: Immune Status Panel and Comprehensive Immune Status Panel CPT Codes: 86359 – T cells, total count 86360 – T cells; absolute CD4 and CD8 count, including ratio Test Includes: CD4(CD3+) and CD8(CD3+) relative percentages, absolute values and a calculated Helper/Suppressor ratio. Logistics Test Indications: This is a minimal antibody panel to monitor immune status. Lab Testing Sections: Flow Cytometry Phone Numbers: MIN Lab: 612-813-6280 STP Lab: 651-220-6550 Test Availability: 3 times weekly determined by volume. Transport collected specimen immediately to Flow Cytometry. Routine testing is not available on weekends or holidays. Therefore, specimens cannot be used if drawn the day before a 3 day weekend such as Memorial Day, Labor Day or major holiday that falls on a Monday or Friday. Turnaround Time: 1 – 3 days Special Instructions: See Test Availability Specimen Specimen Type: Whole blood Container: Lavender top (EDTA) tube Draw Volume: 2 mL blood in a 2 mL Lavender (EDTA) tube Minimum volume: 0.5 mL in an EDTA microtainer Collection: Routine venipuncture Special Processing: Keep sample at room temperature and forward promptly to the laboratory. Do not centrifuge, refrigerate, or freeze sample. Patient Preparation: None Sample Rejection: Specimens will not be processed that are clotted; hemolyzed; greater than 72 hours old; collected in the wrong tube type (0.5 mL in a 2 mL tube), or that have been held or handled at a temperature other than room temperature Interpretive Reference Range: Age-dependant reference ranges provided with results. -

How Relevant Are Bone Marrow-Derived Mast Cells (Bmmcs) As Models for Tissue Mast Cells? a Comparative Transcriptome Analysis of Bmmcs and Peritoneal Mast Cells
cells Article How Relevant Are Bone Marrow-Derived Mast Cells (BMMCs) as Models for Tissue Mast Cells? A Comparative Transcriptome Analysis of BMMCs and Peritoneal Mast Cells 1, 2, 1 1 2,3 Srinivas Akula y , Aida Paivandy y, Zhirong Fu , Michael Thorpe , Gunnar Pejler and Lars Hellman 1,* 1 Department of Cell and Molecular Biology, Uppsala University, The Biomedical Center, Box 596, SE-751 24 Uppsala, Sweden; [email protected] (S.A.); [email protected] (Z.F.); [email protected] (M.T.) 2 Department of Medical Biochemistry and Microbiology, Uppsala University, The Biomedical Center, Box 589, SE-751 23 Uppsala, Sweden; [email protected] (A.P.); [email protected] (G.P.) 3 Department of Anatomy, Physiology and Biochemistry, Swedish University of Agricultural Sciences, Box 7011, SE-75007 Uppsala, Sweden * Correspondence: [email protected]; Tel.: +46-(0)18-471-4532; Fax: +46-(0)18-471-4862 These authors contributed equally to this work. y Received: 29 July 2020; Accepted: 16 September 2020; Published: 17 September 2020 Abstract: Bone marrow-derived mast cells (BMMCs) are often used as a model system for studies of the role of MCs in health and disease. These cells are relatively easy to obtain from total bone marrow cells by culturing under the influence of IL-3 or stem cell factor (SCF). After 3 to 4 weeks in culture, a nearly homogenous cell population of toluidine blue-positive cells are often obtained. However, the question is how relevant equivalents these cells are to normal tissue MCs. By comparing the total transcriptome of purified peritoneal MCs with BMMCs, here we obtained a comparative view of these cells. -

Rational Design of a CD4 Mimic That Inhibits HIV-1 Entry and Exposes Cryptic Neutralization Epitopes
RESEARCH ARTICLE Rational design of a CD4 mimic that inhibits HIV-1 entry and exposes cryptic neutralization epitopes Loïc Martin1, François Stricher1, Dorothée Missé2, Francesca Sironi3, Martine Pugnière4, Philippe Barthe5, Rafael Prado-Gotor5, Isabelle Freulon1, Xavier Magne2, Christian Roumestand5, André Ménez1, Paolo Lusso3, Francisco Veas2, and Claudio Vita1* Published online 16 December 2002; doi:10.1038/nbt768 The conserved surfaces of the human immunodeficiency virus (HIV)-1 envelope involved in receptor binding represent potential targets for the development of entry inhibitors and neutralizing antibodies. Using structural information on a CD4-gp120-17b antibody complex, we have designed a 27-amino acid CD4 mimic, CD4M33, that presents optimal interactions with gp120 and binds to viral particles and diverse HIV-1 envelopes with CD4-like affinity.This mini-CD4 inhibits infection of both immortalized and primary cells by HIV-1, including pri- http://www.nature.com/naturebiotechnology mary patient isolates that are generally resistant to inhibition by soluble CD4. Furthermore, CD4M33 pos- sesses functional properties of CD4, including the ability to unmask conserved neutralization epitopes of gp120 that are cryptic on the unbound glycoprotein. CD4M33 is a prototype of inhibitors of HIV-1 entry and, in complex with envelope proteins, a potential component of vaccine formulations, or a molecular target in phage display technology to develop broad-spectrum neutralizing antibodies. HIV-1 avoids the host immune surveillance by exposing -

The Interleukin-18 Receptor Is Differentially Expressed in Whole
1 The interleukin-18 receptor is differentially expressed in the blood during sepsis. 2 3 Shahan Mamoor, MS1 4 [email protected] East Islip, NY, USA 5 6 We probed published and public microarray datasets1,2 to discover the most significant gene expression changes in the blood of patients with sepsis. We found significant induction 7 expression of IL18RAP and IL18R1, genes encoding subunits of the interleukin-18 receptor, in 8 whole blood from patients with sepsis. 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 Keywords: sepsis, septic shock, interleukin-18, IL18R1, IL18RAP, systems biology of septic 26 shock. 27 28 PAGE 1 OF 13 1 Septic shock is a leading cause of mortality in the United States and worldwide3. We 2 used published and public microarray datasets1,2 to identify differentially expressed genes in the 3 4 blood of patients with sepsis. We identified IL18R1 and IL18RAP as among the genes most 5 differentially expressed in blood in the septic state. 6 7 Methods 8 9 We utilized microarray datasets GSE1001591 and GSE264402 for this differential gene 10 11 expression analysis of blood cells during sepsis. GSE100509 was generated with whole blood 12 using Illumina HumanWG-6 v3.0 expression beadchip technology with n=12 whole blood from 13 control subjects and n=33 whole blood from sepsis patients. GSE26440 was generated using 14 15 Affymetrix Human Genome U133 Plus 2.0 Array technology with n=32 control subjects and 16 n=98 sepsis patients. The Benjamini and Hochberg method of p-value adjustment was used for 17 ranking of differential expression but raw p-values were used for assessment of statistical 18 19 significance of global differential expression. -

Conserved Transcriptomic Profile Between Mouse and Human Colitis
ARTICLE https://doi.org/10.1038/s41467-019-10769-x OPEN Conserved transcriptomic profile between mouse and human colitis allows unsupervised patient stratification Paulo Czarnewski1, Sara M. Parigi1, Chiara Sorini 1, Oscar E. Diaz 1, Srustidhar Das1, Nicola Gagliani1,2,3 & Eduardo J. Villablanca 1,3 1234567890():,; Clinical manifestations and response to therapies in ulcerative colitis (UC) are hetero- geneous, yet patient classification criteria for tailored therapies are currently lacking. Here, we present an unsupervised molecular classification of UC patients, concordant with response to therapy in independent retrospective cohorts. We show that classical clustering of UC patient tissue transcriptomic data sets does not identify clinically relevant profiles, likely due to associated covariates. To overcome this, we compare cross-sectional human data sets with a newly generated longitudinal transcriptome profile of murine DSS-induced colitis. We show that the majority of colitis risk-associated gene expression peaks during the inflammatory rather than the recovery phase. Moreover, we achieve UC patient clustering into two distinct transcriptomic profiles, differing in neutrophil-related gene activation. Notably, 87% of patients in UC1 cluster are unresponsive to two most widely used biological therapies. These results demonstrate that cross-species comparison enables stratification of patients undistinguishable by other molecular approaches. 1 Immunology and Allergy Unit, Department of Medicine, Solna, Karolinska Institute and University Hospital, 17176 Stockholm, Sweden. 2 Department of Medicine and Department of General, Visceral and Thoracic Surgery, University Medical Center Hamburg-Eppendorf, 20246 Hamburg, Germany. 3These authors jointly supervised this work: Nicola Gagliani, Eduardo J. Villablanca. Correspondence and requests for materials should be addressed to E.J.V. -

Tethering IL2 to Its Receptor Il2rb Enhances Antitumor Activity and Expansion of Natural Killer NK92 Cells Youssef Jounaidi, Joseph F
Published OnlineFirst September 15, 2017; DOI: 10.1158/0008-5472.CAN-17-1007 Cancer Therapeutics, Targets, and Chemical Biology Research Tethering IL2 to Its Receptor IL2Rb Enhances Antitumor Activity and Expansion of Natural Killer NK92 Cells Youssef Jounaidi, Joseph F. Cotten, Keith W. Miller, and Stuart A. Forman Abstract IL2 is an immunostimulatory cytokine for key immune cells of IL2 and its receptor IL2Rb joined via a peptide linker (CIRB). including T cells and natural killer (NK) cells. Systemic IL2 NK92 cells expressing CIRB (NK92CIRB) were highly activated and supplementation could enhance NK-mediated immunity in a expanded indefinitely without exogenous IL2. When compared variety of diseases ranging from neoplasms to viral infection. with an IL2-secreting NK92 cell line, NK92CIRB were more acti- However, its systemic use is restricted by its serious side effects and vated, cytotoxic, and resistant to growth inhibition. Direct contact limited efficacy due to activation of T regulatory cells (Tregs). IL2 with cancer cells enhanced the cytotoxic character of NK92CIRB signaling is mediated through interactions with a multi-subunit cells, which displayed superior in vivo antitumor effects in mice. receptor complex containing IL2Ra, IL2Rb, and IL2Rg. Adult Overall, our results showed how tethering IL2 to its receptor natural killer (NK) cells express only IL2Rb and IL2Rg subunits IL2Rb eliminates the need for IL2Ra and IL2Rb, offering a and are therefore relatively insensitive to IL2. To overcome these new tool to selectively activate and empower immune therapy. limitations, we created a novel chimeric IL2-IL2Rb fusion protein Cancer Res; 77(21); 5938–51. Ó2017 AACR. Introduction in order for allogeneic NK cells to be effective, pretransfer lymphodepletion is required to reduce competition for growth Natural killer (NK) cells are lymphocytes endowed with the factors and cytokines (14, 15). -

Estrogen Receptor Alpha (ESR1)-Dependent Regulation of the Mouse Oviductal Transcriptome Katheryn L
University of Kentucky UKnowledge Animal and Food Sciences Faculty Publications Animal and Food Sciences 1-25-2016 Estrogen Receptor Alpha (ESR1)-Dependent Regulation of the Mouse Oviductal Transcriptome Katheryn L. Cerny University of Kentucky, [email protected] Rosanne A. C. Ribeiro University of Kentucky Myoungkun Jeoung University of Kentucky, [email protected] CheMyong Ko University of Illinois at Urbana-Champaign Phillip J. Bridges University of Kentucky, [email protected] Right click to open a feedback form in a new tab to let us know how this document benefits oy u. Follow this and additional works at: https://uknowledge.uky.edu/animalsci_facpub Part of the Animal Sciences Commons, Cell and Developmental Biology Commons, Food Science Commons, and the Physiology Commons Repository Citation Cerny, Katheryn L.; Ribeiro, Rosanne A. C.; Jeoung, Myoungkun; Ko, CheMyong; and Bridges, Phillip J., "Estrogen Receptor Alpha (ESR1)-Dependent Regulation of the Mouse Oviductal Transcriptome" (2016). Animal and Food Sciences Faculty Publications. 7. https://uknowledge.uky.edu/animalsci_facpub/7 This Article is brought to you for free and open access by the Animal and Food Sciences at UKnowledge. It has been accepted for inclusion in Animal and Food Sciences Faculty Publications by an authorized administrator of UKnowledge. For more information, please contact [email protected]. Estrogen Receptor Alpha (ESR1)-Dependent Regulation of the Mouse Oviductal Transcriptome Notes/Citation Information Published in PLOS ONE, v. 11, no. 1, e0147685, p. 1-17. © 2016 Cerny et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.