Proteomics-Based Comparative Mapping of the Human Brown and White Adipocyte Secretome
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptomic Analysis of the Aquaporin (AQP) Gene Family
Pancreatology 19 (2019) 436e442 Contents lists available at ScienceDirect Pancreatology journal homepage: www.elsevier.com/locate/pan Transcriptomic analysis of the Aquaporin (AQP) gene family interactome identifies a molecular panel of four prognostic markers in patients with pancreatic ductal adenocarcinoma Dimitrios E. Magouliotis a, b, Vasiliki S. Tasiopoulou c, Konstantinos Dimas d, * Nikos Sakellaridis d, Konstantina A. Svokos e, Alexis A. Svokos f, Dimitris Zacharoulis b, a Division of Surgery and Interventional Science, Faculty of Medical Sciences, UCL, London, UK b Department of Surgery, University of Thessaly, Biopolis, Larissa, Greece c Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece d Department of Pharmacology, Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece e The Warren Alpert Medical School of Brown University, Providence, RI, USA f Riverside Regional Medical Center, Newport News, VA, USA article info abstract Article history: Background: This study aimed to assess the differential gene expression of aquaporin (AQP) gene family Received 14 October 2018 interactome in pancreatic ductal adenocarcinoma (PDAC) using data mining techniques to identify novel Received in revised form candidate genes intervening in the pathogenicity of PDAC. 29 January 2019 Method: Transcriptome data mining techniques were used in order to construct the interactome of the Accepted 9 February 2019 AQP gene family and to determine which genes members are differentially expressed in PDAC as Available online 11 February 2019 compared to controls. The same techniques were used in order to evaluate the potential prognostic role of the differentially expressed genes. Keywords: PDAC Results: Transcriptome microarray data of four GEO datasets were incorporated, including 142 primary Aquaporin tumor samples and 104 normal pancreatic tissue samples. -

Supplementary Materials
1 Supplementary Materials: Supplemental Figure 1. Gene expression profiles of kidneys in the Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice. (A) A heat map of microarray data show the genes that significantly changed up to 2 fold compared between Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice (N=4 mice per group; p<0.05). Data show in log2 (sample/wild-type). 2 Supplemental Figure 2. Sting signaling is essential for immuno-phenotypes of the Fcgr2b-/-lupus mice. (A-C) Flow cytometry analysis of splenocytes isolated from wild-type, Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice at the age of 6-7 months (N= 13-14 per group). Data shown in the percentage of (A) CD4+ ICOS+ cells, (B) B220+ I-Ab+ cells and (C) CD138+ cells. Data show as mean ± SEM (*p < 0.05, **p<0.01 and ***p<0.001). 3 Supplemental Figure 3. Phenotypes of Sting activated dendritic cells. (A) Representative of western blot analysis from immunoprecipitation with Sting of Fcgr2b-/- mice (N= 4). The band was shown in STING protein of activated BMDC with DMXAA at 0, 3 and 6 hr. and phosphorylation of STING at Ser357. (B) Mass spectra of phosphorylation of STING at Ser357 of activated BMDC from Fcgr2b-/- mice after stimulated with DMXAA for 3 hour and followed by immunoprecipitation with STING. (C) Sting-activated BMDC were co-cultured with LYN inhibitor PP2 and analyzed by flow cytometry, which showed the mean fluorescence intensity (MFI) of IAb expressing DC (N = 3 mice per group). 4 Supplemental Table 1. Lists of up and down of regulated proteins Accession No. -
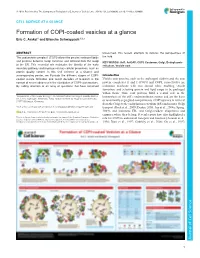
Formation of COPI-Coated Vesicles at a Glance Eric C
© 2018. Published by The Company of Biologists Ltd | Journal of Cell Science (2018) 131, jcs209890. doi:10.1242/jcs.209890 CELL SCIENCE AT A GLANCE Formation of COPI-coated vesicles at a glance Eric C. Arakel1 and Blanche Schwappach1,2,* ABSTRACT unresolved, this review attempts to refocus the perspectives of The coat protein complex I (COPI) allows the precise sorting of lipids the field. and proteins between Golgi cisternae and retrieval from the Golgi KEY WORDS: Arf1, ArfGAP, COPI, Coatomer, Golgi, Endoplasmic to the ER. This essential role maintains the identity of the early reticulum, Vesicle coat secretory pathway and impinges on key cellular processes, such as protein quality control. In this Cell Science at a Glance and accompanying poster, we illustrate the different stages of COPI- Introduction coated vesicle formation and revisit decades of research in the Vesicle coat proteins, such as the archetypal clathrin and the coat context of recent advances in the elucidation of COPI coat structure. protein complexes II and I (COPII and COPI, respectively) are By calling attention to an array of questions that have remained molecular machines with two central roles: enabling vesicle formation, and selecting protein and lipid cargo to be packaged within them. Thus, coat proteins fulfil a central role in the 1Department of Molecular Biology, Universitätsmedizin Göttingen, Humboldtallee homeostasis of the cell’s endomembrane system and are the basis 23, 37073 Göttingen, Germany. 2Max-Planck Institute for Biophysical Chemistry, 37077 Göttingen, Germany. of functionally segregated compartments. COPI operates in retrieval from the Golgi to the endoplasmic reticulum (ER) and in intra-Golgi *Author for correspondence ([email protected]) transport (Beck et al., 2009; Duden, 2003; Lee et al., 2004a; Spang, E.C.A., 0000-0001-7716-7149; B.S., 0000-0003-0225-6432 2009), and maintains ER- and Golgi-resident chaperones and enzymes where they belong. -

Supplementary Figures 1-14 and Supplementary References
SUPPORTING INFORMATION Spatial Cross-Talk Between Oxidative Stress and DNA Replication in Human Fibroblasts Marko Radulovic,1,2 Noor O Baqader,1 Kai Stoeber,3† and Jasminka Godovac-Zimmermann1* 1Division of Medicine, University College London, Center for Nephrology, Royal Free Campus, Rowland Hill Street, London, NW3 2PF, UK. 2Insitute of Oncology and Radiology, Pasterova 14, 11000 Belgrade, Serbia 3Research Department of Pathology and UCL Cancer Institute, Rockefeller Building, University College London, University Street, London WC1E 6JJ, UK †Present Address: Shionogi Europe, 33 Kingsway, Holborn, London WC2B 6UF, UK TABLE OF CONTENTS 1. Supplementary Figures 1-14 and Supplementary References. Figure S-1. Network and joint spatial razor plot for 18 enzymes of glycolysis and the pentose phosphate shunt. Figure S-2. Correlation of SILAC ratios between OXS and OAC for proteins assigned to the SAME class. Figure S-3. Overlap matrix (r = 1) for groups of CORUM complexes containing 19 proteins of the 49-set. Figure S-4. Joint spatial razor plots for the Nop56p complex and FIB-associated complex involved in ribosome biogenesis. Figure S-5. Analysis of the response of emerin nuclear envelope complexes to OXS and OAC. Figure S-6. Joint spatial razor plots for the CCT protein folding complex, ATP synthase and V-Type ATPase. Figure S-7. Joint spatial razor plots showing changes in subcellular abundance and compartmental distribution for proteins annotated by GO to nucleocytoplasmic transport (GO:0006913). Figure S-8. Joint spatial razor plots showing changes in subcellular abundance and compartmental distribution for proteins annotated to endocytosis (GO:0006897). Figure S-9. Joint spatial razor plots for 401-set proteins annotated by GO to small GTPase mediated signal transduction (GO:0007264) and/or GTPase activity (GO:0003924). -

Cogps:Cancer Outlier Gene Profile Sets
coGPS:Cancer Outlier Gene Profile Sets Yingying Wei, Michael Ochs April 24, 2017 1 Introduction Generation of high-throughput genomic data has become routine in modern bi- ology. While the focus remains in many cases on the identification of molecular species, such as mRNA transcripts, that differ between two conditions, the data is often of great interest for elucidating differences in pathway activity. Standard algorithms, such as limma [Smyth 2004]and SAM [Tusher , Tibshirani and Chu 2001], are widely used for identifying transcripts (hereon referred to genes) that dif- fer statistically between two groups. Later, as multiple datasets became avail- able on the same biological process, people began to combine information from multiple studies to improve the power of statistical analysis in each study [Conlon, Song and Liu 2006], [Kendziorski et al. 2003], [Yuan and Kendziorski 2006], [Cui et al. 2005], and [Scharpf et al. 2009]. It is often the case in cancer studies that the identification of important genes by these approaches fails. The realization that genes are often disregulated in only a subset of tumors led to the development of Cancer Outlier Profile Analysis (COPA) [MacDonald, 2006]. Outlier genes are defined to be ones that are over- or under- expressed in a subset of tumor samples compared to normal samples. The difference between outlier differential expressed gene and normal differen- tial expressed gene is that outlier gene may have the same level of expression in majority of tumor samples as that of normals. A few statistics for finding outlier genes have been proposed [MacDonald, 2006], [Tibshirani, R. and Hastie, 2007], and [Wu, 2007]. -

In Vivo Mapping of a GPCR Interactome Using Knockin Mice
In vivo mapping of a GPCR interactome using knockin mice Jade Degrandmaisona,b,c,d,e,1, Khaled Abdallahb,c,d,1, Véronique Blaisb,c,d, Samuel Géniera,c,d, Marie-Pier Lalumièrea,c,d, Francis Bergeronb,c,d,e, Catherine M. Cahillf,g,h, Jim Boulterf,g,h, Christine L. Lavoieb,c,d,i, Jean-Luc Parenta,c,d,i,2, and Louis Gendronb,c,d,i,j,k,2 aDépartement de Médecine, Université de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; bDépartement de Pharmacologie–Physiologie, Université de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; cFaculté de Médecine et des Sciences de la Santé, Université de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; dCentre de Recherche du Centre Hospitalier Universitaire de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; eQuebec Network of Junior Pain Investigators, Sherbrooke, QC J1H 5N4, Canada; fDepartment of Psychiatry and Biobehavioral Sciences, University of California, Los Angeles, CA 90095; gSemel Institute for Neuroscience and Human Behavior, University of California, Los Angeles, CA 90095; hShirley and Stefan Hatos Center for Neuropharmacology, University of California, Los Angeles, CA 90095; iInstitut de Pharmacologie de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; jDépartement d’Anesthésiologie, Université de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; and kQuebec Pain Research Network, Sherbrooke, QC J1H 5N4, Canada Edited by Brian K. Kobilka, Stanford University School of Medicine, Stanford, CA, and approved April 9, 2020 (received for review October 16, 2019) With over 30% of current medications targeting this family of attenuates pain hypersensitivities in several chronic pain models proteins, G-protein–coupled receptors (GPCRs) remain invaluable including neuropathic, inflammatory, diabetic, and cancer pain therapeutic targets. -

A Trafficome-Wide Rnai Screen Reveals Deployment of Early and Late Secretory Host Proteins and the Entire Late Endo-/Lysosomal V
bioRxiv preprint doi: https://doi.org/10.1101/848549; this version posted November 19, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 A trafficome-wide RNAi screen reveals deployment of early and late 2 secretory host proteins and the entire late endo-/lysosomal vesicle fusion 3 machinery by intracellular Salmonella 4 5 Alexander Kehl1,4, Vera Göser1, Tatjana Reuter1, Viktoria Liss1, Maximilian Franke1, 6 Christopher John1, Christian P. Richter2, Jörg Deiwick1 and Michael Hensel1, 7 8 1Division of Microbiology, University of Osnabrück, Osnabrück, Germany; 2Division of Biophysics, University 9 of Osnabrück, Osnabrück, Germany, 3CellNanOs – Center for Cellular Nanoanalytics, Fachbereich 10 Biologie/Chemie, Universität Osnabrück, Osnabrück, Germany; 4current address: Institute for Hygiene, 11 University of Münster, Münster, Germany 12 13 Running title: Host factors for SIF formation 14 Keywords: siRNA knockdown, live cell imaging, Salmonella-containing vacuole, Salmonella- 15 induced filaments 16 17 Address for correspondence: 18 Alexander Kehl 19 Institute for Hygiene 20 University of Münster 21 Robert-Koch-Str. 4148149 Münster, Germany 22 Tel.: +49(0)251/83-55233 23 E-mail: [email protected] 24 25 or bioRxiv preprint doi: https://doi.org/10.1101/848549; this version posted November 19, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. -

Development of an Infectious Clone System to Study the Life Cycle of Hazara Virus
Development of an Infectious Clone System to Study the Life Cycle of Hazara Virus Jack Fuller Submitted in accordance with the requirements for the degree of Doctor of Philosophy The University of Leeds Faculty of Biological Sciences July 2020 i The candidate confirms that the work submitted is his own and that appropriate credit has been given where reference has been made to the work of others. This copy has been supplied on the understanding that it is copyright material and that no quotation from the thesis may be published without proper acknowledgement. © 2020 The University of Leeds and Jack Fuller ii Acknowledgements I would like to thank my supervisors, John, Jamel and Roger for their endless support and fresh ideas throughout my PhD. A special thanks go to John for always providing enthusiasm and encouragement, especially during the tougher times of the project! I would also like to thank everyone in 8.61, not only for making my time at Leeds enjoyable, but for helping me develop as a scientist through useful critique and discussion. A special thanks goes to Francis Hopkins, for welcoming me into the Barr group and providing a constant supply of humour and to Ellie Todd, for listening to my endless whines and gripes, and for providing welcome distractions in the form of her PowerPoint artwork. Outside of the lab, my mum deserves a special mention for always being the voice of reason and support at the end of the phone, but also for pushing me to succeed throughout my entire education. I have no doubts I would not be in the position I am now without her! Finally, a huge thanks to my partner Hannah, who has been there to support me through all the highs and lows of my PhD, and has sacrificed many weekend trips to allow me to finish experiments and to tend to my viruses! iii Abstract Crimean-Congo hemorrhagic fever orthonairovirus (CCHFV) is a negative sense single stranded RNA virus, capable of causing fatal hemorrhagic fever in humans. -

Membrane Trafficking in Health and Disease Rebecca Yarwood*, John Hellicar*, Philip G
© 2020. Published by The Company of Biologists Ltd | Disease Models & Mechanisms (2020) 13, dmm043448. doi:10.1242/dmm.043448 AT A GLANCE Membrane trafficking in health and disease Rebecca Yarwood*, John Hellicar*, Philip G. Woodman‡ and Martin Lowe‡ ABSTRACT KEY WORDS: Disease, Endocytic pathway, Genetic disorder, Membrane traffic, Secretory pathway, Vesicle Membrane trafficking pathways are essential for the viability and growth of cells, and play a major role in the interaction of cells with Introduction their environment. In this At a Glance article and accompanying Membrane trafficking pathways are essential for cells to maintain poster, we outline the major cellular trafficking pathways and discuss critical functions, to grow, and to accommodate to their chemical how defects in the function of the molecular machinery that mediates and physical environment. Membrane flux through these pathways this transport lead to various diseases in humans. We also briefly is high, and in specialised cells in some tissues can be enormous. discuss possible therapeutic approaches that may be used in the For example, pancreatic acinar cells synthesise and secrete amylase, future treatment of trafficking-based disorders. one of the many enzymes they produce, at a rate of approximately 0.5% of cellular protein mass per hour (Allfrey et al., 1953), while in Schwann cells, the rate of membrane protein export must correlate School of Biological Sciences, Faculty of Biology, Medicine and Health, with the several thousand-fold expansion of the cell surface that University of Manchester, Manchester, M13 9PT, UK. occurs during myelination (Pereira et al., 2012). The population of *These authors contributed equally to this work cell surface proteins is constantly monitored and modified via the ‡Authors for correspondence ([email protected]; endocytic pathway. -

Research Article Complex and Multidimensional Lipid Raft Alterations in a Murine Model of Alzheimer’S Disease
SAGE-Hindawi Access to Research International Journal of Alzheimer’s Disease Volume 2010, Article ID 604792, 56 pages doi:10.4061/2010/604792 Research Article Complex and Multidimensional Lipid Raft Alterations in a Murine Model of Alzheimer’s Disease Wayne Chadwick, 1 Randall Brenneman,1, 2 Bronwen Martin,3 and Stuart Maudsley1 1 Receptor Pharmacology Unit, National Institute on Aging, National Institutes of Health, 251 Bayview Boulevard, Suite 100, Baltimore, MD 21224, USA 2 Miller School of Medicine, University of Miami, Miami, FL 33124, USA 3 Metabolism Unit, National Institute on Aging, National Institutes of Health, 251 Bayview Boulevard, Suite 100, Baltimore, MD 21224, USA Correspondence should be addressed to Stuart Maudsley, [email protected] Received 17 May 2010; Accepted 27 July 2010 Academic Editor: Gemma Casadesus Copyright © 2010 Wayne Chadwick et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Various animal models of Alzheimer’s disease (AD) have been created to assist our appreciation of AD pathophysiology, as well as aid development of novel therapeutic strategies. Despite the discovery of mutated proteins that predict the development of AD, there are likely to be many other proteins also involved in this disorder. Complex physiological processes are mediated by coherent interactions of clusters of functionally related proteins. Synaptic dysfunction is one of the hallmarks of AD. Synaptic proteins are organized into multiprotein complexes in high-density membrane structures, known as lipid rafts. These microdomains enable coherent clustering of synergistic signaling proteins. -

Mutations in COPA Lead to Abnormal Trafficking of STING to the Golgi and Interferon Signaling
Mutations in COPA lead to abnormal trafficking of STING to the Golgi and interferon signaling Alice Lepelley, Maria José Martin-Niclos, Melvin Le Bihan, Joseph Marsh, Carolina Uggenti, Gillian Rice, Vincent Bondet, Darragh Duffy, Jonny Hertzog, Jan Rehwinkel, et al. To cite this version: Alice Lepelley, Maria José Martin-Niclos, Melvin Le Bihan, Joseph Marsh, Carolina Uggenti, et al.. Mutations in COPA lead to abnormal trafficking of STING to the Golgi and interferon signal- ing. Journal of Experimental Medicine, Rockefeller University Press, 2020, 217 (11), pp.e20200600. 10.1084/jem.20200600. pasteur-02933253 HAL Id: pasteur-02933253 https://hal-pasteur.archives-ouvertes.fr/pasteur-02933253 Submitted on 8 Sep 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. BRIEF DEFINITIVE REPORT Mutations in COPA lead to abnormal trafficking of STING to the Golgi and interferon signaling Alice Lepelley1, Maria Jose´ Martin-Niclós1, Melvin Le Bihan2, Joseph A. Marsh3, Carolina Uggenti4, Gillian I. Rice5, Vincent Bondet6,7, Darragh Duffy6,7, Jonny Hertzog8, Jan Rehwinkel8, Serge Amselem9,10, Siham Boulisfane-El Khalifi11,MaryBrennan12, Edwin Carter4, Lucienne Chatenoud13,14,15,Stephanie´ Chhun13,14,15, Aurore Coulomb l’Hermine16, Marine Depp4, Marie Legendre9,10, Karen J. -
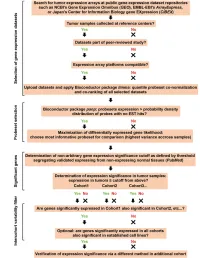
Supplementary Data
SUPPLEMENTAL INFORMATION A study restricted to chemokine receptors as well as a genome-wide transcript analysis uncovered CXCR4 as preferentially expressed in Ewing's sarcoma (Ewing's sarcoma) cells of metastatic origin (Figure 4). Transcriptome analyses showed that in addition to CXCR4, genes known to support cell motility and invasion topped the list of genes preferentially expressed in metastasis-derived cells (Figure 4D). These included kynurenine 3-monooxygenase (KMO), galectin-1 (LGALS1), gastrin-releasing peptide (GRP), procollagen C-endopeptidase enhancer (PCOLCE), and ephrin receptor B (EPHB3). KMO, a key enzyme of tryptophan catabolism, has not been linked to metastasis. Tryptophan and its catabolites, however, are involved in immune evasion by tumors, a process that can assist in tumor progression and metastasis (1). LGALS1, GRP, PCOLCE and EPHB3 have been linked to tumor progression and metastasis of several cancers (2-4). Top genes preferentially expressed in L-EDCL included genes that suppress cell motility and/or potentiate cell adhesion such as plakophilin 1 (PKP1), neuropeptide Y (NPY), or the metastasis suppressor TXNIP (5-7) (Figure 4D). Overall, L-EDCL were enriched in gene sets geared at optimizing nutrient transport and usage (Figure 4D; Supplementary Table 3), a state that may support the early stages of tumor growth. Once tumor growth outpaces nutrient and oxygen supplies, gene expression programs are usually switched to hypoxic response and neoangiogenesis, which ultimately lead to tumor egress and metastasis. Accordingly, gene sets involved in extracellular matrix remodeling, MAPK signaling, and response to hypoxia were up-regulated in M-EDCL (Figure 4D; Supplementary Table 4), consistent with their association to metastasis in other cancers (8, 9).