1 Diversification of Reprogramming Trajectories Revealed by Parallel Single-Cell 2 Transcriptome and Chromatin Accessibility Sequencing
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Down-Regulation of Stem Cell Genes, Including Those in a 200-Kb Gene Cluster at 12P13.31, Is Associated with in Vivo Differentiation of Human Male Germ Cell Tumors
Research Article Down-Regulation of Stem Cell Genes, Including Those in a 200-kb Gene Cluster at 12p13.31, Is Associated with In vivo Differentiation of Human Male Germ Cell Tumors James E. Korkola,1 Jane Houldsworth,1,2 Rajendrakumar S.V. Chadalavada,1 Adam B. Olshen,3 Debbie Dobrzynski,2 Victor E. Reuter,4 George J. Bosl,2 and R.S.K. Chaganti1,2 1Cell Biology Program and Departments of 2Medicine, 3Epidemiology and Biostatistics, and 4Pathology, Memorial Sloan-Kettering Cancer Center, New York, New York Abstract on the degree and type of differentiation (i.e., seminomas, which Adult male germ cell tumors (GCTs) comprise distinct groups: resemble undifferentiated primitive germ cells, and nonseminomas, seminomas and nonseminomas, which include pluripotent which show varying degrees of embryonic and extraembryonic embryonal carcinomas as well as other histologic subtypes patterns of differentiation; refs. 2, 3). Nonseminomatous GCTs are exhibiting various stages of differentiation. Almost all GCTs further subdivided into embryonal carcinomas, which show early show 12p gain, but the target genes have not been clearly zygotic or embryonal-like differentiation, yolk sac tumors and defined. To identify 12p target genes, we examined Affymetrix choriocarcinomas, which exhibit extraembryonal forms of differ- (Santa Clara, CA) U133A+B microarray (f83% coverage of 12p entiation, and teratomas, which show somatic differentiation along genes) expression profiles of 17 seminomas, 84 nonseminoma multiple lineages (3). Both seminomas and embryonal carcinoma GCTs, and 5 normal testis samples. Seventy-three genes on 12p are known to express stem cell markers, such as POU5F1 (4) and were significantly overexpressed, including GLUT3 and REA NANOG (5). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -

Glucocorticoid Receptor Signaling Activates TEAD4 to Promote Breast
Published OnlineFirst July 9, 2019; DOI: 10.1158/0008-5472.CAN-19-0012 Cancer Molecular Cell Biology Research Glucocorticoid Receptor Signaling Activates TEAD4 to Promote Breast Cancer Progression Lingli He1,2, Liang Yuan3,Yang Sun1,2, Pingyang Wang1,2, Hailin Zhang4, Xue Feng1,2, Zuoyun Wang1,2, Wenxiang Zhang1,2, Chuanyu Yang4,Yi Arial Zeng1,2,Yun Zhao1,2,3, Ceshi Chen4,5,6, and Lei Zhang1,2,3 Abstract The Hippo pathway plays a critical role in cell growth and to the TEAD4 promoter to boost its own expression. Func- tumorigenesis. The activity of TEA domain transcription factor tionally, the activation of TEAD4 by GC promoted breast 4 (TEAD4) determines the output of Hippo signaling; how- cancer stem cells maintenance, cell survival, metastasis, and ever, the regulation and function of TEAD4 has not been chemoresistance both in vitro and in vivo. Pharmacologic explored extensively. Here, we identified glucocorticoids (GC) inhibition of TEAD4 inhibited GC-induced breast cancer as novel activators of TEAD4. GC treatment facilitated gluco- chemoresistance. In conclusion, our study reveals a novel corticoid receptor (GR)-dependent nuclear accumulation and regulation and functional role of TEAD4 in breast cancer and transcriptional activation of TEAD4. TEAD4 positively corre- proposes a potential new strategy for breast cancer therapy. lated with GR expression in human breast cancer, and high expression of TEAD4 predicted poor survival of patients with Significance: This study provides new insight into the role breast cancer. Mechanistically, GC activation promoted GR of glucocorticoid signaling in breast cancer, with potential for interaction with TEAD4, forming a complex that was recruited clinical translation. -

Full Text (PDF)
Published OnlineFirst May 1, 2014; DOI: 10.1158/0008-5472.CAN-13-3147 Cancer Tumor and Stem Cell Biology Research Loss of the Polycomb Mark from Bivalent Promoters Leads to Activation of Cancer-Promoting Genes in Colorectal Tumors Maria A. Hahn1, Arthur X. Li2, Xiwei Wu3, Richard Yang1, David A. Drew4, Daniel W. Rosenberg4, and Gerd P. Pfeifer1 Abstract In colon tumors, the transcription of many genes becomes deregulated by poorly defined epigenetic mechanisms that have been studied mainly in established cell lines. In this study, we used frozen human colon tissues to analyze patterns of histone modification and DNA cytosine methylation in cancer and matched normal mucosa specimens. DNA methylation is strongly targeted to bivalent H3K4me3- and H3K27me3-associated promoters, which lose both histone marks and acquire DNA methylation. However, we found that loss of the Polycomb mark H3K27me3 from bivalent promoters was accompanied often by activation of genes associated with cancer progression, including numerous stem cell regulators, oncogenes, and proliferation-associated genes. Indeed, we found many of these same genes were also activated in patients with ulcerative colitis where chronic inflammation predisposes them to colon cancer. Based on our findings, we propose that a loss of Polycomb repression at bivalent genes combined with an ensuing selection for tumor-driving events plays a major role in cancer progression. Cancer Res; 74(13); 3617–29. Ó2014 AACR. Introduction cells (21, 22). The acquisition of the more permanent silencing Tumorigenesis is a complex process that is driven by a mark, 5mC, at the promoters of Polycomb target genes does number of genetic and epigenetic alterations, which often not fundamentally change their expression levels although result in aberrant gene expression (1–3). -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes. -

Methylome and Transcriptome Maps of Human Visceral and Subcutaneous
www.nature.com/scientificreports OPEN Methylome and transcriptome maps of human visceral and subcutaneous adipocytes reveal Received: 9 April 2019 Accepted: 11 June 2019 key epigenetic diferences at Published: xx xx xxxx developmental genes Stephen T. Bradford1,2,3, Shalima S. Nair1,3, Aaron L. Statham1, Susan J. van Dijk2, Timothy J. Peters 1,3,4, Firoz Anwar 2, Hugh J. French 1, Julius Z. H. von Martels1, Brodie Sutclife2, Madhavi P. Maddugoda1, Michelle Peranec1, Hilal Varinli1,2,5, Rosanna Arnoldy1, Michael Buckley1,4, Jason P. Ross2, Elena Zotenko1,3, Jenny Z. Song1, Clare Stirzaker1,3, Denis C. Bauer2, Wenjia Qu1, Michael M. Swarbrick6, Helen L. Lutgers1,7, Reginald V. Lord8, Katherine Samaras9,10, Peter L. Molloy 2 & Susan J. Clark 1,3 Adipocytes support key metabolic and endocrine functions of adipose tissue. Lipid is stored in two major classes of depots, namely visceral adipose (VA) and subcutaneous adipose (SA) depots. Increased visceral adiposity is associated with adverse health outcomes, whereas the impact of SA tissue is relatively metabolically benign. The precise molecular features associated with the functional diferences between the adipose depots are still not well understood. Here, we characterised transcriptomes and methylomes of isolated adipocytes from matched SA and VA tissues of individuals with normal BMI to identify epigenetic diferences and their contribution to cell type and depot-specifc function. We found that DNA methylomes were notably distinct between diferent adipocyte depots and were associated with diferential gene expression within pathways fundamental to adipocyte function. Most striking diferential methylation was found at transcription factor and developmental genes. Our fndings highlight the importance of developmental origins in the function of diferent fat depots. -

Transitions in Cell Potency During Early Mouse Development Are Driven by Notch
bioRxiv preprint doi: https://doi.org/10.1101/451922; this version posted October 24, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Menchero et al. Transitions in cell potency during early mouse development are driven by Notch Sergio Menchero1, Antonio Lopez-Izquierdo1, Isabel Rollan1, Julio Sainz de Aja1, Ϯ, Maria Jose Andreu1, Minjung Kang2, Javier Adan1, Rui Benedito3, Teresa Rayon1, †, Anna-Katerina Hadjantonakis2 and Miguel Manzanares1,* 1 Centro Nacional de Investigaciones Cardiovasculares Carlos III (CNIC), Melchor Fernández Almagro 3, 28029 Madrid, Spain. 2 Developmental Biology Program, Sloan Kettering Institute, New York NY 10065, USA. 3 Molecular Genetics of Angiogenesis Group, Centro Nacional de Investigaciones Cardiovasculares Carlos III (CNIC), Melchor Fernández Almagro 3, 28029 Madrid, Spain. Ϯ Current address: Children's Hospital, Stem Cell Program, 300 Longwood Ave, Boston MA 02115, USA. † Current address: The Francis Crick Institute, 1 Midland Road, London NW1 1AT, UK. * Corresponding author ([email protected]) 1 bioRxiv preprint doi: https://doi.org/10.1101/451922; this version posted October 24, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Menchero et al. Abstract The Notch signalling pathway plays fundamental roles in diverse developmental processes in metazoans, where it is important in driving cell fate and directing differentiation of various cell types. -
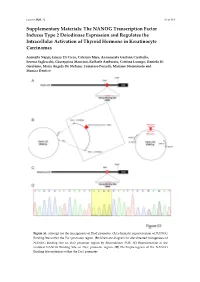
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7