Three Distinct Developmental Pathways for Adaptive and Two IFN-Γ-Producing Γδ T Subsets in Adult Thymus
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Tables
Supp Table 1. Patient characteristics of samples used for colony assays, xenografts and intracellular colony assays Sr.No Age Sex Diagnosis Blast% Cytogenetics Mutations (IPSS) (AF%) Patient samples used for colony assays 1. 68 F Low risk 1% N/A None MDS 2. 78 M RAEB-1 7% N/A DNMT3A 3. 66 F High risk 12.2% 5q-,7q-,+11, DNMT3A MDS 20q- (28%) TP53 (58%) 4. 61 M High risk 6.2% Monosomy 7 ASXL1 (36%) MDS EZH2 (77%) RUNX1(18%) 5. 63 M Low risk Normal ETV6 (34%), MDS KRAS(15%), RUNX1 (40%), SRSF2 (43%), ZRSR2 (86%) 6. 62 F RAEB-2 Del 5q TP53 (7%) 7. 76 F t-MDS <1% Normal TET2(10%) 8. 80 M Low risk 5% Normal SF3B1(21%) MDS TET2(8%) ZRZR2(62%) 9. 74 F MPN 5% JAK2V617F+ 10. 76 M Low risk <1% Deletion Y None MDS 11. 84 M Low risk N/A N/A N/A MDS 12. 86 M Int-2 MDS 4-8% Normal U2AF1 (43%) blasts CBL (15%) 13. 64 M low risk 1-3% 20q deletion ASXL1 (17%) MDS SETBP1(17%) U2AF1(15%) 14. 81 F Low risk <1% Normal None MDS Patient samples used for PDX 15. 87 F Int-2 risk 1.2% Complex SETBP1 (38%) MDS cytogenetics, del 7, dup 11, del 13q 16. 59 F High-risk 7-10% Complex NRAS (12%), MDS cytogenetics RUNX1 (20%), (-5q31, -7q31, SRSF2 (22%), trisomy 8, del STAG2 (17%) 11q23 17. 79 F Int-2 risk 5-8% None MDS 18. 67 M MPN 6.6% Normal CALR (51%) IDH1(47%) PDGFRB (47%) Patient samples used for intracellular ASO uptake 19. -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -

Early Growth Response 1 Regulates Hematopoietic Support and Proliferation in Human Primary Bone Marrow Stromal Cells
Hematopoiesis SUPPLEMENTARY APPENDIX Early growth response 1 regulates hematopoietic support and proliferation in human primary bone marrow stromal cells Hongzhe Li, 1,2 Hooi-Ching Lim, 1,2 Dimitra Zacharaki, 1,2 Xiaojie Xian, 2,3 Keane J.G. Kenswil, 4 Sandro Bräunig, 1,2 Marc H.G.P. Raaijmakers, 4 Niels-Bjarne Woods, 2,3 Jenny Hansson, 1,2 and Stefan Scheding 1,2,5 1Division of Molecular Hematology, Department of Laboratory Medicine, Lund University, Lund, Sweden; 2Lund Stem Cell Center, Depart - ment of Laboratory Medicine, Lund University, Lund, Sweden; 3Division of Molecular Medicine and Gene Therapy, Department of Labora - tory Medicine, Lund University, Lund, Sweden; 4Department of Hematology, Erasmus MC Cancer Institute, Rotterdam, the Netherlands and 5Department of Hematology, Skåne University Hospital Lund, Skåne, Sweden ©2020 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2019.216648 Received: January 14, 2019. Accepted: July 19, 2019. Pre-published: August 1, 2019. Correspondence: STEFAN SCHEDING - [email protected] Li et al.: Supplemental data 1. Supplemental Materials and Methods BM-MNC isolation Bone marrow mononuclear cells (BM-MNC) from BM aspiration samples were isolated by density gradient centrifugation (LSM 1077 Lymphocyte, PAA, Pasching, Austria) either with or without prior incubation with RosetteSep Human Mesenchymal Stem Cell Enrichment Cocktail (STEMCELL Technologies, Vancouver, Canada) for lineage depletion (CD3, CD14, CD19, CD38, CD66b, glycophorin A). BM-MNCs from fetal long bones and adult hip bones were isolated as reported previously 1 by gently crushing bones (femora, tibiae, fibulae, humeri, radii and ulna) in PBS+0.5% FCS subsequent passing of the cell suspension through a 40-µm filter. -

Novel Gene Signatures Predictive of Patient Recurrence-Free Survival and Castration Resistance in Prostate Cancer Jun A, Baotong Zhang, Zhiqian Zhang, Jin-Tang Dong
Supplementary Materials: Novel Gene Signatures Predictive of Patient Recurrence-Free Survival and Castration Resistance in Prostate Cancer Jun A, Baotong Zhang, Zhiqian Zhang, Jin-Tang Dong Figure S1. Overlap differentially expressed genes (DEGs) between our study and previously re- ported microarray experiments. Figure S2. The diagnosis effectiveness of the six CRPCPS genes. ROC curves for the six CRPCPS genes in the training cohort (a) and the validation cohort (b). Cancers 2021, 13 S2 of S13 Figure S3. Evaluation of the CRPCPS in the internal validation cohort and the entire TCGA cohort. (a) Distribution of CRPCPS score (left), patients’ recurrent status (center), and expression profiles of the six CRPCPS genes (right) in the internal validation cohort. (b) Receiver operating character- istic (ROC) curves were used to evaluate the predictability of RFS at 3-, 5-, and 8-year by the CRPCPS score, Gleason score, and pathological tumor stage in the internal validation cohort. (c) Distribution of CRPCPS score (left), patients' survival status (center), and expression profiles of the six CRPCPS genes that constitute the CRPCPS (right) in the entire TCGA cohort. (d) Receiver oper- ating characteristic (ROC) curves were used to evaluate the predictability of RFS at 3-, 5-, and 8- year by the CRPCPS score, Gleason score, and pathological tumor stage in the entire TCGA cohort. Cancers 2021, 13 S3 of S13 Figure S4. Association of CRPCPS with tumor stage (a-d), Gleason score (e-h), and lymph node status (i-k) in different patient cohorts. Figure S5. Association of CRPCPS with patients’ age in the TCGA training cohort (a), the TCGA validation cohort (b), the entire TCGA cohort (c), the MSKCC cohort (d), the Cambridge cohort (GSE70768) (e), and the Belfast cohort (GSE116918) (f). -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -

LITAF and TNFSF15, Two Downstream Targets of AMPK, Exert Inhibitory Effects on Tumor Growth
Oncogene (2011) 30, 1892–1900 & 2011 Macmillan Publishers Limited All rights reserved 0950-9232/11 www.nature.com/onc ORIGINAL ARTICLE LITAF and TNFSF15, two downstream targets of AMPK, exert inhibitory effects on tumor growth J Zhou1, Z Yang2, T Tsuji3, J Gong1, J Xie1, C Chen3,WLi1, S Amar4 and Z Luo1 1Department of Biochemistry, Boston University School of Medicine, Boston, MA, USA; 2Department of Biochemistry and Molecular Biology, College of Life Sciences, Shaanxi Normal University, Xi’an, PR China; 3Department of Radiation Oncology, Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, MA, USA and 4Department of Periodontology and Oral Biology, Boston University School of Dental Medicine, Boston, MA, USA Lipopolysaccharide (LPS)-induced tumor necrosis factor Keywords: AMPK; LITAF; TNFSF15; p53; tumor (TNF) a factor (LITAF) is a multiple functional molecule suppressor; tumorigenesis whose sequence is identical to the small integral membrane protein of the lysosome/late endosome. LITAF was initially identified as a transcription factor that activates transcription of proinflammatory cytokine in Introduction macrophages in response to LPS. Mutations of the LITAF gene are associated with a genetic disease, called Lipopolysaccharide (LPS)-induced tumor necrosis fac- Charcot–Marie–Tooth syndrome. Recently, we have tor (TNF) a factor (LITAF) was initially characterized reported that mRNA levels of LITAF and TNF super- as a transcription factor that activates transcription of family member 15 (TNFSF15) are upregulated by 50 cytokines such as TNFa, interleukin 6, soluble TNF- adenosine monophosphate (AMP)-activated protein ki- receptor II (sTNF-RII) and chemokine interleukin-8 in nase (AMPK). The present study further assesses their macrophages in response to LPS (Myokai et al., 1999; biological functions. -

A Single-Cell Transcriptome Atlas of the Mouse Glomerulus
RAPID COMMUNICATION www.jasn.org A Single-Cell Transcriptome Atlas of the Mouse Glomerulus Nikos Karaiskos,1 Mahdieh Rahmatollahi,2 Anastasiya Boltengagen,1 Haiyue Liu,1 Martin Hoehne ,2 Markus Rinschen,2,3 Bernhard Schermer,2,4,5 Thomas Benzing,2,4,5 Nikolaus Rajewsky,1 Christine Kocks ,1 Martin Kann,2 and Roman-Ulrich Müller 2,4,5 Due to the number of contributing authors, the affiliations are listed at the end of this article. ABSTRACT Background Three different cell types constitute the glomerular filter: mesangial depending on cell location relative to the cells, endothelial cells, and podocytes. However, to what extent cellular heteroge- glomerular vascular pole.3 Because BP ad- neity exists within healthy glomerular cell populations remains unknown. aptation and mechanoadaptation of glo- merular cells are key determinants of kidney Methods We used nanodroplet-based highly parallel transcriptional profiling to function and dysregulated in kidney disease, characterize the cellular content of purified wild-type mouse glomeruli. we tested whether glomerular cell type sub- Results Unsupervised clustering of nearly 13,000 single-cell transcriptomes identi- sets can be identified by single-cell RNA fied the three known glomerular cell types. We provide a comprehensive online sequencing in wild-type glomeruli. This atlas of gene expression in glomerular cells that can be queried and visualized using technique allows for high-throughput tran- an interactive and freely available database. Novel marker genes for all glomerular scriptome profiling of individual cells and is cell types were identified and supported by immunohistochemistry images particularly suitable for identifying novel obtained from the Human Protein Atlas. -

Overview of Research on Fusion Genes in Prostate Cancer
2011 Review Article Overview of research on fusion genes in prostate cancer Chunjiao Song1,2, Huan Chen3 1Medical Research Center, Shaoxing People’s Hospital, Shaoxing University School of Medicine, Shaoxing 312000, China; 2Shaoxing Hospital, Zhejiang University School of Medicine, Shaoxing 312000, China; 3Key Laboratory of Microorganism Technology and Bioinformatics Research of Zhejiang Province, Zhejiang Institute of Microbiology, Hangzhou 310000, China Contributions: (I) Conception and design: C Song; (II) Administrative support: Shaoxing Municipal Health and Family Planning Science and Technology Innovation Project (2017CX004) and Shaoxing Public Welfare Applied Research Project (2018C30058); (III) Provision of study materials or patients: None; (IV) Collection and assembly of data: C Song; (V) Data analysis and interpretation: H Chen; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. Correspondence to: Chunjiao Song. No. 568 Zhongxing Bei Road, Shaoxing 312000, China. Email: [email protected]. Abstract: Fusion genes are known to drive and promote carcinogenesis and cancer progression. In recent years, the rapid development of biotechnologies has led to the discovery of a large number of fusion genes in prostate cancer specimens. To further investigate them, we summarized the fusion genes. We searched related articles in PubMed, CNKI (Chinese National Knowledge Infrastructure) and other databases, and the data of 92 literatures were summarized after preliminary screening. In this review, we summarized approximated 400 fusion genes since the first specific fusion TMPRSS2-ERG was discovered in prostate cancer in 2005. Some of these are prostate cancer specific, some are high-frequency in the prostate cancer of a certain ethnic group. This is a summary of scientific research in related fields and suggests that some fusion genes may become biomarkers or the targets for individualized therapies. -
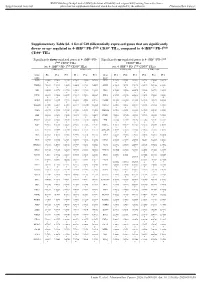
Supplementary Table S6. a List of 520 Differentially Expressed Genes
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Immunother Cancer Supplementary Table S6. A list of 520 differentially expressed genes that are significantly down- or up- regulated in 4-1BBpos PD-1high CD39+ TILs, compared to 4-1BBneg PD-1high CD39+ TILs Significantly down-regulated genes in 4-1BBpos PD- Significantly up-regulated genes in 4-1BBpos PD-1high 1high CD39+ TILs CD39+ TILs (vs. 4-1BBneg PD-1high CD39+ TILs) (vs. 4-1BBneg PD-1high CD39+ TILs) 4-1BBneg cells 4-1BBpos cells 4-1BBneg cells 4-1BBpos cells Gene Pt 1 Pt 2 Pt 3 Pt 1 Pt 2 Pt 3 Gene Pt 1 Pt 2 Pt 3 Pt 1 Pt 2 Pt 3 name name FOSB 9.92496 9.80436 9.61180 9.76085 9.55460 8.47625 SPC25 2.82975 2.53201 1.83699 3.05751 3.79646 3.66172 5 9 4 4 3 9 8 5 5 6 7 PTGER4 7.26693 7.28598 7.19060 6.84482 6.61319 5.90088 GMNN 4.55416 5.05351 3.79176 4.95671 5.63164 4.93921 3 5 1 4 8 7 8 5 3 3 ABI3 6.24122 6.45779 6.27735 5.85419 5.81360 5.18958 ING3 4.78602 5.23034 4.49070 5.22964 5.62759 5.34212 1 3 5 6 6 2 3 6 5 8 7 1 LITAF 9.06689 9.17420 8.99478 8.79217 8.74965 8.08497 KIF14 4.58732 4.59859 4.62494 5.12658 5.70643 5.26241 4 4 9 8 7 5 7 9 3 7 4 ZNF10 4.90912 4.98940 4.74775 4.33282 4.40018 3.50431 CENPE 6.54529 6.55450 6.47696 6.84945 7.22848 6.96595 8 7 8 8 2 8 5 7 9 5 3 2 TNFAIP3 11.4201 11.2484 11.4068 11.1877 10.9900 10.5350 CDCA7 3.80074 4.20586 3.92787 4.43496 5.46931 4.40814 9 4 2 9 7 9 9 5 8 NAPA 5.94770 5.79864 5.64849 -

Oxidized Phospholipids Regulate Amino Acid Metabolism Through MTHFD2 to Facilitate Nucleotide Release in Endothelial Cells
ARTICLE DOI: 10.1038/s41467-018-04602-0 OPEN Oxidized phospholipids regulate amino acid metabolism through MTHFD2 to facilitate nucleotide release in endothelial cells Juliane Hitzel1,2, Eunjee Lee3,4, Yi Zhang 3,5,Sofia Iris Bibli2,6, Xiaogang Li7, Sven Zukunft 2,6, Beatrice Pflüger1,2, Jiong Hu2,6, Christoph Schürmann1,2, Andrea Estefania Vasconez1,2, James A. Oo1,2, Adelheid Kratzer8,9, Sandeep Kumar 10, Flávia Rezende1,2, Ivana Josipovic1,2, Dominique Thomas11, Hector Giral8,9, Yannick Schreiber12, Gerd Geisslinger11,12, Christian Fork1,2, Xia Yang13, Fragiska Sigala14, Casey E. Romanoski15, Jens Kroll7, Hanjoong Jo 10, Ulf Landmesser8,9,16, Aldons J. Lusis17, 1234567890():,; Dmitry Namgaladze18, Ingrid Fleming2,6, Matthias S. Leisegang1,2, Jun Zhu 3,4 & Ralf P. Brandes1,2 Oxidized phospholipids (oxPAPC) induce endothelial dysfunction and atherosclerosis. Here we show that oxPAPC induce a gene network regulating serine-glycine metabolism with the mitochondrial methylenetetrahydrofolate dehydrogenase/cyclohydrolase (MTHFD2) as a cau- sal regulator using integrative network modeling and Bayesian network analysis in human aortic endothelial cells. The cluster is activated in human plaque material and by atherogenic lipo- proteins isolated from plasma of patients with coronary artery disease (CAD). Single nucleotide polymorphisms (SNPs) within the MTHFD2-controlled cluster associate with CAD. The MTHFD2-controlled cluster redirects metabolism to glycine synthesis to replenish purine nucleotides. Since endothelial cells secrete purines in response to oxPAPC, the MTHFD2- controlled response maintains endothelial ATP. Accordingly, MTHFD2-dependent glycine synthesis is a prerequisite for angiogenesis. Thus, we propose that endothelial cells undergo MTHFD2-mediated reprogramming toward serine-glycine and mitochondrial one-carbon metabolism to compensate for the loss of ATP in response to oxPAPC during atherosclerosis. -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses By
Downloaded from http://www.jimmunol.org/ by guest on September 23, 2021 Inducing is online at: average * The Journal of Immunology , 11 of which you can access for free at: 2013; 191:2956-2966; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication August 2013; doi: 10.4049/jimmunol.1300376 http://www.jimmunol.org/content/191/6/2956 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol cites 49 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 This article http://www.jimmunol.org/content/191/6/2956.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 23, 2021. The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN. -
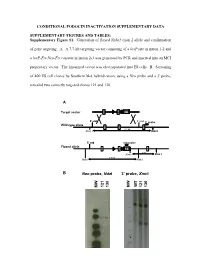
Supplementary Figure S1. Generation of Floxed Nphs2 Exon 2 Allele and Confirmation
CONDITIONAL PODOCIN INACTIVATION SUPPLEMENTARY DATA SUPPLEMENTARY FIGURES AND TABLES: Supplementary Figure S1. Generation of floxed Nphs2 exon 2 allele and confirmation of gene targeting. A. A 7.7-kb targeting vector consisting of a loxP site in intron 1-2 and a loxP-Frt-Neo-Frt cassette in intron 2-3 was generated by PCR and inserted into an MCI proprietary vector. The linearized vector was electroporated into ES cells. B. Screening of 400 ES cell clones by Southern blot hybridization, using a Neo probe and a 3' probe, revealed two correctly targeted clones 121 and 130. A Target vector 1 2 NEO 5’ end 3’ end 3’ probe Wild-type allele 1 2 3 9.2 kb Xmn I Xmn I 5’ end neo probe Floxed allele 1 2 NEO 3 5 kb Xmn I Xmn I 7.8 kb Nde I Nde I B Neo probe, NdeI 3’ probe, XmnI WT 121 130 121 130 MW MW - 1 - Supplementary Figure S2. Generation of Nphs2lox2/-,Cre+ mice and excision of exon 2 upon Cre recombinase induction. A. Triallelic Nphs2lox2/-,Cre+ mice were obtained by mating phenotypically normal Nphs2lox2/lox2 mice with Nphs2+/-,Cre+ mice. Mendelian inheritance of these alleles was observed. B. Genotypes were verified by multiplex PCR of tail genomic DNA. C. Cre recombinase activity was induced upon tamoxifen administration, leading to excision of the floxed exon 2 of the Nphs2 gene. D. Cre recombinase activity in the kidney was verified by PCR using a set of forward and reverse primers designed around exon 2 and demonstrating a 692-bp product (before Cre) and a 316-bp product (after Cre) using genomic DNA extracted from the renal cortex.