In K-Ras–Mutated Colorectal Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Profiling Data
Compound Name DiscoveRx Gene Symbol Entrez Gene Percent Compound Symbol Control Concentration (nM) JNK-IN-8 AAK1 AAK1 69 1000 JNK-IN-8 ABL1(E255K)-phosphorylated ABL1 100 1000 JNK-IN-8 ABL1(F317I)-nonphosphorylated ABL1 87 1000 JNK-IN-8 ABL1(F317I)-phosphorylated ABL1 100 1000 JNK-IN-8 ABL1(F317L)-nonphosphorylated ABL1 65 1000 JNK-IN-8 ABL1(F317L)-phosphorylated ABL1 61 1000 JNK-IN-8 ABL1(H396P)-nonphosphorylated ABL1 42 1000 JNK-IN-8 ABL1(H396P)-phosphorylated ABL1 60 1000 JNK-IN-8 ABL1(M351T)-phosphorylated ABL1 81 1000 JNK-IN-8 ABL1(Q252H)-nonphosphorylated ABL1 100 1000 JNK-IN-8 ABL1(Q252H)-phosphorylated ABL1 56 1000 JNK-IN-8 ABL1(T315I)-nonphosphorylated ABL1 100 1000 JNK-IN-8 ABL1(T315I)-phosphorylated ABL1 92 1000 JNK-IN-8 ABL1(Y253F)-phosphorylated ABL1 71 1000 JNK-IN-8 ABL1-nonphosphorylated ABL1 97 1000 JNK-IN-8 ABL1-phosphorylated ABL1 100 1000 JNK-IN-8 ABL2 ABL2 97 1000 JNK-IN-8 ACVR1 ACVR1 100 1000 JNK-IN-8 ACVR1B ACVR1B 88 1000 JNK-IN-8 ACVR2A ACVR2A 100 1000 JNK-IN-8 ACVR2B ACVR2B 100 1000 JNK-IN-8 ACVRL1 ACVRL1 96 1000 JNK-IN-8 ADCK3 CABC1 100 1000 JNK-IN-8 ADCK4 ADCK4 93 1000 JNK-IN-8 AKT1 AKT1 100 1000 JNK-IN-8 AKT2 AKT2 100 1000 JNK-IN-8 AKT3 AKT3 100 1000 JNK-IN-8 ALK ALK 85 1000 JNK-IN-8 AMPK-alpha1 PRKAA1 100 1000 JNK-IN-8 AMPK-alpha2 PRKAA2 84 1000 JNK-IN-8 ANKK1 ANKK1 75 1000 JNK-IN-8 ARK5 NUAK1 100 1000 JNK-IN-8 ASK1 MAP3K5 100 1000 JNK-IN-8 ASK2 MAP3K6 93 1000 JNK-IN-8 AURKA AURKA 100 1000 JNK-IN-8 AURKA AURKA 84 1000 JNK-IN-8 AURKB AURKB 83 1000 JNK-IN-8 AURKB AURKB 96 1000 JNK-IN-8 AURKC AURKC 95 1000 JNK-IN-8 -

Gene Expression Profiling of Micrornas Associated with UCA1 in Bladder Cancer Cells
INTERNATIONAL JOURNAL OF ONCOLOGY 48: 1617-1627, 2016 Gene expression profiling of microRNAs associated with UCA1 in bladder cancer cells XIAOJUAN XIE1,2, JINGJING PAN1, LIQIANG WEI2, SHOUZHEN WU3, HUILIAN HOU4, XU LI3 and WEI CHEN1 1Department of Clinical Laboratory, The First Affiliated Hospital of Xi'an Jiaotong University, Xi'an, Shaanxi 710061; 2Shaanxi Center for Clinical Laboratory, Shaanxi Provincial People's Hospital, Xi'an, Shaanxi 710068; 3Center for Translational Medicine, 4Department of Pathology, The First Affiliated Hospital of Xi'an Jiaotong University, Xi'an, Shaanxi 710061, P.R. China Received November 11, 2015; Accepted December 27, 2015 DOI: 10.3892/ijo.2016.3357 Abstract. Emerging evidence indicates that non-coding and positively correlated with miR-196a, whereas UCA1 and RNAs, such as lncRNAs and microRNAs, play important miR-196a were negatively correlated with p27kip1, which was roles in diverse diseases, such as cancer, immune diseases downregulated in bladder cancer patients. Thus, our findings and cardiovascular diseases. Interestingly, lncRNAs could provided valuable information on miRNAs associated with directly or indirectly regulate the expression of miRNAs. UCA1 in bladder cancer, which could be helpful to further However, the expression profiling of miRNAs associated with explore the related genes and molecular networks fundamental UCA1 in bladder cancer remains unknown. Here, we used in bladder cancer progression. Illumina deep sequencing to sequence miRNA libraries from both the UCA1 knockdown and normal high-expression 5637 Introduction cells. We identified 225 and 235 miRNAs expressed in 5637 cells of normal high-expression and knockdown of UCA1, Recently, emerging studies have demonstrated that 70-90% of respectively. -

New Insights in RBM20 Cardiomyopathy
Current Heart Failure Reports (2020) 17:234–246 https://doi.org/10.1007/s11897-020-00475-x TRANSLATIONAL RESEARCH IN HEART FAILURE (J BACKS & M VAN DEN HOOGENHOF, SECTION EDITORS) New Insights in RBM20 Cardiomyopathy D. Lennermann1,2 & J. Backs1,2 & M. M. G. van den Hoogenhof1,2 Published online: 13 August 2020 # The Author(s) 2020 Abstract Purpose of Review This review aims to give an update on recent findings related to the cardiac splicing factor RNA-binding motif protein 20 (RBM20) and RBM20 cardiomyopathy, a form of dilated cardiomyopathy caused by mutations in RBM20. Recent Findings While most research on RBM20 splicing targets has focused on titin (TTN), multiple studies over the last years have shown that other splicing targets of RBM20 including Ca2+/calmodulin-dependent kinase IIδ (CAMK2D) might be critically involved in the development of RBM20 cardiomyopathy. In this regard, loss of RBM20 causes an abnormal intracellular calcium handling, which may relate to the arrhythmogenic presentation of RBM20 cardiomyopathy. In addition, RBM20 presents clinically in a highly gender-specific manner, with male patients suffering from an earlier disease onset and a more severe disease progression. Summary Further research on RBM20, and treatment of RBM20 cardiomyopathy, will need to consider both the multitude and relative contribution of the different splicing targets and related pathways, as well as gender differences. Keywords RBM20 . Dilated cardiomyopathy . CaMKIIδ . Calcium handling . Gender differences . Titin Introduction (ARVC), where a small number of genes account for most of the genetic causes, DCM-causing mutations have been ob- Dilated cardiomyopathy (DCM), as defined by left ventricular served in a variety of genes of diverse ontology [2]. -

Transcriptional Regulatory Networks in Neocortical Development
Review From trans to cis: transcriptional regulatory networks in neocortical development 1 1 1,2 Mikihito Shibata , Forrest O. Gulden , and Nenad Sestan 1 Department of Neurobiology and Kavli Institute for Neuroscience, Yale School of Medicine, New Haven, CT 06510, USA 2 Department of Psychiatry and Program in Cellular Neuroscience, Neurodegeneration and Repair, Yale School of Medicine, New Haven, CT 06510, USA Transcriptional mechanisms mediated by the binding of tions among TFs and CREs form the central core of tran- transcription factors (TFs) to cis-acting regulatory ele- scriptional regulatory networks (TRNs) (Box 1) [1]. ments (CREs) in DNA play crucial roles in directing gene Interactions between TFs and CREs are complex, dynam- expression. While TFs have been extensively studied, ic,andmodulatedbyvariousepigeneticprocesses[12].While less effort has gone towards the identification and func- individual CREs may be regulated by the competitive or tional characterization of CREs and associated epigenet- cooperative binding of many TFs, multiple CREs may also ic modulation. However, owing to methodological and confer transcriptional control over the expression of any analytical advances, more comprehensive studies of single gene [12]. Furthermore, TFs and CREs regulate regulatory elements and mechanisms are now possible. transcription together with their cognate transcriptional We summarize recent progress in integrative analyses of cofactors, chromatin regulators, epigenetic modifications, these regulatory components in the development of the RNA-binding proteins, and non-coding RNAs [2,13]. This cerebral neocortex, the part of the brain involved in combinatorial and multi-level transcriptional regulation cognition and complex behavior. These studies are greatly increases the complexity of gene expression regula- uncovering not only the underlying transcriptional reg- tion, facilitating the unique spatio-temporal patterns nec- ulatory networks, but also how these networks are essary for tissue development and function [3]. -

Supplementary Table 9. Functional Annotation Clustering Results for the Union (GS3) of the Top Genes from the SNP-Level and Gene-Based Analyses (See ST4)
Supplementary Table 9. Functional Annotation Clustering Results for the union (GS3) of the top genes from the SNP-level and Gene-based analyses (see ST4) Column Header Key Annotation Cluster Name of cluster, sorted by descending Enrichment score Enrichment Score EASE enrichment score for functional annotation cluster Category Pathway Database Term Pathway name/Identifier Count Number of genes in the submitted list in the specified term % Percentage of identified genes in the submitted list associated with the specified term PValue Significance level associated with the EASE enrichment score for the term Genes List of genes present in the term List Total Number of genes from the submitted list present in the category Pop Hits Number of genes involved in the specified term (category-specific) Pop Total Number of genes in the human genome background (category-specific) Fold Enrichment Ratio of the proportion of count to list total and population hits to population total Bonferroni Bonferroni adjustment of p-value Benjamini Benjamini adjustment of p-value FDR False Discovery Rate of p-value (percent form) Annotation Cluster 1 Enrichment Score: 3.8978262119731335 Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_CC_DIRECT GO:0005886~plasma membrane 383 24.33290978 5.74E-05 SLC9A9, XRCC5, HRAS, CHMP3, ATP1B2, EFNA1, OSMR, SLC9A3, EFNA3, UTRN, SYT6, ZNRF2, APP, AT1425 4121 18224 1.18857065 0.038655922 0.038655922 0.086284383 UP_KEYWORDS Membrane 626 39.77128335 1.53E-04 SLC9A9, HRAS, -

Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association Between BMI and Adult-Onset Non- Atopic
Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association between BMI and Adult-Onset Non- Atopic Asthma Ayoung Jeong 1,2, Medea Imboden 1,2, Akram Ghantous 3, Alexei Novoloaca 3, Anne-Elie Carsin 4,5,6, Manolis Kogevinas 4,5,6, Christian Schindler 1,2, Gianfranco Lovison 7, Zdenko Herceg 3, Cyrille Cuenin 3, Roel Vermeulen 8, Deborah Jarvis 9, André F. S. Amaral 9, Florian Kronenberg 10, Paolo Vineis 11,12 and Nicole Probst-Hensch 1,2,* 1 Swiss Tropical and Public Health Institute, 4051 Basel, Switzerland; [email protected] (A.J.); [email protected] (M.I.); [email protected] (C.S.) 2 Department of Public Health, University of Basel, 4001 Basel, Switzerland 3 International Agency for Research on Cancer, 69372 Lyon, France; [email protected] (A.G.); [email protected] (A.N.); [email protected] (Z.H.); [email protected] (C.C.) 4 ISGlobal, Barcelona Institute for Global Health, 08003 Barcelona, Spain; [email protected] (A.-E.C.); [email protected] (M.K.) 5 Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain 6 CIBER Epidemiología y Salud Pública (CIBERESP), 08005 Barcelona, Spain 7 Department of Economics, Business and Statistics, University of Palermo, 90128 Palermo, Italy; [email protected] 8 Environmental Epidemiology Division, Utrecht University, Institute for Risk Assessment Sciences, 3584CM Utrecht, Netherlands; [email protected] 9 Population Health and Occupational Disease, National Heart and Lung Institute, Imperial College, SW3 6LR London, UK; [email protected] (D.J.); [email protected] (A.F.S.A.) 10 Division of Genetic Epidemiology, Medical University of Innsbruck, 6020 Innsbruck, Austria; [email protected] 11 MRC-PHE Centre for Environment and Health, School of Public Health, Imperial College London, W2 1PG London, UK; [email protected] 12 Italian Institute for Genomic Medicine (IIGM), 10126 Turin, Italy * Correspondence: [email protected]; Tel.: +41-61-284-8378 Int. -

Appendix 2. Significantly Differentially Regulated Genes in Term Compared with Second Trimester Amniotic Fluid Supernatant
Appendix 2. Significantly Differentially Regulated Genes in Term Compared With Second Trimester Amniotic Fluid Supernatant Fold Change in term vs second trimester Amniotic Affymetrix Duplicate Fluid Probe ID probes Symbol Entrez Gene Name 1019.9 217059_at D MUC7 mucin 7, secreted 424.5 211735_x_at D SFTPC surfactant protein C 416.2 206835_at STATH statherin 363.4 214387_x_at D SFTPC surfactant protein C 295.5 205982_x_at D SFTPC surfactant protein C 288.7 1553454_at RPTN repetin solute carrier family 34 (sodium 251.3 204124_at SLC34A2 phosphate), member 2 238.9 206786_at HTN3 histatin 3 161.5 220191_at GKN1 gastrokine 1 152.7 223678_s_at D SFTPA2 surfactant protein A2 130.9 207430_s_at D MSMB microseminoprotein, beta- 99.0 214199_at SFTPD surfactant protein D major histocompatibility complex, class II, 96.5 210982_s_at D HLA-DRA DR alpha 96.5 221133_s_at D CLDN18 claudin 18 94.4 238222_at GKN2 gastrokine 2 93.7 1557961_s_at D LOC100127983 uncharacterized LOC100127983 93.1 229584_at LRRK2 leucine-rich repeat kinase 2 HOXD cluster antisense RNA 1 (non- 88.6 242042_s_at D HOXD-AS1 protein coding) 86.0 205569_at LAMP3 lysosomal-associated membrane protein 3 85.4 232698_at BPIFB2 BPI fold containing family B, member 2 84.4 205979_at SCGB2A1 secretoglobin, family 2A, member 1 84.3 230469_at RTKN2 rhotekin 2 82.2 204130_at HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 81.9 222242_s_at KLK5 kallikrein-related peptidase 5 77.0 237281_at AKAP14 A kinase (PRKA) anchor protein 14 76.7 1553602_at MUCL1 mucin-like 1 76.3 216359_at D MUC7 mucin 7, -
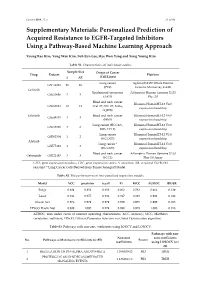
Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach
Cancers 2019, 11, x S1 of S9 Supplementary Materials: Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach Young Rae Kim, Yong Wan Kim, Suh Eun Lee, Hye Won Yang and Sung Young Kim Table S1. Characteristics of individual studies. Sample Size Origin of Cancer Drug Dataset Platform S AR (Cell Lines) Lung cancer Agilent-014850 Whole Human GSE34228 26 26 (PC9) Genome Microarray 4x44K Gefitinib Epidermoid carcinoma Affymetrix Human Genome U133 GSE10696 3 3 (A431) Plus 2.0 Head and neck cancer Illumina HumanHT-12 V4.0 GSE62061 12 12 (Cal-27, SSC-25, FaDu, expression beadchip SQ20B) Erlotinib Head and neck cancer Illumina HumanHT-12 V4.0 GSE49135 3 3 (HN5) expression beadchip Lung cancer (HCC827, Illumina HumanHT-12 V3.0 GSE38310 3 6 ER3, T15-2) expression beadchip Lung cancer Illumina HumanHT-12 V3.0 GSE62504 1 2 (HCC827) expression beadchip Afatinib Lung cancer * Illumina HumanHT-12 V4.0 GSE75468 1 3 (HCC827) expression beadchip Head and neck cancer Affymetrix Human Genome U133 Cetuximab GSE21483 3 3 (SCC1) Plus 2.0 Array GEO, gene expression omnibus; GSE, gene expression series; S, sensitive; AR, acquired EGFR-TKI resistant; * Lung Cancer Cells Derived from Tumor Xenograft Model. Table S2. The performances of four penalized regression models. Model ACC precision recall F1 MCC AUROC BRIER Ridge 0.889 0.852 0.958 0.902 0.782 0.964 0.129 Lasso 0.944 0.957 0.938 0.947 0.889 0.991 0.042 Elastic Net 0.978 0.979 0.979 0.979 0.955 0.999 0.023 EPSGO Elastic Net 0.989 1.000 0.979 0.989 0.978 1.000 0.018 AUROC, area under curve of receiver operating characteristic; ACC, accuracy; MCC, Matthews correlation coefficient; EPSGO, Efficient Parameter Selection via Global Optimization algorithm. -

Characterization of the Small Molecule Kinase Inhibitor SU11248 (Sunitinib/ SUTENT in Vitro and in Vivo
TECHNISCHE UNIVERSITÄT MÜNCHEN Lehrstuhl für Genetik Characterization of the Small Molecule Kinase Inhibitor SU11248 (Sunitinib/ SUTENT in vitro and in vivo - Towards Response Prediction in Cancer Therapy with Kinase Inhibitors Michaela Bairlein Vollständiger Abdruck der von der Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt der Technischen Universität München zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften genehmigten Dissertation. Vorsitzender: Univ. -Prof. Dr. K. Schneitz Prüfer der Dissertation: 1. Univ.-Prof. Dr. A. Gierl 2. Hon.-Prof. Dr. h.c. A. Ullrich (Eberhard-Karls-Universität Tübingen) 3. Univ.-Prof. A. Schnieke, Ph.D. Die Dissertation wurde am 07.01.2010 bei der Technischen Universität München eingereicht und durch die Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt am 19.04.2010 angenommen. FOR MY PARENTS 1 Contents 2 Summary ................................................................................................................................................................... 5 3 Zusammenfassung .................................................................................................................................................... 6 4 Introduction .............................................................................................................................................................. 8 4.1 Cancer .............................................................................................................................................................. -

Apoptotic Cells Inflammasome Activity During the Uptake of Macrophage
Downloaded from http://www.jimmunol.org/ by guest on September 29, 2021 is online at: average * The Journal of Immunology , 26 of which you can access for free at: 2012; 188:5682-5693; Prepublished online 20 from submission to initial decision 4 weeks from acceptance to publication April 2012; doi: 10.4049/jimmunol.1103760 http://www.jimmunol.org/content/188/11/5682 Complement Protein C1q Directs Macrophage Polarization and Limits Inflammasome Activity during the Uptake of Apoptotic Cells Marie E. Benoit, Elizabeth V. Clarke, Pedro Morgado, Deborah A. Fraser and Andrea J. Tenner J Immunol cites 56 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription http://www.jimmunol.org/content/suppl/2012/04/20/jimmunol.110376 0.DC1 This article http://www.jimmunol.org/content/188/11/5682.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2012 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 29, 2021. The Journal of Immunology Complement Protein C1q Directs Macrophage Polarization and Limits Inflammasome Activity during the Uptake of Apoptotic Cells Marie E. -

Cellular Processes Regulating Cytoskeletal
CELLULAR PROCESSES REGULATING CYTOSKELETAL SIGNALING, INSULIN RESISTANCE AND CALCIUM SIGNALING ARE COMMONLY MISREGULATED IN TYPE 1 AND TYPE 2 MYOTONIC DYSTROPHY PATIENTS by NRIPESH PRASAD A DISSERTATION Submitted in partial fulfillment of the requirements for the Degree of Doctor of Philosophy in The Biotechnology Science and Engineering Program to The School of Graduate Studies of The University of Alabama in Huntsville HUNTSVILLE, ALABAMA 2014 ACKNOWLEDGEMENT The journey we take is supported by many others. Acknowledgement for a few might be just a trifle thing written on a piece of paper. Nevertheless, in the true essence, it gives us an opportunity to remember and express our feelings to those, whom we love, revere and share our secrets. Here I get a great chance to express my token of thanks to people who helped and supported me to complete this journey. It is my sublime duty to express my deepest sense of gratitude and veneration to my advisor Dr. Shawn E. Levy, Director, Genomics Service Lab, HudsonAlpha Institute for Biotechnology for his sincere and indelible inspiration, constant encouragement, constructive criticism, meticulous guidance, sustained interest, immense patience, and supportive nature throughout the investigation of the the research and preparation of this manuscript. I express my deepest sense of reverence and indebtedness to the esteemed members of my Advisory Committee, Dr(s). Joseph Ng, Debra Moriarity, Devin Absher, Gerg Cooper and Luis Cruz-Vera for their valuable suggestions and encouragement at various stages of my research and thesis writing. Sincere regards are due to, Dean, Graduate School, University of Alabama in Huntsville, Huntsville, AL, USA for providing all the necessary help and support.