University of Cincinnati
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Variant Requirements for DNA Repair Proteins in Cancer Cell Lines That Use
Variant requirements for DNA repair proteins in cancer cell lines that use alternative lengthening of telomere mechanisms of elongation DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Alaina Rae Martinez Biomedical Sciences Graduate Program The Ohio State University 2016 Dissertation Committee: Dr. Jeffrey D. Parvin, Advisor Dr. Joanna Groden Dr. Amanda E. Toland Dr. Kay F. Huebner Copyright by Alaina Rae Martinez 2016 Abstract The human genome relies on DNA repair proteins and the telomere to maintain genome stability. Genome instability is recognized as a hallmark of cancer, as is limitless replicative capacity. Cancer cells require telomere maintenance to enable this uncontrolled growth. Most often telomerase is activated, although a subset of human cancers depend on recombination-based mechanisms known as Alternative Lengthening of Telomeres (ALT). ALT depends invariably on recombination and its associated DNA repair proteins to extend telomeres. This study tested the hypothesis that the requirement for those requisite recombination proteins include other types of DNA repair proteins. These functions were tested in ALT cell lines using C-circle abundance as a marker of ALT. The requirement for homologous recombination proteins and other DNA repair proteins varied between ALT cell lines compared. Several proteins essential for homologous recombination were dispensable for C-circle production in some ALT cell lines, while proteins grouped into excision DNA repair processes were required for C- circle production. The MSH2 mismatch repair protein was required for telomere recombination by intertelomeric exchange. In sum, our study suggests that ALT proceeds by multiple mechanisms that differ between human cancer cell lines and that some of these depend on DNA repair proteins not associated with homologous recombination pathways. -

Structure and Function of the Human Recq DNA Helicases
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2005 Structure and function of the human RecQ DNA helicases Garcia, P L Posted at the Zurich Open Repository and Archive, University of Zurich ZORA URL: https://doi.org/10.5167/uzh-34420 Dissertation Published Version Originally published at: Garcia, P L. Structure and function of the human RecQ DNA helicases. 2005, University of Zurich, Faculty of Science. Structure and Function of the Human RecQ DNA Helicases Dissertation zur Erlangung der naturwissenschaftlichen Doktorw¨urde (Dr. sc. nat.) vorgelegt der Mathematisch-naturwissenschaftlichen Fakultat¨ der Universitat¨ Z ¨urich von Patrick L. Garcia aus Unterseen BE Promotionskomitee Prof. Dr. Josef Jiricny (Vorsitz) Prof. Dr. Ulrich H ¨ubscher Dr. Pavel Janscak (Leitung der Dissertation) Z ¨urich, 2005 For my parents ii Summary The RecQ DNA helicases are highly conserved from bacteria to man and are required for the maintenance of genomic stability. All unicellular organisms contain a single RecQ helicase, whereas the number of RecQ homologues in higher organisms can vary. Mu- tations in the genes encoding three of the five human members of the RecQ family give rise to autosomal recessive disorders called Bloom syndrome, Werner syndrome and Rothmund-Thomson syndrome. These diseases manifest commonly with genomic in- stability and a high predisposition to cancer. However, the genetic alterations vary as well as the types of tumours in these syndromes. Furthermore, distinct clinical features are observed, like short stature and immunodeficiency in Bloom syndrome patients or premature ageing in Werner Syndrome patients. Also, the biochemical features of the human RecQ-like DNA helicases are diverse, pointing to different roles in the mainte- nance of genomic stability. -

Folic Acid Antagonists: Antimicrobial and Immunomodulating Mechanisms and Applications
International Journal of Molecular Sciences Review Folic Acid Antagonists: Antimicrobial and Immunomodulating Mechanisms and Applications Daniel Fernández-Villa 1, Maria Rosa Aguilar 1,2 and Luis Rojo 1,2,* 1 Instituto de Ciencia y Tecnología de Polímeros, Consejo Superior de Investigaciones Científicas, CSIC, 28006 Madrid, Spain; [email protected] (D.F.-V.); [email protected] (M.R.A.) 2 Consorcio Centro de Investigación Biomédica en Red de Bioingeniería, Biomateriales y Nanomedicina, 28029 Madrid, Spain * Correspondence: [email protected]; Tel.: +34-915-622-900 Received: 18 September 2019; Accepted: 7 October 2019; Published: 9 October 2019 Abstract: Bacterial, protozoan and other microbial infections share an accelerated metabolic rate. In order to ensure a proper functioning of cell replication and proteins and nucleic acids synthesis processes, folate metabolism rate is also increased in these cases. For this reason, folic acid antagonists have been used since their discovery to treat different kinds of microbial infections, taking advantage of this metabolic difference when compared with human cells. However, resistances to these compounds have emerged since then and only combined therapies are currently used in clinic. In addition, some of these compounds have been found to have an immunomodulatory behavior that allows clinicians using them as anti-inflammatory or immunosuppressive drugs. Therefore, the aim of this review is to provide an updated state-of-the-art on the use of antifolates as antibacterial and immunomodulating agents in the clinical setting, as well as to present their action mechanisms and currently investigated biomedical applications. Keywords: folic acid antagonists; antifolates; antibiotics; antibacterials; immunomodulation; sulfonamides; antimalarial 1. -

The Bloom Syndrome Protein Limits the Lethality Associated with RAD51 Deficiency
Published OnlineFirst March 9, 2010; DOI: 10.1158/1541-7786.MCR-09-0534 Molecular DNA Damage and Cellular Stress Responses Cancer Research The Bloom Syndrome Protein Limits the Lethality Associated with RAD51 Deficiency Kenza Lahkim Bennani-Belhaj1,2, Sébastien Rouzeau1,2, Géraldine Buhagiar-Labarchède1,2, Pauline Chabosseau1,2, Rosine Onclercq-Delic1,2, Emilie Bayart1, Fabrice Cordelières3,4, Jérôme Couturier5,6, and Mounira Amor-Guéret1,2 Abstract Little is known about the functional interaction between the Bloom's syndrome protein (BLM) and the re- combinase RAD51 within cells. Using RNA interference technology, we provide the first demonstration that RAD51 acts upstream from BLM to prevent anaphase bridge formation. RAD51 downregulation was associated with an increase in the frequency of BLM-positive anaphase bridges, but not of BLM-associated ultrafine bridges. Time-lapse live microscopy analysis of anaphase bridge cells revealed that BLM promoted cell survival in the absence of Rad51. Our results directly implicate BLM in limiting the lethality associated with RAD51 deficiency through the processing of anaphase bridges resulting from the RAD51 defect. These findings provide insight into the molecular basis of some cancers possibly associated with variants of the RAD51 gene family. Mol Cancer Res; 8(3); 385–94. ©2010 AACR. Introduction cently, SUMOylation of BLM has been shown to regulate its association with RAD51 and its function in HR-medi- Bloom's syndrome displays one of the strongest known ated repair of damaged replication forks (13). In several correlations between chromosomal instability and a high models, it has been proposed that BLM restarts replication risk of cancer at an early age. -

A National Strategy for the Elimination of Hepatitis B and C: Phase Two Report
THE NATIONAL ACADEMIES PRESS This PDF is available at http://www.nap.edu/24731 SHARE A National Strategy for the Elimination of Hepatitis B and C: Phase Two Report DETAILS 296 pages | 6 x 9 | PAPERBACK ISBN 978-0-309-45729-3 | DOI: 10.17226/24731 CONTRIBUTORS GET THIS BOOK Gillian J. Buckley and Brian L. Strom, Editors; Committee on a National Strategy for the Elimination of Hepatitis B and C; Board on Population Health and Public Health Practice; Health and Medicine FIND RELATED TITLES Division; National Academies of Sciences, Engineering, and Medicine Visit the National Academies Press at NAP.edu and login or register to get: – Access to free PDF downloads of thousands of scientific reports – 10% off the price of print titles – Email or social media notifications of new titles related to your interests – Special offers and discounts Distribution, posting, or copying of this PDF is strictly prohibited without written permission of the National Academies Press. (Request Permission) Unless otherwise indicated, all materials in this PDF are copyrighted by the National Academy of Sciences. Copyright © National Academy of Sciences. All rights reserved. A National Strategy for the Elimination of Hepatitis B and C: Phase Two Report Gillian J. Buckley and Brian L. Strom, Editors Committee on a National Strategy for the Elimination of Hepatitis B and C Board on Population Health and Public Health Practice Health and Medicine Division A Report of Copyright © National Academy of Sciences. All rights reserved. A National Strategy for the Elimination of Hepatitis B and C: Phase Two Report THE NATIONAL ACADEMIES PRESS 500 Fifth Street, NW Washington, DC 20001 This activity was supported by the American Association for the Study of Liver Diseases, the Infectious Diseases Society of America, the National Viral Hepatitis Roundtable, and the U.S. -

Pharmacology
Step I Pharmacology 'WSMU~ is a ~aint program 0~the lfederatlan° a"• State Meaieal Boaras a~the I!Jntitea States • lne, and the Natianal Baara a1 Medieal Examiners. USMLE·Step 1 Pharmacology Lecture Notes 2006-2007 Edition KAPLA~. I meulca • USMLE is a joint program of the Federation of State Medical Boards of the United States, Inc. and the National Board of Medical Examiners. ©2006 Kaplan, Inc. All rights reserved. No part of this book may be reproduced in any form, by photostat, microfilm, xerography or any other means, or incorporated into any information retrieval system, electronic or mechanical, without the written permission of Kaplan, Inc. Not for resale. Author Lionel P. Rayman, Pharm.D., Ph.D. Department of Pathology Forensic Toxicology Laboratory University of Miami School of Medicine Miami, FL Contributing Authors Director of Medical Curriculum Sonia Reichert, M.D. Craig Davis, Ph.D. Associate Professor Editorial Director University of South Carolina School of Medicine Department of Pharmacology, Physiology, and Neuroscience Kathlyn McGreevy Columbia, SC Production Manager Maris Victor Nora, Pharm.D., Ph.D. Michael Wolff Associate Professor Rush Medical College Production Editor Chicago.Tl. William Ng Anthony Trevor, Ph.D. Cover Design Professor Emeritus Joanna Myllo Department of Cellular and Molecular Pharmacology University of California Cover Art San Francisco, CA Christine Schaar Steven R. Harris, Ph.D. Associate Dean for Basic Sciences Associate Professor of Pharmacology Pikeville College School of Osteopathic Medicine -

Key Enzymes in Cancer: Mechanism of Action and Inhibition with Anticancer Agents
University of Texas Rio Grande Valley ScholarWorks @ UTRGV Chemistry Faculty Publications and Presentations College of Sciences 2018 Key Enzymes in Cancer: Mechanism of Action and Inhibition With Anticancer Agents Debasish Bandyopadhyay The University of Texas Rio Grande Valley, [email protected] Gabriel Lopez The University of Texas Rio Grande Valley Stephanie Cantu The University of Texas Rio Grande Valley Samantha Balboa The University of Texas Rio Grande Valley Annabel Garcia The University of Texas Rio Grande Valley See next page for additional authors Follow this and additional works at: https://scholarworks.utrgv.edu/chem_fac Part of the Chemistry Commons, and the Life Sciences Commons Recommended Citation Debasish Bandyopadhyay, Gabriel Lopez, Stephanie Cantu, Samantha Balboa, Annabel Garcia, Christina Silva, Diandra Valdes. In Chemistry Research and Applications (Vol. 2): Organic and Medicinal Chemistry, Chapter 9; Key Enzymes in Cancer: Mechanism of Action and Inhibition with Anticancer Agents. 2018, Nova Science Publishers, Inc., Hauppauge, New York, USA (ISBN: 978-1-53614-855-8). This Book is brought to you for free and open access by the College of Sciences at ScholarWorks @ UTRGV. It has been accepted for inclusion in Chemistry Faculty Publications and Presentations by an authorized administrator of ScholarWorks @ UTRGV. For more information, please contact [email protected], [email protected]. Authors Debasish Bandyopadhyay, Gabriel Lopez, Stephanie Cantu, Samantha Balboa, Annabel Garcia, Christina Silva, and Diandra Valdes This book is available at ScholarWorks @ UTRGV: https://scholarworks.utrgv.edu/chem_fac/131 In: Organic and Medicinal Chemistry, Volume 2 ISBN: 978-1-53614-855-8 Editor: Bimal Krishna Banik © 2019 Nova Science Publishers, Inc. -

Curriculum Vitae
CURRICULUM VITAE NAME: Patricia Lynn Opresko BUSINESS ADDRESS: University of Pittsburgh Graduate School of Public Health Department of Environmental and UPMC Hillman Cancer Center 5117 Centre Avenue, Suite 2.6a Pittsburgh, PA15213-1863 Phone: 412-623-7764 Fax: 412-623-7761 E-mail: [email protected] EDUCATION AND TRAINING Undergraduate 1990 - 1994 DeSales University B.S., 1994 Chemistry and Center Valley, PA Biology Graduate 1994 - 2000 Pennsylvania State Ph.D., 2000 Biochemistry and University, College of Molecular Biology Medicine, Hershey, PA Post-Graduate 3/2000 - 5/2000 Pennsylvania State Postdoctoral Dr. Kristin Eckert, University, College of Fellow Mutagenesis and Medicine, Jake Gittlen Cancer etiology Cancer Research Institute Hershey, PA 2000-2005 National Institute on IRTA Postdoctoral Dr. Vilhelm Bohr Aging, National Fellow Molecular Institutes of Health, Gerontology and Baltimore, MD DNA Repair 1 APPOINTMENTS AND POSITIONS Academic 8/1/2018 – Co-leader Genome Stability Program, UPMC present Hillman Cancer Center 5/1/2018- Tenured Professor Pharmacology and Chemical Biology, present School of Medicine, University of Pittsburgh, Pittsburgh, PA 2/1/2018- Tenured Professor Environmental and Occupational Health, present Graduate School of Public Health, University of Pittsburgh, Pittsburgh, PA 2014 – Tenured Associate Environmental and Occupational Health, 1/31/2018 Professor Graduate School of Public Health, University of Pittsburgh, Pittsburgh, PA 2005 - 2014 Assistant Professor Environmental and Occupational Health, Graduate School -

Scaffolding Protein SPIDR/KIAA0146 Connects the Bloom Syndrome Helicase with Homologous Recombination Repair
Scaffolding protein SPIDR/KIAA0146 connects the Bloom syndrome helicase with homologous recombination repair Li Wan1, Jinhua Han1, Ting Liu1, Shunli Dong, Feng Xie, Hongxia Chen, and Jun Huang2 Life Sciences Institute, Zhejiang University, Hangzhou, Zhejiang 310058, China Edited by James E. Cleaver, University of California, San Francisco, CA, and approved February 26, 2013 (received for review December 1, 2012) The Bloom syndrome gene product, BLM, is a member of the highly of the SDSA pathway (6, 7). The ability of BLM to yield non- conserved RecQ family. An emerging concept is the BLM helicase crossover products is thought to play a critical role in the avoidance collaborates with the homologous recombination (HR) machinery to of chromosomal rearrangements during the homolog-directed re- help avoid undesirable HR events and to achieve a high degree of pair of chromosomal lesions. As a result, cells defective for BLM fidelity during the HR reaction. However, exactly how such coordina- exhibit elevated rates of sister chromatid exchange (SCE) (19–21). tion occurs in vivo is poorly understood. Here, we identified a protein Upon the occurrence of DNA damage, BLM is able to form termed SPIDR (scaffolding protein involved in DNA repair) as the link discrete foci, where it colocalizes with other DNA repair proteins between BLM and the HR machinery. SPIDR independently interacts (22, 23). However, mechanistically how BLM is recruited to sites with BLM and RAD51 and promotes the formation of a BLM/RAD51- of DNA damage and how it collaborates with other proteins to containing complex of biological importance. Consistent with its role mediate recombination repair remain largely unexplored. -
Neurodegeneration in Accelerated Aging
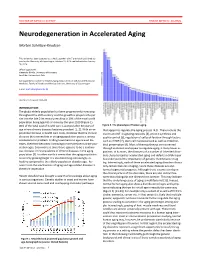
DOCTOR OF MEDICAL SCIENCE DANISH MEDICAL JOURNAL Neurodegeneration in Accelerated Aging Morten Scheibye-Knudsen This review has been accepted as a thesis together with 7 previously published pa- pers by the University of Copenhagen, October 16, 2014 and defended on January 14, 2016 Official opponents: Alexander Bürkle, University of Konstanz Lars Eide, University of Oslo Correspondence: Center for Healthy Aging, Department of Cellular and Molecular Medicine, Faculty of Health and Medical Sciences, University of Copenhagen E-mail: [email protected] Dan Med J 2016;63(11):B5308 INTRODUCTION The global elderly population has been progressively increasing throughout the 20th century and this growth is projected to per- sist into the late 21st century resulting in 20% of the total world population being aged 65 or more by the year 2100 (Figure 1). 80% of the total cost of health care is accrued after 40 years of Figure 2. The phenotype of human aging. age where chronic diseases become prevalent [1, 2]. With an ex- that appear to regulate the aging process [4,5]. These include the ponential increase in health care costs, it follows that the chronic insulin and IGF-1 signaling cascades [4], protein synthesis and diseases that accumulate in an aging population poses a serious quality control [6], regulation of cell proliferation through factors socioeconomic problem. Finding treatments to age related dis- such as mTOR [7], stem cell maintenance 8 as well as mitochon- eases, therefore becomes increasingly more pertinent as the pop- drial preservation [9]. Most of these pathways are conserved ulation ages. Even more so since there appears to be a continu- through evolution and appear to regulate aging in many lower or- ous increase in the prevalence of chronic diseases in the aging ganisms. -

Haplotype Patterns in Cancer-Related Genes with Long-Range Linkage Disequilibrium: No Evidence of Association with Breast Cancer Or Positive Selection
European Journal of Human Genetics (2008) 16, 252–260 & 2008 Nature Publishing Group All rights reserved 1018-4813/08 $30.00 www.nature.com/ejhg ARTICLE Haplotype patterns in cancer-related genes with long-range linkage disequilibrium: no evidence of association with breast cancer or positive selection Gloria Ribas*,1, Roger L Milne2, Anna Gonzalez-Neira2 and Javier Benı´tez1,2 1Human Genetics Group, Human Cancer Genetics Programme, Spanish National Cancer Centre (CNIO), Madrid, Spain; 2National Genotyping Centre (CeGen), Human Cancer Genetics Programme, Spanish National Cancer Centre (CNIO), Madrid, Spain The average length of linkage disequilibrium (LD) blocks in European populations is about 22 kb. In this study, we have selected 20 genes with LD blocks larger than 60 kb (with a median length of 88 kb) from a total of 121 cancer-related genes. We observed limited haplotype diversity, with an average of three haplotypes per gene accounting for more than 90% of the diversity, two of these being a Yin–Yang pair in 95% of the LD blocks. The mean frequency of the most common haplotype in the Spanish population was just below 50%, similar to those for the HapMap CEU and African samples, but lower than the 60% observed in Asian samples. Genes involved in the regulation of nucleobases and nucleic acid metabolism were overrepresented among these 20 genes with long LD blocks (eight genes ATM, BRCA1, BRCA2, ERCC6, MLH1, MSH3, RAD54B and XRCC4) relative to the other 101 cancer-related genes studied (P ¼ 1.23 Â 10À6). The ancestral haplotype was observed at a frequency greater than 3 in 67% of the genes either in the Spanish or one of the HapMap sampled populations. -

Acetylation of BLM Protein Regulates Its Function in Response to DNA Damage Cite This: RSC Adv.,2017,7,55301 Yankun Wang and Jianyuan Luo *
RSC Advances View Article Online PAPER View Journal | View Issue Acetylation of BLM protein regulates its function in response to DNA damage Cite this: RSC Adv.,2017,7,55301 Yankun Wang and Jianyuan Luo * Bloom syndrome is an autosomal recessive disease with phenotypes of cancer predisposition and premature aging caused by mutations of the blm gene. BLM belongs to the RecQ DNA helicase family and functions in maintaining genomic stability. In this study, we found that several lysine residues of BLM were acetylated in cells. The dynamic acetylation levels of BLM were regulated by CBP/p300 and SIRT1. Received 15th June 2017 We further identified that five lysines, K476, K863, K1010, K1329, and K1411, are the major acetylation Accepted 29th November 2017 sites. Treating cells with different DNA damage agents found that acetylation of BLM was different in DOI: 10.1039/c7ra06666j response to etoposide and hydroxyurea, suggesting that BLM acetylation may have multiple functions in rsc.li/rsc-advances DNA repair. Creative Commons Attribution-NonCommercial 3.0 Unported Licence. Introduction recombination and makes DNA back to integrated condition.14 On the other hand, BLM interacts with 53BP1 and completes Bloom syndrome protein (BLM), coded by the blm gene, is the repair in the NHEJ pathway.11 It has been found that BLM is a 1417 amino acid protein. Mutations or deletions of the blm sensitive to multiple stress factors, including hydroxyurea (HU), gene lead to Bloom Syndrome (BS).1 It is an inherited etoposide and ionizing radiation (IR) which all