1 Research Article 29 July 2015 Werner Syndrome Helicase
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Structure and Function of the Human Recq DNA Helicases
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2005 Structure and function of the human RecQ DNA helicases Garcia, P L Posted at the Zurich Open Repository and Archive, University of Zurich ZORA URL: https://doi.org/10.5167/uzh-34420 Dissertation Published Version Originally published at: Garcia, P L. Structure and function of the human RecQ DNA helicases. 2005, University of Zurich, Faculty of Science. Structure and Function of the Human RecQ DNA Helicases Dissertation zur Erlangung der naturwissenschaftlichen Doktorw¨urde (Dr. sc. nat.) vorgelegt der Mathematisch-naturwissenschaftlichen Fakultat¨ der Universitat¨ Z ¨urich von Patrick L. Garcia aus Unterseen BE Promotionskomitee Prof. Dr. Josef Jiricny (Vorsitz) Prof. Dr. Ulrich H ¨ubscher Dr. Pavel Janscak (Leitung der Dissertation) Z ¨urich, 2005 For my parents ii Summary The RecQ DNA helicases are highly conserved from bacteria to man and are required for the maintenance of genomic stability. All unicellular organisms contain a single RecQ helicase, whereas the number of RecQ homologues in higher organisms can vary. Mu- tations in the genes encoding three of the five human members of the RecQ family give rise to autosomal recessive disorders called Bloom syndrome, Werner syndrome and Rothmund-Thomson syndrome. These diseases manifest commonly with genomic in- stability and a high predisposition to cancer. However, the genetic alterations vary as well as the types of tumours in these syndromes. Furthermore, distinct clinical features are observed, like short stature and immunodeficiency in Bloom syndrome patients or premature ageing in Werner Syndrome patients. Also, the biochemical features of the human RecQ-like DNA helicases are diverse, pointing to different roles in the mainte- nance of genomic stability. -

Curriculum Vitae
CURRICULUM VITAE NAME: Patricia Lynn Opresko BUSINESS ADDRESS: University of Pittsburgh Graduate School of Public Health Department of Environmental and UPMC Hillman Cancer Center 5117 Centre Avenue, Suite 2.6a Pittsburgh, PA15213-1863 Phone: 412-623-7764 Fax: 412-623-7761 E-mail: [email protected] EDUCATION AND TRAINING Undergraduate 1990 - 1994 DeSales University B.S., 1994 Chemistry and Center Valley, PA Biology Graduate 1994 - 2000 Pennsylvania State Ph.D., 2000 Biochemistry and University, College of Molecular Biology Medicine, Hershey, PA Post-Graduate 3/2000 - 5/2000 Pennsylvania State Postdoctoral Dr. Kristin Eckert, University, College of Fellow Mutagenesis and Medicine, Jake Gittlen Cancer etiology Cancer Research Institute Hershey, PA 2000-2005 National Institute on IRTA Postdoctoral Dr. Vilhelm Bohr Aging, National Fellow Molecular Institutes of Health, Gerontology and Baltimore, MD DNA Repair 1 APPOINTMENTS AND POSITIONS Academic 8/1/2018 – Co-leader Genome Stability Program, UPMC present Hillman Cancer Center 5/1/2018- Tenured Professor Pharmacology and Chemical Biology, present School of Medicine, University of Pittsburgh, Pittsburgh, PA 2/1/2018- Tenured Professor Environmental and Occupational Health, present Graduate School of Public Health, University of Pittsburgh, Pittsburgh, PA 2014 – Tenured Associate Environmental and Occupational Health, 1/31/2018 Professor Graduate School of Public Health, University of Pittsburgh, Pittsburgh, PA 2005 - 2014 Assistant Professor Environmental and Occupational Health, Graduate School -
Neurodegeneration in Accelerated Aging
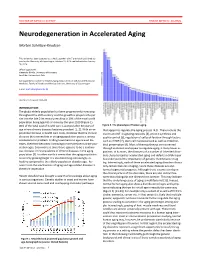
DOCTOR OF MEDICAL SCIENCE DANISH MEDICAL JOURNAL Neurodegeneration in Accelerated Aging Morten Scheibye-Knudsen This review has been accepted as a thesis together with 7 previously published pa- pers by the University of Copenhagen, October 16, 2014 and defended on January 14, 2016 Official opponents: Alexander Bürkle, University of Konstanz Lars Eide, University of Oslo Correspondence: Center for Healthy Aging, Department of Cellular and Molecular Medicine, Faculty of Health and Medical Sciences, University of Copenhagen E-mail: [email protected] Dan Med J 2016;63(11):B5308 INTRODUCTION The global elderly population has been progressively increasing throughout the 20th century and this growth is projected to per- sist into the late 21st century resulting in 20% of the total world population being aged 65 or more by the year 2100 (Figure 1). 80% of the total cost of health care is accrued after 40 years of Figure 2. The phenotype of human aging. age where chronic diseases become prevalent [1, 2]. With an ex- that appear to regulate the aging process [4,5]. These include the ponential increase in health care costs, it follows that the chronic insulin and IGF-1 signaling cascades [4], protein synthesis and diseases that accumulate in an aging population poses a serious quality control [6], regulation of cell proliferation through factors socioeconomic problem. Finding treatments to age related dis- such as mTOR [7], stem cell maintenance 8 as well as mitochon- eases, therefore becomes increasingly more pertinent as the pop- drial preservation [9]. Most of these pathways are conserved ulation ages. Even more so since there appears to be a continu- through evolution and appear to regulate aging in many lower or- ous increase in the prevalence of chronic diseases in the aging ganisms. -

Werner Syndrome Protein Participates in a Complex with RAD51, RAD54
Erratum Werner syndrome protein participates in a complex with RAD51, RAD54, RAD54B and ATR in response to ICL-induced replication arrest Marit Otterlei, Per Bruheim, Byungchan Ahn, Wendy Bussen, Parimal Karmakar, Kathy Baynton and Vilhelm A. Bohr Journal of Cell Science 119, 5215 (2006) doi:10.1242/jcs.03359 There was an error in the first e-press version of the article published in J. Cell Sci. 119, 5137-5146. The first e-press version of this article gave the page range as 5114-5123, whereas it should have been 5137-5146. We apologise for this mistake. Research Article 5137 Werner syndrome protein participates in a complex with RAD51, RAD54, RAD54B and ATR in response to ICL-induced replication arrest Marit Otterlei1,2,*, Per Bruheim1,3,§, Byungchan Ahn1,4,§, Wendy Bussen5, Parimal Karmakar1,6, Kathy Baynton7 and Vilhelm A. Bohr1 1Laboratory of Molecular Gerontology, National Institute on Aging, NIH, 5600 Nathan Shock Dr., Baltimore, MD 21224, USA 2Department of Cancer Research and Molecular Medicine, Laboratory Centre, Faculty of Medicine, Norwegian University of Science and Technology, Erling Skjalgsons gt. 1, N-7006 Trondheim, Norway 3Department of Biotechnology, Norwegian University of Science and Technology, N-7491 Trondheim, Norway 4Department of Life Sciences, University of Ulsan, Ulsan 680-749, Korea 5Department of Molecular Biophysics and Biochemistry, Yale University School of Medicine, 333 Cedar St, SHM-C130, New Haven, CT 06515, USA 6Department of Life Science and Biotechnology, Jadavpur University, Kolkata-700 032, WB, -

Human DNA Repair Genes Relatively Late-Diverging, Major Eukaryotic Taxa Whose Exact Order of Radiation Is Difficult to Deter- Richard D
A NALYSIS OF G ENOMIC I NFORMATION likely GTPases, as indicated by the activity of CIITA 27. L. Aravind, H. Watanabe, D. J. Lipman, E. V. Koonin, of the Integrated Protein Index and A. Uren for critical and HET-E [E. V. Koonin, L. Aravind, Trends Biochem. Proc. Natl. Acad. Sci. U.S.A. 97, 11319 (2000). reading of the manuscript and useful comments. The Sci. 25, 223 (2000)]. 28. J. R. Brown, W. F. Doolittle, Microbiol. Mol. Biol. Rev. release of the unpublished WormPep data set by The 14. T. L. Beattie, W. Zhou, M. O. Robinson, L. Harrington, 61, 456 (1997). Sanger Center is acknowledged and greatly appreciated. Curr. Biol. 8, 177 (1998). 29. We thank E. Birney and A. Bateman (The Sanger Center, 15. E. Diez, Z. Yaraghi, A. MacKenzie, P. Gros, J. Immunol. Hinxton, UK) for kindly providing the preliminary version 25 October 2000; accepted 18 January 2001 164, 1470 (2000). 16. A. M. Verhagen et al., Cell 102, 43 (2000). 17. L. Goyal, K. McCall, J. Agapite, E. Hartwieg, H. Steller, EMBO J. 19, 589 (2000). 18. The eukaryotic crown group is the assemblage of Human DNA Repair Genes relatively late-diverging, major eukaryotic taxa whose exact order of radiation is difficult to deter- Richard D. Wood,1* Michael Mitchell,2 John Sgouros,2 mine with confidence. The crown group includes the 1 multicellular eukaryotes (animals, fungi, and plants) Tomas Lindahl and some unicellular eukaryotic lineages such as slime molds and Acanthamoebae [A. H. Knoll, Science 256, 622 (1992); S. Kumar, A. Rzhetsky, J. Mol. -

University of Cincinnati
UNIVERSITY OF CINCINNATI _____________ , 20 _____ I,______________________________________________, hereby submit this as part of the requirements for the degree of: ________________________________________________ in: ________________________________________________ It is entitled: ________________________________________________ ________________________________________________ ________________________________________________ ________________________________________________ Approved by: ________________________ ________________________ ________________________ ________________________ ________________________ BLM is a Suppressor of DNA Recombination A dissertation submitted to the Division of Research and Advanced Studies Of the University of Cincinnati In partial fulfillment of the Requirements for the degree of Doctorate of Philosophy (Ph.D.) In the Department of Molecular Genetics, Microbiology, and Biochemistry Of the College of Arts and Sciences 2002 by Joel E. Straughen B.S., The Ohio State University, 1985 M.D., University of Cincinnati, 2002 Committee Chair: Joanna Groden, Ph.D. i ABSTRACT Bloom’s syndrome (BS) is a rare, recessive chromosome breakage disorder characterized by small stature, sun sensitivity, facial erythema, immunodeficiency, female subfertility, male infertility, and a predisposition to a variety of cancers. When this body of work was started, the gene for Bloom’s syndrome (BLM)hadyettobe identified. This work presents characterization of the genomic region at BLM and the identification of BLM. With the cloning -

The Werner Syndrome Protein Is Distinguished from the Bloom Syndrome Protein by Its Capacity to Tightly Bind Diverse DNA Structures
The Werner Syndrome Protein Is Distinguished from the Bloom Syndrome Protein by Its Capacity to Tightly Bind Diverse DNA Structures Ashwini Kamath-Loeb1, Lawrence A. Loeb1, Michael Fry2* 1 Department of Pathology, The Gottstein Memorial Cancer Research Center, University of Washington, Seattle, Washington, United States of America, 2 Department of Biochemistry, Rappaport Faculty of Medicine, Technion - Israel Institute of Technology, Haifa, Israel Abstract Loss of Werner syndrome helicase-exonuclease (WRN) or of its homolog Bloom syndrome helicase (BLM) results in different inherited disorders. Whereas Werner syndrome is characterized by premature onset of aging and age-associated diseases, Bloom syndrome involves developmental abnormalities and increased predisposition to diverse malignancies. To identify biochemical differences between WRN and BLM that might contribute to the dissimilar outcomes of their loss, we compared their abilities to unwind and bind in vitro diverse DNA structures. Full-length recombinant WRN and BLM proteins expressed in and purified from Sf9 insect cells unwound to comparable extents and with similar Km values partial DNA duplex, splayed arm DNA and G’2 bimolecular quadruplex DNA. However, WRN resolved bubble DNA ,25-fold more efficiently than BLM. The two enzymes were mainly distinguished by their contrasting abilities to bind DNA. WRN bound partial duplexes, bubble and splayed arm DNA and G’2 bimolecular and G4 four-molecular quadruplexes with dissociation constants of 0.25 to 25 nM. By contrast, BLM formed substantial complexes with only G4 quadruplex DNA while binding only marginally other DNA structures. We raise the possibility that in addition to its enzymatic activities WRN may act as a scaffold for the assembly on DNA of additional DNA processing proteins. -

Crystal Structure of the Werner's Syndrome Helicase
bioRxiv preprint doi: https://doi.org/10.1101/2020.05.04.075176; this version posted May 5, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Crystal Structure of the Werner’s Syndrome Helicase Joseph A. Newman1, Angeline E. Gavard1, Simone Lieb2, Madhwesh C. Ravichandran2, Katja Hauer2, Patrick Werni2, Leonhard Geist2, Jark Böttcher2, John. R. Engen3, Klaus Rumpel2, Matthias Samwer2, Mark Petronczki2 and Opher Gileadi1,* 1- Structural Genomics Consortium, University of Oxford, ORCRB, Roosevelt Drive, Oxford, United Kingdom. 2- Boehringer Ingelheim RCV GmbH & Co KG, Vienna, Austria. 3- Department of Chemistry and Chemical Biology, Northeastern University, Boston, MA 02115, USA. Abstract Werner syndrome helicase (WRN) plays important roles in DNA repair and the maintenance of genome integrity. Germline mutations in WRN give rise to Werner syndrome, a rare autosomal recessive progeroid syndrome that also features cancer predisposition. Interest in WRN as a therapeutic target has increased massively following the identification of WRN as the top synthetic lethal target for microsatellite instable (MSI) cancers. High throughput screens have identified candidate WRN helicase inhibitors, but the development of potent, selective inhibitors would be significantly enhanced by high-resolution structural information.. In this study we have further characterized the functions of WRN that are required for survival of MSI cancer cells, showing that ATP binding and hydrolysis are required for complementation of siRNA-mediated WRN silencing. A crystal structure of the WRN helicase core at 2.2 Å resolutionfeatures an atypical mode of nucleotide binding with extensive contacts formed by motif VI, which in turn defines the relative positioning of the two RecA like domains. -

Roles of the Werner Syndrome Recq Helicase in DNA Replication
dna repair 7 (2008) 1776–1786 available at www.sciencedirect.com journal homepage: www.elsevier.com/locate/dnarepair Mini-review Roles of the Werner syndrome RecQ helicase in DNA replication Julia M. Sidorova ∗ Department of Pathology, University of Washington, Seattle, WA 98195-7705, USA article info abstract Article history: Congenital deficiency in the WRN protein, a member of the human RecQ helicase family, Received 22 July 2008 gives rise to Werner syndrome, a genetic instability and cancer predisposition disorder with Accepted 23 July 2008 features of premature aging. Cellular roles of WRN are not fully elucidated. WRN has been Published on line 6 September 2008 implicated in telomere maintenance, homologous recombination, DNA repair, and other processes. Here I review the available data that directly address the role of WRN in preserv- Keywords: ing DNA integrity during replication and propose that WRN can function in coordinating Werner syndrome replication fork progression with replication stress-induced fork remodeling. I further dis- RecQ helicase cuss this role of WRN within the contexts of damage tolerance group of regulatory pathways, Human cell culture and redundancy and cooperation with other RecQ helicases. Replication stress Published by Elsevier B.V. Damage tolerance Contents 1. Introduction ................................................................................................................. 1777 1.1. Is S phase prolonged in WRN-deficient cells? ...................................................................... 1777 1.2. What is the mechanism of the S phase extension in the absence of WRN? ...................................... 1778 1.3. Are all forks equal when it comes to WRN? ........................................................................ 1779 1.4. A mechanistic model for WRN role in replication elongation ..................................................... 1779 1.5. WRN and damage tolerance pathways ............................................................................. 1781 1.6. -

WRN-Mutated Colorectal Cancer Is Characterized by a Distinct Genetic Phenotype
cancers Article WRN-Mutated Colorectal Cancer Is Characterized by a Distinct Genetic Phenotype Kai Zimmer 1, Alberto Puccini 2 , Joanne Xiu 3, Yasmine Baca 3, Gilbert Spizzo 1,4, Heinz-Josef Lenz 2, Francesca Battaglin 2 , Richard M. Goldberg 5 , Axel Grothey 6, Anthony F. Shields 7, Mohamed E. Salem 8, John L. Marshall 9, W. Michael Korn 3, Dominik Wolf 1, Florian Kocher 1 and Andreas Seeber 1,* 1 Department of Hematology and Oncology, Comprehensive Cancer Center Innsbruck, Innsbruck Medical University, 6020 Innsbruck, Austria; [email protected] (K.Z.); [email protected] (G.S.); [email protected] (D.W.); fl[email protected] (F.K.) 2 Norris Comprehensive Cancer Center, Keck School of Medicine, University of Southern California, Los Angeles, CA 90033, USA; [email protected] (A.P.); [email protected] (H.-J.L.); [email protected] (F.B.) 3 Caris Life Sciences, Phoenix, AZ 85040, USA; [email protected] (J.X.); [email protected] (Y.B.); [email protected] (W.M.K.) 4 Department of Internal Medicine, Oncologic Day Hospital, Bressanone Hospital (SABES-ASDAA), 39042 Bressanone-Brixen, Italy 5 West Virginia University Cancer Institute, Morgantown, WV 26506, USA; [email protected] 6 West Cancer Center, Germantown, TN 38138, USA; [email protected] 7 Department of Oncology, Karmanos Cancer Institute, Wayne State University, Detroit, MI 48201, USA; [email protected] 8 Levine Cancer Institute, Charlotte, NC 28204, USA; [email protected] 9 Ruesch Center for the Cure of Gastrointestinal Cancers, Lombardi Comprehensive Cancer Center, Georgetown University Medical Center, Washington, DC 20057, USA; [email protected] * Correspondence: [email protected]; Tel.: +43-50504-23001 Received: 20 March 2020; Accepted: 19 May 2020; Published: 22 May 2020 Abstract: Werner syndrome gene (WRN) contributes to DNA repair. -

A Study of Genomic Instability in Chronic Lymphocytic Leukaemia
A Study of Genomic Instability in Chronic Lymphocytic Leukaemia (CLL) Thesis submitted in accordance with the requirements of the University of Liverpool for the degree of Doctor of Philosophy by Faris Jamal M Tayeb March, 2018 0 Abstract of thesis Chronic Lymphocytic Leukaemia (CLL) is a malignancy of mature B cells. The median age at diagnosis is 70-years old, mostly seen in Western Societies. Half of the patients show an indolent phenotype and watchful waiting is the recommended approach for their management. However, once treated, patients are heterogeneous in terms of response and relapse. Regarding prognosis, genetic testing, alongside current clinical staging, is important to guide treatment and prognosis. Deletion of the short arm of chromosome 17 (17p) is one of the worst prognostic markers for CLL and usually involves loss of heterozygosity (LOH) and mutations in the TP53 gene. P53 is one of the cell-cycle regulators that activates senescence, cell-cycle arrest, DNA repair or apoptosis as part of the DNA damage response (DDR). In some severe cases of CLL with inactivated P53, the DDR is affected in favour of CLL survival. However, there is a little knowledge regarding the relationship between TP53 and mutations affecting other DNA repair genes and whether these could lead to a synergistic effect. It was, therefore, the aim of this study to address this important question. Genomic DNA was extracted from blood CLL cells of 10 patients, all of whom were bearing mutation(s) in TP53 as identified with Sanger sequencing and had progressive disease at the time of sampling. 194 known human DNA maintenance genes were identified and biotinylated-cRNA probes designed (Agilent SureSelect) to enrich DNA from their exons (2786 regions - total size = 500kb) for sequencing using an Ion Torrent Personal Genome Machine (PGM). -

Phd Thesis, University of Helsinki, Finland, 2008
FACULTY OF HEALTH AND MEDICAL SCIENCES UNIVERSITY OF COPENHAGEN An analysis of DNA repair mechanisms and mitochondrial function in accelerated aging disorders Ph.D. thesis Martin Borch Jensen Submitted: January 15, 2013 Supervisors: Professor Lene Juel Rasmussen & Dr. Professor Vilhelm A. Bohr An analysis of DNA repair mechanisms and mitochondrial function in accelerated aging disorders Martin Borch Jensen January 15th 2013 Preface & acknowledgements This thesis is submitted as part of the Ph.D. program at University of Copenhagen. Along with coursework comprising 30 ECTS (half of one academic year), the research presented herein is the product of three years as a Ph.D. student, starting January 2010, where I have worked partly at the Center for Healthy Aging, University of Copenhagen, and partly at the Laboratory of Molecular Gerontology, National Institute on Aging, NIH. In this period I was supervised by Professor Lene Juel Rasmussen and Doctor Professor Vilhelm A. Bohr, and have been attached to the \Molecular Mechanisms of Disease" research school at the University of Copenhagen. First and foremost, I would like to thank my advisors Vilhelm and Lene for giving me the opportunity to spend three years doing research on a topic that I would no doubt have pursued regardless. I am fairly confident that both my enjoyment and scientific output has been significantly higher than if I had been tackling the problem of aging in my back yard, and receiving a salary throughout this period can only be described as a luxury. Though not my advisor in any real capacity, I feel fortunate that Mansour Akbari ar- rived at the Copenhagen lab not long after I did.