Rab2 Antibody [Discontinued, View Alternatives] (R31430)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Note Hominoid Fission of Chromosome 14/15 and Role Of
Supplemental Note Supplemental Note Hominoid fission of chromosome 14/15 and role of segmental duplications Giuliana Giannuzzi, Michele Pazienza, John Huddleston, Francesca Antonacci, Maika Malig, Laura Vives, Evan E. Eichler and Mario Ventura 1. Analysis of the macaque contig spanning the hominoid 14/15 fission site We grouped contig clones based on their FISH pattern on human and macaque chromosomes (Groups F1, F2, F3, G1, G2, G3, and G4). Group F2 clones (Hsa15b- and Hsa15c-positive) showed a single signal on macaque 7q and three signal clusters on human chromosome 15: at the orthologous 15q26, as expected, as well as at 15q11–14 and 15q24–25, which correspond to the actual and ancestral pericentromeric regions, respectively (Ventura et al. 2003). Indeed, this locus contains an LCR15 copy (Pujana et al. 2001) in both the macaque and human genomes. Most BAC clones were one-end anchored in the human genome (chr15:100,028–100,071 kb). In macaque this region experienced a 64 kb duplicative insertion from chromosome 17 (orthologous to human chromosome 13) (Figure 2), with the human configuration (absence of insertion) likely being the ancestral state because it is identical in orangutan and marmoset. Four Hsa15b- positive clones mapped on macaque and human chromosome 19, but neither human nor macaque assemblies report the STS Hsa15b duplicated at this locus. The presence of assembly gaps in macaque may explain why the STS is not annotated in this region. We aligned the sequence of macaque CH250-70H12 (AC187495.2) versus its human orthologous sequence (hg18 chr15:100,039k-100,170k) and found a 12 kb human expansion through tandem duplication of a ~100 bp unit corresponding to a portion of exon 20 of DNM1 (Figure S3). -

Differential Expression Profiling of Gene Response to Ionizing Radiation in Two Endometrial Cancer Cell Lines with Distinct Radiosensitivities
625-634 28/1/2009 12:32 ÌÌ ™ÂÏ›‰·625 ONCOLOGY REPORTS 21: 625-634, 2009 625 Differential expression profiling of gene response to ionizing radiation in two endometrial cancer cell lines with distinct radiosensitivities XUE-LIAN DU1,2, TAO JIANG2, ZE-QING WEN1, QING-SHUI LI2, RONG GAO2 and FEI WANG1 1Department of Gynecologic Oncology, Shandong Tumor Hospital, Shandong University, Jinan 250117; 2Department of Obstetrics and Gynecology, Provincial Hospital Affiliated to Shandong University, Jinan 250021, P.R. China Received November 4, 2008; Accepted December 17, 2008 DOI: 10.3892/or_00000265 Abstracts. Although radiotherapy is routinely administered Introduction to high-risk endometrial carcinoma and offer a significant disease-free survival advantage, the therapeutic effect is Endometrial cancer is one of the most common gynecological sometimes limited by the occurrence of radioresistance. To malignancies worldwide. Surgery is the preferred initial determine the patterns of gene expression responsible for treatment and most women with early-stage, low-risk disease the radioresistance and to search for potential target genes for will do well without adjuvant radiotherapy. However, both radiotherapy, we selected two cell lines with distinct radio- intermediate-risk and high-risk patients are at risk for local- sensitivities using colony-formation assay from four endo- regional relapse and therefore adjuvant radiotherapy, such as metrial cancer cell lines. The cell cycle distribution showed pelvic radiation, vaginal brachytherapy, and whole-abdomen higher fractions of G2/M phase cells in the radiosensitive cell radiation, is essential for local control (1). Johnson and line KLE after radiation compared with the radioresistant cell colleagues reported that adjuvant external-beam pelvic radio- line ISK. -

Protein Interaction Network of Alternatively Spliced Isoforms from Brain Links Genetic Risk Factors for Autism
ARTICLE Received 24 Aug 2013 | Accepted 14 Mar 2014 | Published 11 Apr 2014 DOI: 10.1038/ncomms4650 OPEN Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism Roser Corominas1,*, Xinping Yang2,3,*, Guan Ning Lin1,*, Shuli Kang1,*, Yun Shen2,3, Lila Ghamsari2,3,w, Martin Broly2,3, Maria Rodriguez2,3, Stanley Tam2,3, Shelly A. Trigg2,3,w, Changyu Fan2,3, Song Yi2,3, Murat Tasan4, Irma Lemmens5, Xingyan Kuang6, Nan Zhao6, Dheeraj Malhotra7, Jacob J. Michaelson7,w, Vladimir Vacic8, Michael A. Calderwood2,3, Frederick P. Roth2,3,4, Jan Tavernier5, Steve Horvath9, Kourosh Salehi-Ashtiani2,3,w, Dmitry Korkin6, Jonathan Sebat7, David E. Hill2,3, Tong Hao2,3, Marc Vidal2,3 & Lilia M. Iakoucheva1 Increased risk for autism spectrum disorders (ASD) is attributed to hundreds of genetic loci. The convergence of ASD variants have been investigated using various approaches, including protein interactions extracted from the published literature. However, these datasets are frequently incomplete, carry biases and are limited to interactions of a single splicing isoform, which may not be expressed in the disease-relevant tissue. Here we introduce a new interactome mapping approach by experimentally identifying interactions between brain-expressed alternatively spliced variants of ASD risk factors. The Autism Spliceform Interaction Network reveals that almost half of the detected interactions and about 30% of the newly identified interacting partners represent contribution from splicing variants, emphasizing the importance of isoform networks. Isoform interactions greatly contribute to establishing direct physical connections between proteins from the de novo autism CNVs. Our findings demonstrate the critical role of spliceform networks for translating genetic knowledge into a better understanding of human diseases. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supporting Information
Supporting Information Edgar et al. 10.1073/pnas.1601895113 SI Methods (Actimetrics), and recordings were analyzed using LumiCycle Mice. Sample size was determined using the resource equation: Data Analysis software (Actimetrics). E (degrees of freedom in ANOVA) = (total number of exper- – Cell Cycle Analysis of Confluent Cell Monolayers. NIH 3T3, primary imental animals) (number of experimental groups), with −/− sample size adhering to the condition 10 < E < 20. For com- WT, and Bmal1 fibroblasts were sequentially transduced − − parison of MuHV-4 and HSV-1 infection in WT vs. Bmal1 / with lentiviral fluorescent ubiquitin-based cell cycle indicators mice at ZT7 (Fig. 2), the investigator did not know the genotype (FUCCI) mCherry::Cdt1 and amCyan::Geminin reporters (32). of the animals when conducting infections, bioluminescence Dual reporter-positive cells were selected by FACS (Influx Cell imaging, and quantification. For bioluminescence imaging, Sorter; BD Biosciences) and seeded onto 35-mm dishes for mice were injected intraperitoneally with endotoxin-free lucif- subsequent analysis. To confirm that expression of mCherry:: Cdt1 and amCyan::Geminin correspond to G1 (2n DNA con- erin (Promega E6552) using 2 mg total per mouse. Following < ≤ anesthesia with isofluorane, they were scanned with an IVIS tent) and S/G2 (2 n 4 DNA content) cell cycle phases, Lumina (Caliper Life Sciences), 15 min after luciferin admin- respectively, cells were stained with DNA dye DRAQ5 (abcam) and analyzed by flow cytometry (LSR-Fortessa; BD Biosci- istration. Signal intensity was quantified using Living Image ences). To examine dynamics of replicative activity under ex- software (Caliper Life Sciences), obtaining maximum radiance perimental confluent conditions, synchronized FUCCI reporter for designated regions of interest (photons per second per − − − monolayers were observed by time-lapse live cell imaging over square centimeter per Steradian: photons·s 1·cm 2·sr 1), relative 3 d (Nikon Eclipse Ti-E inverted epifluorescent microscope). -

GOLGA2/GM130 Is a Novel Target for Neuroprotection Therapy in Intracerebral Hemorrhage
GOLGA2/GM130 is a Novel Target for Neuroprotection Therapy in Intracerebral Hemorrhage Shuwen Deng Second Xiangya Hospital Qing Hu Second Xiangya Hospital Qiang He Second Xiangya Hospital Xiqian Chen Second Xiangya Hospital Wei Lu ( [email protected] ) Second Xiangya Hospital, central south university https://orcid.org/0000-0002-3760-1550 Research Article Keywords: Golgi apparatus, therapy, intracerebral hemorrhage, autophagy, blood–brain barrier Posted Date: June 1st, 2021 DOI: https://doi.org/10.21203/rs.3.rs-547422/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/25 Abstract Blood–brain barrier (BBB) impairment after intracerebral hemorrhage (ICH) can lead to secondary brain injury and aggravate neurological decits. Currently, there are no effective methods for its prevention or treatment partly because of to our lack of understanding of the mechanism of ICH injury to the BBB. Here, we explored the role of Golgi apparatus protein GM130 in the BBB and neurological function after ICH. The levels of the tight junction-associated proteins ZO-1 and occludin decreased, whereas those of LC3-II, an autophagosome marker, increased in hemin-treated Bend.3 cells (p < 0.05). Additionally, GM130 overexpression increased ZO-1 and occludin levels, while decreasing LC3-II levels (p < 0.05). GM130 silencing reversed these effects and mimicked the effect of hemin treatment (p < 0.05). Moreover, tight junctions were disrupted after hemin treatment or GM130 silencing and repaired by GM130 overexpression. GM130 silencing in Bend.3 cells increased autophagic ux, whereas GM130 overexpression downregulated this activity. Furthermore, GM130 silencing-induced tight junction disruption was partially restored by 3-methyladenine (an autophagy inhibitor) administration. -

Roles of TBC1D1 and TBC1D4 in Insulin- and Exercise-Stimulated Glucose Transport of Skeletal Muscle
Diabetologia (2015) 58:19–30 DOI 10.1007/s00125-014-3395-5 REVIEW Roles of TBC1D1 and TBC1D4 in insulin- and exercise-stimulated glucose transport of skeletal muscle Gregory D. Cartee Received: 30 June 2014 /Accepted: 7 August 2014 /Published online: 4 October 2014 # Springer-Verlag Berlin Heidelberg 2014 Abstract This review focuses on two paralogue Rab GTPase mechanism for greater TBC1D4 phosphorylation in insulin- activating proteins known as TBC1D1 Tre-2/BUB2/cdc 1 stimulated muscles after acute exercise is uncertain, and a domain family (TBC1D) 1 and TBC1D4 (also called Akt causal link between enhanced TBC1D4 phosphorylation and Substrate of 160 kDa, AS160) and their roles in controlling increased post-exercise insulin sensitivity has yet to be skeletal muscle glucose transport in response to the indepen- established. In summary, TBC1D1 and TBC1D4 have impor- dent and combined effects of insulin and exercise. Convincing tant, but distinct roles in regulating muscle glucose transport evidence implicates Akt2-dependent TBC1D4 phosphoryla- in response to insulin and exercise. tion on T642 as a key part of the mechanism for insulin- stimulated glucose uptake by skeletal muscle. TBC1D1 phos- Keywords Akt substrate of 160 kDa . Diabetes . Glucose phorylation on several insulin-responsive sites (including transport .High-fatdiet .Insulinresistance .Obesity .Physical T596, a site corresponding to T642 in TBC1D4) does not activity . Review appear to be essential for in vivo insulin-stimulated glucose uptake by skeletal muscle. In vivo exercise or ex vivo con- Abbreviations traction of muscle result in greater TBC1D1 phosphorylation AMPK 5' AMP-activated kinase on S237 that is likely to be secondary to increased AMP- APPL2 Adaptor protein containing PH domain, activated protein kinase activity and potentially important for PTB domain and leucine zipper motif 2 contraction-stimulated glucose uptake. -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

Nuclear PTEN Safeguards Pre-Mrna Splicing to Link Golgi Apparatus for Its Tumor Suppressive Role
ARTICLE DOI: 10.1038/s41467-018-04760-1 OPEN Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role Shao-Ming Shen1, Yan Ji2, Cheng Zhang1, Shuang-Shu Dong2, Shuo Yang1, Zhong Xiong1, Meng-Kai Ge1, Yun Yu1, Li Xia1, Meng Guo1, Jin-Ke Cheng3, Jun-Ling Liu1,3, Jian-Xiu Yu1,3 & Guo-Qiang Chen1 Dysregulation of pre-mRNA alternative splicing (AS) is closely associated with cancers. However, the relationships between the AS and classic oncogenes/tumor suppressors are 1234567890():,; largely unknown. Here we show that the deletion of tumor suppressor PTEN alters pre-mRNA splicing in a phosphatase-independent manner, and identify 262 PTEN-regulated AS events in 293T cells by RNA sequencing, which are associated with significant worse outcome of cancer patients. Based on these findings, we report that nuclear PTEN interacts with the splicing machinery, spliceosome, to regulate its assembly and pre-mRNA splicing. We also identify a new exon 2b in GOLGA2 transcript and the exon exclusion contributes to PTEN knockdown-induced tumorigenesis by promoting dramatic Golgi extension and secretion, and PTEN depletion significantly sensitizes cancer cells to secretion inhibitors brefeldin A and golgicide A. Our results suggest that Golgi secretion inhibitors alone or in combination with PI3K/Akt kinase inhibitors may be therapeutically useful for PTEN-deficient cancers. 1 Department of Pathophysiology, Key Laboratory of Cell Differentiation and Apoptosis of Chinese Ministry of Education, Shanghai Jiao Tong University School of Medicine (SJTU-SM), Shanghai 200025, China. 2 Institute of Health Sciences, Shanghai Institutes for Biological Sciences of Chinese Academy of Sciences and SJTU-SM, Shanghai 200025, China. -
Download Validation Data
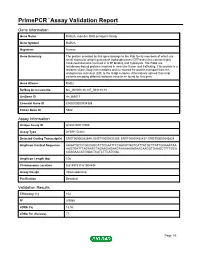
PrimePCR™Assay Validation Report Gene Information Gene Name RAB2A, member RAS oncogene family Gene Symbol RAB2A Organism Human Gene Summary The protein encoded by this gene belongs to the Rab family members of which are small molecular weight guanosine triphosphatases (GTPases) that contain highly conserved domains involved in GTP binding and hydrolysis. The Rabs are membrane-bound proteins involved in vesicular fusion and trafficking. This protein is a resident of pre-Golgi intermediates and is required for protein transport from the endoplasmic reticulum (ER) to the Golgi complex. Alternatively spliced transcript variants encoding different isoforms have been found for this gene. Gene Aliases RAB2 RefSeq Accession No. NC_000008.10, NT_008183.19 UniGene ID Hs.369017 Ensembl Gene ID ENSG00000104388 Entrez Gene ID 5862 Assay Information Unique Assay ID qHsaCID0011980 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000262646, ENST00000531289, ENST00000452437, ENST00000543829 Amplicon Context Sequence AAGATGCCCGCCAGCATTCCAATTCCAACATGGTCATTATGCTTATTGGAAATAA AAGTGATTTAGAATCTAGAAGAGAAGTAAAAAAAGAAGAAGGTGAAGCTTTTGCA CGAGAACATGGACTCATCTTCATGGA Amplicon Length (bp) 106 Chromosome Location 8:61497317-61504494 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 104 R2 0.9988 cDNA Cq 18.53 cDNA Tm (Celsius) 77 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 43.2 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report -

E-Mutpath: Computational Modelling Reveals the Functional Landscape of Genetic Mutations Rewiring Interactome Networks
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.22.262386; this version posted August 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. e-MutPath: Computational modelling reveals the functional landscape of genetic mutations rewiring interactome networks Yongsheng Li1, Daniel J. McGrail1, Brandon Burgman2,3, S. Stephen Yi2,3,4,5 and Nidhi Sahni1,6,7,8,* 1Department oF Systems Biology, The University oF Texas MD Anderson Cancer Center, Houston, TX 77030, USA 2Department oF Oncology, Livestrong Cancer Institutes, Dell Medical School, The University oF Texas at Austin, Austin, TX 78712, USA 3Institute For Cellular and Molecular Biology (ICMB), The University oF Texas at Austin, Austin, TX 78712, USA 4Institute For Computational Engineering and Sciences (ICES), The University oF Texas at Austin, Austin, TX 78712, USA 5Department oF Biomedical Engineering, Cockrell School of Engineering, The University oF Texas at Austin, Austin, TX 78712, USA 6Department oF Epigenetics and Molecular Carcinogenesis, The University oF Texas MD Anderson Science Park, Smithville, TX 78957, USA 7Department oF BioinFormatics and Computational Biology, The University oF Texas MD Anderson Cancer Center, Houston, TX 77030, USA 8Program in Quantitative and Computational Biosciences (QCB), Baylor College oF Medicine, Houston, TX 77030, USA *To whom correspondence should be addressed. Nidhi Sahni. Tel: +1 512 2379506; Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.08.22.262386; this version posted August 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

A SARS-Cov-2-Human Protein-Protein Interaction Map Reveals Drug Targets and Potential Drug- Repurposing
bioRxiv preprint doi: https://doi.org/10.1101/2020.03.22.002386; this version posted March 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. A SARS-CoV-2-Human Protein-Protein Interaction Map Reveals Drug Targets and Potential Drug- Repurposing David E. Gordon1,2,3,4, Gwendolyn M. Jang1,2,3,4, Mehdi Bouhaddou1,2,3,4, Jiewei Xu1,2,3,4, Kirsten Obernier1,2,3,4, Matthew J. O'Meara5, Jeffrey Z. Guo1,2,3,4, Danielle L. Swaney1,2,3,4, Tia A. Tummino1,2,6, Ruth Huettenhain1,2,3,4, Robyn Kaake1,2,3,4, Alicia L. Richards1,2,3,4, Beril Tutuncuoglu1,2,3,4, Helene Foussard1,2,3,4, Jyoti Batra1,2,3,4, Kelsey Haas1,2,3,4, Maya Modak1,2,3,4, Minkyu Kim1,2,3,4, Paige Haas1,2,3,4, Benjamin J. Polacco1,2,3,4, Hannes Braberg1,2,3,4, Jacqueline M. Fabius1,2,3,4, Manon Eckhardt1,2,3,4, Margaret Soucheray1,2,3,4, Melanie J. Bennett1,2,3,4, Merve Cakir1,2,3,4, Michael J. McGregor1,2,3,4, Qiongyu Li1,2,3,4, Zun Zar Chi Naing1,2,3,4, Yuan Zhou1,2,3,4, Shiming Peng1,2,6, Ilsa T. Kirby1,4,7, James E. Melnyk1,4,7, John S. Chorba1,4,7, Kevin Lou1,4,7, ShiZhong A. Dai1,4,7, Wenqi Shen1,4,7, Ying Shi1,4,7, Ziyang Zhang1,4,7, Inigo Barrio-HernandeZ8, Danish Memon8, Claudia Hernandez-Armenta8, Christopher J.P.