Genetic Structure and Molecular Diversity of Brazilian Grapevine Germplasm
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
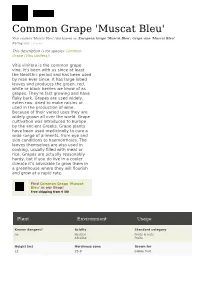
Common Grape 'Muscat Bleu' Vitis Vinifera 'Muscat Bleu' Also Known As: European Grape 'Muscat Bleu', Grape Vine 'Muscat Bleu' Rating: 0.0 ( 0 Votes)
Common Grape 'Muscat Bleu' Vitis vinifera 'Muscat Bleu' Also known as: European Grape 'Muscat Bleu', Grape vine 'Muscat Bleu' Rating: 0.0 ( 0 votes) This description is for species Common Grape (Vitis vinifera): Vitis vinifera is the common grape vine. It's been with us since at least the Neolithic period and has been used by man ever since. It has large lobed leaves and produces the green, red, white or black berries we know of as grapes. They're fast growing and have flaky bark. Grapes are used widely, eaten raw, dried to make raisins or used in the production of wine. Because of their varied uses they are widely grown all over the world. Grape cultivation was introduced to Europe by the ancient Greeks. Grape plants have been used medicinally to cure a wide range of ailments, from eye and skin conditions to haemorrhoids. The leaves themselves are also used in cooking, usually filled with meat or rice. Grapes are actually reasonably hardy, but if you do live in a cooler climate it's advisable to grow them in a greenhouse where they will flourish and grow at a rapid rate. Find Common Grape 'Muscat Bleu' in our Shop! Free shipping from € 50! Plant Environment Usage Known dangers? Acidity Standard category no Neutral Fruits & nuts Alkaline Fruits Height [m] Hardiness zone Grown for 12 Z5-9 Edible fruit Plant Environment Usage Spread [m] Heat zone Creative category 4 H9-6 Kid Approved Simple food Outdoor food Tough survivors Dominant flower colour Winter temperatures [°C] Garden type Inconspicuous or absent -29 - -1 Cottage garden Kitchen garden Mediterranean garden Flower Fragrance Heat days Garden spaces No, neutral please 45 - 150 Walls, trellises and pergolas Camouflage Flowering seasons Moisture Gardening expertise Early summer well-drained intermediate Mid summer Foliage in summer Soil type Time to reach full size Green sandy up to 20 years Clay chalky Propagation methods Sun requirements grafting Full sun seed layering Hardwood cuttings Growth habit Exposure Climbing Sheltered Twining . -

Das Weinjahr L’Année Viticole L’Anno Viticolo 2014
Eidgenössisches Departement für Wirtschaft, Bildung und Forschung WBF Bundesamt für Landwirtschaft BLW Département fédéral de l’économie, de la formation et de la recherche DEFR Office fédéral de l’agriculture OFAG Dipartimento federale dell’economia, della formazione e della ricerca DEFR Ufficio federale dell'agricoltura UFAG Das Weinjahr L’année viticole L’anno viticolo 2014 Weinwirtschaftliche Statistik Statistiques vitivinicoles Statistiche vitivinicole April / avril / aprile 2015 Inhaltsverzeichnis / Sommaire / Indice 1 Einleitung (deutsch, S. 4) 1.1 Vorbemerkungen 1.2 Zusammenfassung 1.2.1 Rebfläche 1.2.2 Ernte 1.2.3 Weinvorräte am 31. Dezember 1.2.4 Weinkonsum 1.2.5 Verarbeitungswein 1.2.6 Weineinfuhren 1 Introduction (français, p. 8) 1.1 Remarques préliminaires 1.2 Résumé 1.2.1 Surface viticole 1.2.2 Récolte 1.2.3 Stocks de vins au 31 décembre 1.2.4 Consommation de vin 1.2.5 Vin industriel 1.2.6 Importations de vins 1 Introduzione (italiano, p. 12) 1.1 Premesse 1.2 Riassunto 1.2.1 Superficie viticola 1.2.2 Raccolto 1.2.3 Scorte di vino al 31 dicembre 1.2.4 Consumo di vino 1.2.5 Vino industriale 1.2.6 Importazioni di vino 2 Rebfläche / Surface viticole 2.1 Gesamtergebnisse / Résultats généraux 2.2 Rebflächen weisser Sorten pro Kanton / Surfaces des cépages blancs par canton 2.3 Rebflächen roter Sorten pro Kanton / Surfaces des cépages rouges par canton 2.4 Rebflächen nach Sorten / Surfaces viticoles par cépages 3 Ernte / Récolte 3.1 Weinlese exkl. Traubensaft / Vendanges sans jus de raisin 3.2 Mittlere Brix-Prozente / Pour-cent -

Identification and Characterization of Swiss Grapevine Cultivars Using Microsatellite Markers
Mitteilungen Klosterneuburg 56 (2006): 147-156 Identification and characterization of Swiss grapevine cultivars using microsatellite markers ANDREA FREI, NAOMI A. PORRET, JUÈ RG E. FREY and JUÈ RG GAFNER Agroscope ACWChangins-Wa Èdenswil Forschungsanstalt fuÈ r Obst-, Wein- und Gartenbau CH-8820 WaÈdenswil, Postfach 185 E-mail: [email protected] Microsatellite analysis with up to 12 markers was performed on 463 grapevine samples mostly collected from Swiss cultivar collections, viticulturists, winegrowers and private collectors, using an optimized and fast method based on quick DNA extraction, six-plex PCR and automated fragment analysis. Several already published genetic profiles were confirmed, misnamed plants were identified, and additional new profiles are presented in this study. Some of the misnamed plants could be named correctly, while cultivar allocation of others needs confirmation. We found that 'Briegler' seems to be a synonym for 'Bondola nera', possibly the German name for a cultivar grown in Sou- thern Switzerland. In the case of 'Erlenbacher Schwarz' we found types with two different profiles. The ones from Eastern Switzerland are identical to 'Hitzkircher', while the type collected in Western Switzerland is different. Fur- ther data collection is necessary to elucidate which type is the true-to-type cultivar. Also an old and rare cultivar na- med 'MoÈrchel' was shown to be identical to 'Trollinger' (syn. 'Schiava Grossa', 'Gross-Vernatsch' or 'Frankenthal Noir'). Several 'Chasselas' and 'Muscat' types were allocated -

Weineinkaufen Bei Weinart
Die Marke der AlpenweinKultur Das Portfolio im Detail - Edition 2015 - XI Alpentaverne mit Magnothek Eigenmarken Swiss Made • Ausgezeichnete Grands Crus • Spitzen- BIO- PIWI- und Bergweine der Alpenländer • Autochthone Sorten • Raritäten • Trouvaillen • Grossflaschen VINOVITAS à votre santé ! WeinArt® No1 in Valais VS und Vaud VD Champagne SWISS SPARKLING Prosecco DOCG PIWI & BIO Weine bis zu den höchsten Rebbergen Europas Grands Vins des Alpes du Mont Ventoux via Grand Châtelard bis Leithaberg TOP Preis-Leistung! AlpenweinKultur Wissenswertes Herkunftsbezeichnungen AOC Appelation d`Origine Controlée VdT Vin de Table / Vino di Tavola VdP Vin du Pays / Landwein IGT Indicazione Geografica Tipica DOC Denominazione di Origine Cotrollata DOCG Denominazione di Origine Cotrollata e Garantita QW Qualitätswein Abkürzungen ◊ Weisswein ◊● Rosé Wein ● Rotwein * Wein mit Ausbau im grossen Holzfass ** Wein mit Ausbau im Barrique = 225 l Holzfass BVS Flasche mit Schraubverschluss DIAM Flaschenverschluss aus Recycling Korken Brut Prosecco Le Bertole Brut mit 12g/l Restzucker Dry Prosecco Supreme mit 25g/l Restzucker R Rarität E Exklusivität 1822 Le Pot Vaudois 1822 ist die Waadtländer Masseinheit = 140cl 1913 In Zusammenarbeit mit den Experten der 1913 SA - Saillon RP Robert Parker Auszeichnung TOP Dieser Vermerk wird bei Weinen angezeigt, welche unabhängig von der Qualitätsstufe ein ausgesprochen gutes Verhältnis zwischen Preis und Leistung aufweisen. Was bedeutet für Sie AlpenweinKultur? Philosophie? Geschichte? Lebenshaltung? Sprechen Sie mit uns, um mehr darüber zu erfahren. Wasser und Wein ist Leben! Gletscherwasser & Alpenwein ist Lebenskultur! Das Salz dazu erhalten Sie bei WeinArt, den Experten in AlpenweinKultur, für Ihr Wohlbefinden und Ihre Gesundheit, sei es bei einer Verkostung ohne Brot oder einem Seminar nicht nur mit Wasser. -

Muscat Bleu N
Catalogue of grapevines cultivated in France © UMT Géno-Vigne® INRA - IFV - Montpellier SupAgro http://plantgrape.plantnet-project.org Edited on 28/09/2021 Muscat Bleu N Name of the variety in France Muscat Bleu Origin Muscat bleu or 83/2 Garnier was obtained by M. Garnier in Geneva, Switzerland, in the 1930's. Based on genetic analyses carried out in Montpellier, this interspecific hybrid probably comes from a crossbreeding between 15/6 Garnier (Villard noir x Müller-Thurgau) and Muscat de Saint-Vallier (20473 Seyve-Villard). Synonyms In Austria, this variety is officially designated as "Muskat bleu". This synonym is officially recognized in France regarding plant propagation material. Legal information In France, Muscat Bleu is officially listed in the "Catalogue of vine varieties" since 2013 on the A list. This variety is also listed in the catalogue of Austria. Use Table grape variety. Evolution of cultivated areas in France 2018 ha 0.1 Descriptive elements The identification is based on: - the tip of the young shoot with no or a very low density of prostate hairs, - the young leaves with bronze spots and no prostate hairs, - the shoots with red or red-striped internodes, - the adult leaves entire or with three lobes, an open V- or U-shaped petiole sinus, large teeth, long compared to their width at the base with straight or convex sides, a low anthocyanin coloration of veins, an involute or sometimes twisted, finely blistered, slightly goffered leaf blade, and on the lower side of the leaves, no erect hairs and prostate hairs, - the round-shaped or slightly ellipsoid berries. -

European Project Grapegen 06 - Grapevine Genetic Resources - Version 21 January 2011 P
European Project GrapeGen 06 - Grapevine Genetic Resources European Grapevine Catalogue: Towards a Comprehensive List T. Lacombe, L. Audeguin, M. Boselli, B. Bucchetti, F. Cabello, M. Crespan, C. D’Onofrio, J. Eiras Dias, S. Ercisli, M. Gardiman, MS. Grando, S. Imazio, O. Jandurova, A. Jung, E. Kiss, P. Kozma, E. Maul, D. Maghradze, C. Martinez, G. Muñoz, J-K. Pátková, I. Pejic, E. Peterlunger, D. Pitsoli, D. Preiner, S. Raimondi, F. Regner, G. Savin, S. Savvides, A. Schneider, J-L. Spring, A. Szoke, A. Veres, J-M. Boursiquot, R. Bacilieri and P. This Annex 3 B : Official national catalogues of grapevine varieties for Member States of the European Union and the Third Countries partner of the GrapeGen 06 Project Legend : before the arrows, name of the variety as registered in the country . After the arrows, common prime name of the variety according to VIVC database when referenced, # identification number of the variety, species of the variety, sex (H = hermaphrodite, F = female, M = male), colour of berry skin (B = yellow-green, N = blue-black, Rg = red, Rs = rose, G = grey). Austria AUT National Catalogue version 2008 Alphonse-Lavalle (AUT) >>> ALPHONSE LAVALLEE # 349 - vinifera - H - N Angela (AUT) >>> ANGELA # 20342 - interspecific cross - H - B Aron (AUT) >>> ARON # 14014 - interspecific cross - - B Attica (AUT) >>> ATTIKA SEEDLESS # 17309 - vinifera - - Rg Attila (AUT) >>> ATTILA # 756 - vinifera - - B Bacchus (AUT) >>> BACCHUS WEISS # 851 - - H - B Bianca (AUT) >>> BIANCA # 1321 - interspecific cross - H - B Birstaler Muskat (AUT) -

Varietal Response to Sour Bunch Rot in Polish Grapevine Genetic Resources
agronomy Article Varietal Response to Sour Bunch Rot in Polish Grapevine Genetic Resources Jerzy Lisek * and Anna Lisek The National Institute of Horticultural Research, Konstytucji 3 Maja 1/3 Str., 96-100 Skierniewice, Poland; [email protected] * Correspondence: [email protected] Abstract: The aim of this study was to assess the resistance to sour rot of twenty-eight valuable cultivars of grapevine for wine production and twenty-five cultivars of table grapevine with diverse geographic and genetic origins, and to explain the causes of varied resistance based on the features related to the morphology, biology and ecology of assessed genotypes. The study was conducted for six years in the grapevine field collection of the National Institute of Horticultural Research in Skierniewice (Poland, latitude 51.9627 N, longitude 20.1666 E). Sour rot was severe in three seasons with abundant rainfall during the berry ripening stage. The number of wine and table cultivars in particular classes of resistance (mean value for three years) was as follows: very little or little—9 (wine) and 9 (table), medium—9 (wine) and 3 (table), high or very high—10 (wine) and 13 (table). The severity of bunch sour rot was positively correlated with single berry weight (moderate or weak correlation), bunch density and single bunch weight (very weak or weak correlation), and negatively correlated with thickness of berry skin (strong correlation) and the time of the beginning of veraison (weak correlation). Cultivars that were characterized by such agrobiological and ecological features as easy detachment of the berry from the pedicel, sensitivity to berry skin cracking, frequent damage to the skin by insects, and sensitivity to sunburn, were more heavily exposed to sour rot. -

Growing Dessert Grapes Outdoors
Dessert grapes outdoors Growing dessert grapes outdoors 1 2 3 4 With the right choice of cultivar, it is possible to grow and ripen dessert grapes outdoors successfully in many regions of the UK Author: Sarah Bell, owner of Sunnybank Vine Nursery in Herefordshire and holder of the Plant Heritage National Plant Collection of Vitis vinifera and hybrids here can be few sights more evocative of a Mediterranean climate than that of a Dessert grapes for a luxuriant grapevine, perhaps scrambling T range of locations (see over a sunny pergola, dripping with succulent fruit. overleaf) taken from the This scene is one that can, with consideration, be 5 6 7 8 National Plant Collection created in many parts of the UK. In fact vines have been grown outdoors in the 1 Vitis ‘Muscat Bleu’ British Isles since pre-Roman times. In recent years 2 V. vinifera ‘Lakemont’ the UK-based wine industry has been expanding at 3 V. ‘Trollhaugen’ a rapid rate, with more than 3 million vines planted 4 V. vinifera ‘Exalta’ in 2019 alone. However, most gardeners will want to 5 V. ‘Reliance’ grow selections that can be plucked and enjoyed 6 V. ‘Alden’ directly from the vine: dessert grapes. These may 7 V. ‘Glenora’ include cultivars that are usually associated with 8 V. ‘New York winemaking; many grapes are dual purpose. Muscat’ agm Whether a grape is good for winemaking or eating 9 V. ‘Brant’ agm depends on its acidity and sugar levels: dessert MARIANNE MAJERUS 10 V. ‘Boskoop Glory’ agm grapes do not need the higher levels of acidity Grapevines are common 11 V. -

Raisin De Table
Compte rendu 2011 Code essai : CEFEL 11 RAI Var 31-6 Espèce : Raisin de table Responsable essai : Agriculture Biologique Daniel LAVIGNE Evaluation variétale en culture biologique Rédigé par : Approuvé par : Page 1 sur 10 Daniel LAVIGNE Pascale WESTERCAMP Emis le 13 juin 2012 Raisin de table Evaluation variétale en culture biologique COMPTE RENDU ESSAI 2011 Muscat Bleu Fanny Candin Etude subventionnée par le Conseil Régional Midi-Pyrénées CEFEL Code essai : 11 RAI Var 31-6 Page 1 sur 10 I - Objectif de l’essai Evaluer de nouvelles variétés tolérantes aux maladies en conditions de culture biologique. II - Matériel et méthode Lieu - Parcelle producteur - Lauzerte (82) - Sol argilo-calcaire profond - Situation de vallée Matériel - Choix de variétés issues du niveau 1 du CEFEL (cf annexes 1 à 3 - fiches variétales) Dispositif - Implantation de 5 souches par variété, greffées sur R110. - Comparaison à des variétés sensibles : Muscat de Hambourg, Ribol, Alval. Dispositif de la parcelle - Culture conduite en semi-pergola (3 m x 1 m - 3330 s/ha) - Irrigation par micro-jet - Protection par filet anti-grêle (installation 2012). - Apport de matière organique sur le rang (3 t/ha) et de compost en plein (20 t/ha) Modalités comparées 3 variétés ont été plantées en juin 2011 : Candin, Fanny et Muscat Bleu (1ère production en 2013). D’autres variétés pourraient également être évaluées suivant les résultats obtenus en niveau 1 (décision du comité de pilotage raisin), comme Angela ou des variétés type Labrusca. La variété Garanth a été plantée en juin 2012. III - Résultats et discussion Premières récoltes en 2013 Les résultats de l’évaluation en niveau 1 sur le site du CEFEL St Laurent (non bio) sont donnés en annexe 4 et 5. -

2.4 Rebflächen Nach Sorten / Surfaces Viticoles Par Cépages 2013
2.4 Rebflächen nach Sorten / Surfaces viticoles par cépages 2013 Fläche Fläche Fläche Sorten / Cépages Farbe1 Sorten / Cépages Farbe1 Sorten / Cépages Farbe1 m2 m2 m 2 Acolon r 26'610 Fr 503-89r r 0 Pinot blanc w 1'065'543 Alicante Bouschet r 1'134 Freisa r 1'044 Pinot gris / Malvoisie w 2'259'591 Aligoté w 230'229 Freisamer / Freiburger w 46'362 Pinot Meunier r 1'170 Alphonse Lavallée r 15 Friulano w 256 Pinotage r 9'987 Altesse w 40'829 Frühburgunder r 10'175 Pinotin r 1'865 Amigne w 421'513 Fumin r 4'239 Piroso r 2'080 Ancellotta r 237'127 Galotta r 262'709 Primitivo / Zinfandel r 0 Angela w 900 Gamaret r 4'133'015 Prinzipal w 0 Arinarnoa r 14'105 Gamay r 14'306'805 Prior r 11'946 Arneis w 1'485 Garanoir r 2'185'276 RAC 3209 r 500 Arvine (petite) w 1'661'320 Garganega w 107 Räuschling w 225'751 Aurore w 60 Gewürztraminer w 488'204 Rebo r 2'726 Auxerrois w 32'145 GF 48-12 w 11'378 Regent r 399'461 Bacchus w 16'249 Goron de Bovernier r 698 Reichensteiner w 2'670 Baco noir r 12'799 Gouais / Gwäss /Heunisch w 8'183 Réselle w 10'877 Barbera r 6'001 Grenache r 8'861 Rèze w 23'993 Baron r 1'900 Grosse Arvine w 490 Riesling w 156'281 Bernarde w 0 Grüner Veltliner w 8'973 Rondo r 7'836 Bianca w 21'544 Helios w 2'340 Roter Milan w 0 Birstaler Muskat w 9'966 Himbertscha w 2'289 Roter Traminer r 0 Blauburgunder / Pinot noir / Clevner / Serva r 43'010'998 Humagne blanc w 301'630 Roussanne w 26'215 Blaufränkisch / Lemberger r 36'097 Humagne rouge r 1'336'920 Saint Laurent r 28'270 Bondola r 115'062 INRA8502Bx r 0 Sangiovese / Montepulciano r 9'382 -

Jean Gissinger Au Cœur De Votre Jardin
TARIFS TARIFS 2019/20 PARTICULIERS 2019/20 PARTICULIERS Jean Gissinger au cœur de votre Jardin Plus de 100 ans d’expérience pour une production locale de végétaux d’ornements et fruitiers - Certifiés Plante Bleue HPF www.jean-gissinger.fr Catalogue Automne 2019 / Printemps 2020 Catalogue Automne 2019 /Printemps 2020 Chez nous, vous disposez de végétaux acclimatés cultivés dans la région. Des atouts importants pour la réussite de vos plantations. ACCES • Depuis Mulhouse ou Strasbourg: RD 83 prendre sortie Rouffach Sud. Une fois rentré dans Rouffach, prendre au stop à gauche. • Depuis Freiburg: D 415 direction Colmar puis A 35 direction Mulhouse, sortie N° 28 (Guebwiller, Niederhergheim), puis direction Rouffach. A l’entrée de Rouffach, prendre la RD 83 direction Belfort sortie Rouffach Sud. Une fois rentré dans Rouffach, prendre au stop à gauche. • Coordonnées GPS Latitude: 47.9543437 Longitude: 7.29 Catalogue Automne 2019 /Printemps 2020 2 3 Pépinières Jean GISSINGER Le plus grand professionnel de l’Est de la France, tout près de chez vous ! VISITER NOTRE JARDIN D’EXPOSITION Nous vous invitons à visiter notre jardin d’exposition et de vente de 2000 m² ouvert toute l’année : Lundi au vendredi de 8h00 à 12h00 et de 13h30 à 17h30 Samedi de 8h00 à 12h00 et de 13h30 à 16h30 (HORMIS LE SAMEDI APRES-MIDI EN JANVIER EXEMPLE DE VEGETAUX FEUILLAGE COLORE OU DECORATIF Albizia julibrissin ‘Summer Chocolat’ – fleurs roses parfumées, feuillage couleur chocolat Liquidambar styraciflua ‘Manon’ = Copalme d’Amérique – port pyramidal, feuillage vert bordé -

DIVERSIDADE GENÉTICA DE ACESSOS DO BANCO ATIVO DE GERMOPLASMA DE Vitis Spp
INSTITUTO AGRONÔMICO PROGRAMA DE PÓS-GRADUAÇÃO AGRICULTURA TROPICAL E SUBTROPICAL DIVERSIDADE GENÉTICA DE ACESSOS DO BANCO ATIVO DE GERMOPLASMA DE Vitis spp. DO INSTITUTO AGRONÔMICO GEOVANI LUCIANO DE OLIVEIRA Orientadora: Dra. Mara Fernandes Moura Coorientadora: Dra. Lívia Moura de Souza Dissertação apresentada como requisito para a obtenção do título de Mestre em Agricultura Tropical e Subtropical, Área de Concentração em Genética, Melhoramento Vegetal e Biotecnologia. Campinas, SP 2020 Ficha elaborada pela bibliotecária do Núcleo de Informação e Documentação do Instituto Agronômico O48d Oliveira, Geovani Luciano de Diversidade genética de acessos do banco ativo de germoplasma de Vitis spp. do Instituto Agronômico / Geovani Luciano de Oliveira. Campinas, 2020. 101 fls. Orientadora: Mara Fernandes Moura Co-orientadora: Lívia Moura de Souza Dissertação (Mestrado) Agricultura Tropical e Subtropical – Instituto Agronômico 1. Melhoramento genético. 2. Coleção nuclear . 3. Recursos genéticos 3. Marcador molecular 4. SSR. I. Moura, Mara Fernandes II. Souza, Lívia Moura de III. Título CDD. 574.182 ii DEDICO A Deus e a minha família por serem minha base. Em especial aos meus pais, Adelar Bueno de Oliveira e Eva Maria de Oliveira. iii AGRADECIMENTOS Primeiramente a Deus, por ter me dado saúde, força e ânimo durante essa etapa e por ter permitido mais esta vitória. Aos meus pais Adelar e Eva, pelo amor, incentivo е apoio incondicional. Agradeço por toda ajuda que me deram durante a realização deste trabalho. Ao meu irmão Gustavo, pelo companheirismo e apoio durante toda a vida. A minha namorada Bianca, por acreditar em mim e em meu potencial, pela compreensão, carinho e atenção. Obrigado por ser essa pessoa incrível, agradeço a Deus todos os dias por ter você em minha vida.