8137.Full.Pdf
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Global Architecture Shaping the Heterogeneity and Tissue-Dependency of the MHC Class I Immunopeptidome Is Evolutionarily Conserved
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.28.317750; this version posted September 29, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. The Global Architecture Shaping the Heterogeneity and Tissue-Dependency of the MHC Class I Immunopeptidome is Evolutionarily Conserved Authors Peter Kubiniok†1, Ana Marcu†2,3, Leon Bichmann†2,4, Leon Kuchenbecker4, Heiko Schuster1,5, David Hamelin1, Jérome Despault1, Kevin Kovalchik1, Laura Wessling1, Oliver Kohlbacher4,7,8,9,10 Stefan Stevanovic2,3,6, Hans-Georg Rammensee2,3,6, Marian C. Neidert11, Isabelle Sirois1, Etienne Caron1,12* Affiliations *Corresponding and Leading author: Etienne Caron ([email protected]) †Equal contribution to this work 1CHU Sainte-Justine Research Center, Montreal, QC H3T 1C5, Canada 2Department of Immunology, Interfaculty Institute for Cell Biology, University of Tübingen, Tübingen, Baden-Württemberg, 72076, Germany. 3Cluster of Excellence iFIT (EXC 2180) "Image-Guided and Functionally Instructed Tumor Therapies", University of Tübingen, Tübingen, Baden-Württemberg, 72076, Germany. 4Applied Bioinformatics, Dept. of Computer Science, University of Tübingen, Tübingen, Baden- Württemberg, 72074, Germany. 5Immatics Biotechnologies GmbH, Tübingen, 72076, Baden-Württemberg, Germany. 6DKFZ Partner Site Tübingen, German Cancer Consortium (DKTK), Tübingen, Baden- Württemberg, 72076, Germany. 7Institute for Bioinformatics and Medical Informatics, -

Supplemental Materials Supplemental Table 1
Electronic Supplementary Material (ESI) for RSC Advances. This journal is © The Royal Society of Chemistry 2016 Supplemental Materials Supplemental Table 1. The differentially expressed proteins from rat pancreas identified by proteomics (SAP vs. SO) No. Protein name Gene name ratio P value 1 Metallothionein Mt1m 3.35 6.34E-07 2 Neutrophil antibiotic peptide NP-2 Defa 3.3 8.39E-07 3 Ilf2 protein Ilf2 3.18 1.75E-06 4 Numb isoform o/o rCG 3.12 2.73E-06 5 Lysozyme Lyz2 3.01 5.63E-06 6 Glucagon Gcg 2.89 1.17E-05 7 Serine protease HTRA1 Htra1 2.75 2.97E-05 8 Alpha 2 macroglobulin cardiac isoform (Fragment) 2.75 2.97E-05 9 Myosin IF (Predicted) Myo1f 2.65 5.53E-05 10 Neuroendocrine secretory protein 55 Gnas 2.61 7.60E-05 11 Matrix metallopeptidase 8 Mmp8 2.57 9.47E-05 12 Protein Tnks1bp1 Tnks1bp1 2.53 1.22E-04 13 Alpha-parvin Parva 2.47 1.78E-04 14 C4b-binding protein alpha chain C4bpa 2.42 2.53E-04 15 Protein KTI12 homolog Kti12 2.41 2.74E-04 16 Protein Rab11fip5 Rab11fip5 2.41 2.84E-04 17 Protein Mcpt1l3 Mcpt1l3 2.33 4.43E-04 18 Phospholipase B-like 1 Plbd1 2.33 4.76E-04 Aldehyde dehydrogenase (NAD), cytosolic 19 2.32 4.93E-04 (Fragments) 20 Protein Dpy19l2 Dpy19l2 2.3 5.68E-04 21 Regenerating islet-derived 3 alpha, isoform CRA_a Reg3a 2.27 6.74E-04 22 60S acidic ribosomal protein P1 Rplp1 2.26 7.22E-04 23 Serum albumin Alb 2.25 7.98E-04 24 Ribonuclease 4 Rnase4 2.24 8.25E-04 25 Cct-5 protein (Fragment) Cct5 2.24 8.52E-04 26 Protein S100-A9 S100a9 2.22 9.71E-04 27 Creatine kinase M-type Ckm 2.21 1.00E-03 28 Protein Larp4b Larp4b 2.18 1.25E-03 -

Leishmania (L.) Amazonensis Peptidase Activities Inside the Living Cells and in Their Lysates
Molecular & Biochemical Parasitology 184 (2012) 82–89 Contents lists available at SciVerse ScienceDirect Molecular & Biochemical Parasitology Leishmania (L.) amazonensis peptidase activities inside the living cells and in their lysates a a b c a Elide E. Caroselli , Diego M. Assis , Clara L. Barbiéri , Wagner A.S. Júdice , Maria A. Juliano , d a,∗ Marcos L. Gazarini , Luiz Juliano a Department of Biophysics, Escola Paulista de Medicina, Universidade Federal de São Paulo, SP, Brazil b Department of Microbiology, Immunology and Parasitology, Escola Paulista de Medicina, Universidade Federal de São Paulo, SP, Brazil c Centro Interdisciplinar de Investigac¸ ão Bioquímica, Universidade de Mogi das Cruzes, Av. Dr. Cândido Xavier de Almeida Souza 200, 08780-911 Mogi das Cruzes, Brazil d Department of Biosciences, Universidade Federal de São Paulo, Santos, Brazil a r t i c l e i n f o a b s t r a c t Article history: In this study we investigated the peptidase activity in Leishmania (L.) amazonensis live amastigote by con- Received 24 November 2011 focal microscopy using peptidyl-MCA as substrates, the hydrolysis of which releases the MCA fluorophore Received in revised form 13 March 2012 inside the cells. Cell pre-treatment with peptidase inhibitors indicated the presence of cysteine and ser- Accepted 27 April 2012 ine peptidases. It was noteworthy that Leishmania amastigotes incorporate only substrates (Z-FR-MCA, Available online 6 May 2012 Z-RR-MCA) or inhibitors (E64, TLCK) containing positively charged groups. The peptidase activities in the supernatants of amastigotes and promastigotes lysates were also evaluated with the same peptidyl-MCA Keywords: substrates and inhibitors in the pH range 4.5–9.0. -

Supplementary Table 9. Functional Annotation Clustering Results for the Union (GS3) of the Top Genes from the SNP-Level and Gene-Based Analyses (See ST4)
Supplementary Table 9. Functional Annotation Clustering Results for the union (GS3) of the top genes from the SNP-level and Gene-based analyses (see ST4) Column Header Key Annotation Cluster Name of cluster, sorted by descending Enrichment score Enrichment Score EASE enrichment score for functional annotation cluster Category Pathway Database Term Pathway name/Identifier Count Number of genes in the submitted list in the specified term % Percentage of identified genes in the submitted list associated with the specified term PValue Significance level associated with the EASE enrichment score for the term Genes List of genes present in the term List Total Number of genes from the submitted list present in the category Pop Hits Number of genes involved in the specified term (category-specific) Pop Total Number of genes in the human genome background (category-specific) Fold Enrichment Ratio of the proportion of count to list total and population hits to population total Bonferroni Bonferroni adjustment of p-value Benjamini Benjamini adjustment of p-value FDR False Discovery Rate of p-value (percent form) Annotation Cluster 1 Enrichment Score: 3.8978262119731335 Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_CC_DIRECT GO:0005886~plasma membrane 383 24.33290978 5.74E-05 SLC9A9, XRCC5, HRAS, CHMP3, ATP1B2, EFNA1, OSMR, SLC9A3, EFNA3, UTRN, SYT6, ZNRF2, APP, AT1425 4121 18224 1.18857065 0.038655922 0.038655922 0.086284383 UP_KEYWORDS Membrane 626 39.77128335 1.53E-04 SLC9A9, HRAS, -

The Microbiota-Produced N-Formyl Peptide Fmlf Promotes Obesity-Induced Glucose
Page 1 of 230 Diabetes Title: The microbiota-produced N-formyl peptide fMLF promotes obesity-induced glucose intolerance Joshua Wollam1, Matthew Riopel1, Yong-Jiang Xu1,2, Andrew M. F. Johnson1, Jachelle M. Ofrecio1, Wei Ying1, Dalila El Ouarrat1, Luisa S. Chan3, Andrew W. Han3, Nadir A. Mahmood3, Caitlin N. Ryan3, Yun Sok Lee1, Jeramie D. Watrous1,2, Mahendra D. Chordia4, Dongfeng Pan4, Mohit Jain1,2, Jerrold M. Olefsky1 * Affiliations: 1 Division of Endocrinology & Metabolism, Department of Medicine, University of California, San Diego, La Jolla, California, USA. 2 Department of Pharmacology, University of California, San Diego, La Jolla, California, USA. 3 Second Genome, Inc., South San Francisco, California, USA. 4 Department of Radiology and Medical Imaging, University of Virginia, Charlottesville, VA, USA. * Correspondence to: 858-534-2230, [email protected] Word Count: 4749 Figures: 6 Supplemental Figures: 11 Supplemental Tables: 5 1 Diabetes Publish Ahead of Print, published online April 22, 2019 Diabetes Page 2 of 230 ABSTRACT The composition of the gastrointestinal (GI) microbiota and associated metabolites changes dramatically with diet and the development of obesity. Although many correlations have been described, specific mechanistic links between these changes and glucose homeostasis remain to be defined. Here we show that blood and intestinal levels of the microbiota-produced N-formyl peptide, formyl-methionyl-leucyl-phenylalanine (fMLF), are elevated in high fat diet (HFD)- induced obese mice. Genetic or pharmacological inhibition of the N-formyl peptide receptor Fpr1 leads to increased insulin levels and improved glucose tolerance, dependent upon glucagon- like peptide-1 (GLP-1). Obese Fpr1-knockout (Fpr1-KO) mice also display an altered microbiome, exemplifying the dynamic relationship between host metabolism and microbiota. -

Studies of Structure and Function of Tripeptidyl-Peptidase II
Till familj och vänner List of Papers This thesis is based on the following papers, which are referred to in the text by their Roman numerals. I. Eriksson, S.; Gutiérrez, O.A.; Bjerling, P.; Tomkinson, B. (2009) De- velopment, evaluation and application of tripeptidyl-peptidase II se- quence signatures. Archives of Biochemistry and Biophysics, 484(1):39-45 II. Lindås, A-C.; Eriksson, S.; Josza, E.; Tomkinson, B. (2008) Investiga- tion of a role for Glu-331 and Glu-305 in substrate binding of tripepti- dyl-peptidase II. Biochimica et Biophysica Acta, 1784(12):1899-1907 III. Eklund, S.; Lindås, A-C.; Hamnevik, E.; Widersten, M.; Tomkinson, B. Inter-species variation in the pH dependence of tripeptidyl- peptidase II. Manuscript IV. Eklund, S.; Kalbacher, H.; Tomkinson, B. Characterization of the endopeptidase activity of tripeptidyl-peptidase II. Manuscript Paper I and II were published under maiden name (Eriksson). Reprints were made with permission from the respective publishers. Contents Introduction ..................................................................................................... 9 Enzymes ..................................................................................................... 9 Enzymes and pH dependence .............................................................. 11 Peptidases ................................................................................................. 12 Serine peptidases ................................................................................. 14 Intracellular protein -

Handbook of Proteolytic Enzymes Second Edition Volume 1 Aspartic and Metallo Peptidases
Handbook of Proteolytic Enzymes Second Edition Volume 1 Aspartic and Metallo Peptidases Alan J. Barrett Neil D. Rawlings J. Fred Woessner Editor biographies xxi Contributors xxiii Preface xxxi Introduction ' Abbreviations xxxvii ASPARTIC PEPTIDASES Introduction 1 Aspartic peptidases and their clans 3 2 Catalytic pathway of aspartic peptidases 12 Clan AA Family Al 3 Pepsin A 19 4 Pepsin B 28 5 Chymosin 29 6 Cathepsin E 33 7 Gastricsin 38 8 Cathepsin D 43 9 Napsin A 52 10 Renin 54 11 Mouse submandibular renin 62 12 Memapsin 1 64 13 Memapsin 2 66 14 Plasmepsins 70 15 Plasmepsin II 73 16 Tick heme-binding aspartic proteinase 76 17 Phytepsin 77 18 Nepenthesin 85 19 Saccharopepsin 87 20 Neurosporapepsin 90 21 Acrocylindropepsin 9 1 22 Aspergillopepsin I 92 23 Penicillopepsin 99 24 Endothiapepsin 104 25 Rhizopuspepsin 108 26 Mucorpepsin 11 1 27 Polyporopepsin 113 28 Candidapepsin 115 29 Candiparapsin 120 30 Canditropsin 123 31 Syncephapepsin 125 32 Barrierpepsin 126 33 Yapsin 1 128 34 Yapsin 2 132 35 Yapsin A 133 36 Pregnancy-associated glycoproteins 135 37 Pepsin F 137 38 Rhodotorulapepsin 139 39 Cladosporopepsin 140 40 Pycnoporopepsin 141 Family A2 and others 41 Human immunodeficiency virus 1 retropepsin 144 42 Human immunodeficiency virus 2 retropepsin 154 43 Simian immunodeficiency virus retropepsin 158 44 Equine infectious anemia virus retropepsin 160 45 Rous sarcoma virus retropepsin and avian myeloblastosis virus retropepsin 163 46 Human T-cell leukemia virus type I (HTLV-I) retropepsin 166 47 Bovine leukemia virus retropepsin 169 48 -
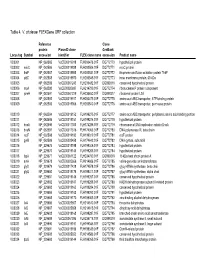
Table 4. V. Cholerae Flexgene ORF Collection
Table 4. V. cholerae FLEXGene ORF collection Reference Clone protein PlasmID clone GenBank Locus tag Symbol accession identifier FLEX clone name accession Product name VC0001 NP_062585 VcCD00019918 FLH200476.01F DQ772770 hypothetical protein VC0002 mioC NP_062586 VcCD00019938 FLH200506.01F DQ772771 mioC protein VC0003 thdF NP_062587 VcCD00019958 FLH200531.01F DQ772772 thiophene and furan oxidation protein ThdF VC0004 yidC NP_062588 VcCD00019970 FLH200545.01F DQ772773 inner membrane protein, 60 kDa VC0005 NP_062589 VcCD00061243 FLH236482.01F DQ899316 conserved hypothetical protein VC0006 rnpA NP_062590 VcCD00025697 FLH214799.01F DQ772774 ribonuclease P protein component VC0007 rpmH NP_062591 VcCD00061229 FLH236450.01F DQ899317 ribosomal protein L34 VC0008 NP_062592 VcCD00019917 FLH200475.01F DQ772775 amino acid ABC transporter, ATP-binding protein VC0009 NP_062593 VcCD00019966 FLH200540.01F DQ772776 amino acid ABC transproter, permease protein VC0010 NP_062594 VcCD00019152 FLH199275.01F DQ772777 amino acid ABC transporter, periplasmic amino acid-binding portion VC0011 NP_062595 VcCD00019151 FLH199274.01F DQ772778 hypothetical protein VC0012 dnaA NP_062596 VcCD00017363 FLH174286.01F DQ772779 chromosomal DNA replication initiator DnaA VC0013 dnaN NP_062597 VcCD00017316 FLH174063.01F DQ772780 DNA polymerase III, beta chain VC0014 recF NP_062598 VcCD00019182 FLH199319.01F DQ772781 recF protein VC0015 gyrB NP_062599 VcCD00025458 FLH174642.01F DQ772782 DNA gyrase, subunit B VC0016 NP_229675 VcCD00019198 FLH199346.01F DQ772783 hypothetical protein -

Proteases in MHC Class I Presentation and Cross-Presentation Kenneth L
Proteases in MHC Class I Presentation and Cross-Presentation Kenneth L. Rock, Diego J. Farfán-Arribas and Lianjun Shen This information is current as J Immunol 2010; 184:9-15; ; of October 1, 2021. doi: 10.4049/jimmunol.0903399 http://www.jimmunol.org/content/184/1/9 References This article cites 79 articles, 36 of which you can access for free at: Downloaded from http://www.jimmunol.org/content/184/1/9.full#ref-list-1 Why The JI? Submit online. http://www.jimmunol.org/ • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: by guest on October 1, 2021 http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2010 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Proteases in MHC Class I Presentation and Cross-Presentation Kenneth L. Rock, Diego J. Farfa´n-Arribas, and Lianjun Shen Cells that have mutated their genes or are virally is tolerant to them. However, if cells are synthesizing mutant infected are a potential threat to a host. Consequently, proteins or ones from viruses, then peptides from these gene the immune system has evolved mechanisms for CD8 T products will also be displayed, and this allows effector CD8 lymphocytes to identify such cells and eliminate them. -

Supplementary Table 2: Infection-Sensitive Genes
Supplementary Table 2: Infection-Sensitive Genes Supplementary Table 2: Upregulated with Adenoviral Infection (pages 1 - 7)- genes that were significant by ANOVA as well as significantly increased in control compared to both 'infected control' (Ad-LacZ) and 'calcineurin infected' (Ad-aCaN). Downregulated with Adenoviral Infection (pages 7 - 13) as above except for direction of change. Columns: Probe Set- Affymetrix probe set identifier for RG-U34A microarray, Symbol and Title- annotated information for above probe set (annotation downloaded June, 2004), ANOVA- p-value for 1-way Analysis of Variance, Uninfected, Ad-LacZ and Ad- aCaN- mean ± SEM expression for uninfected control, adenovirus mediated LacZ treated control, and adenovirus mediated calcineurin treated cultures respectively. Upregulated with Adenoviral Infection Probe Set Symbol Title ANOVA Uninfected Ad-LacZ Ad-aCaN Z48444cds_at Adam10 a disintegrin and metalloprotease domain 10 0.00001 837 ± 31 1107 ± 24 1028 ± 29 L26986_at Adcy8 adenylyl cyclase 8 0.00028 146 ± 12 252 ± 21 272 ± 21 U94340_at Adprt ADP-ribosyltransferase 1 0.00695 2096 ± 100 2783 ± 177 2586 ± 118 U01914_at Akap8 A kinase anchor protein 8 0.00080 993 ± 44 1194 ± 65 1316 ± 44 AB008538_at Alcam activated leukocyte cell adhesion molecule 0.01012 2775 ± 136 3580 ± 216 3429 ± 155 M34176_s_at Ap2b1 adaptor-related protein complex 2, beta 1 subunit 0.01389 2408 ± 199 2947 ± 143 3071 ± 116 D44495_s_at Apex1 apurinic/apyrimidinic endonuclease 1 0.00089 4959 ± 185 5816 ± 202 6057 ± 158 U16245_at Aqp5 aquaporin 5 0.02710 -

Genome-Wide Profiling of Druggable Active Tumor Defense Mechanisms to Enhance Cancer Immunotherapy
bioRxiv preprint doi: https://doi.org/10.1101/843185; this version posted November 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Genome-wide profiling of druggable active tumor defense mechanisms to enhance cancer immunotherapy Rigel J. Kishton1,2,*,#, Shashank J. Patel1,2,†,*, Suman K. Vodnala1,2, Amy E. Decker3, Yogin Patel1,2, Madhusudhanan Sukumar1,2, Tori N. Yamamoto1,2,4, Zhiya Yu1,2, Michelle Ji1,2, Amanda N. Henning1,2, Devikala Gurusamy1,2, Douglas C. Palmer1,2, Winifred Lo1, Anna Pasetto1, Parisa Malekzadeh1, Drew C. Deniger1, Kris C. Wood3, Neville E. Sanjana5,6, Nicholas P. Restifo1,2, #, § 1Surgery Branch, Center for Cancer Research, National Cancer Institute, Bethesda, MD 20892, USA 2Center for Cell-Based Therapy, National Cancer Institute, Bethesda, MD 20892, USA 3Department of Pharmacology & Cancer Biology, Duke University School of Medicine, Durham, NC, USA 4Immunology Graduate Group, University of Pennsylvania, Philadelphia, PA 19104, USA 5New York Genome Center, New York, NY 10013 USA 6Department of Biology, New York University, New York, NY 10003, USA *These authors contributed equally to this work. †Present address: NextCure Inc., Beltsville, MD 20705, USA §Present address: Lyell Immunopharma, South San Francisco, CA 94080, USA #Corresponding authors. NPR: [email protected]. RJK: [email protected]. bioRxiv preprint doi: https://doi.org/10.1101/843185; this version posted November 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. -

Pa28αβ: the Enigmatic Magic Ring of the Proteasome?
Biomolecules 2014, 4, 566-584; doi:10.3390/biom4020566 OPEN ACCESS biomolecules ISSN 2218-273X www.mdpi.com/journal/biomolecules/ Review PA28: The Enigmatic Magic Ring of the Proteasome? Paolo Cascio Department of Veterinary Sciences, University of Turin, Grugliasco 10095, Italy; E-Mail: [email protected]; Tel.: +39-011-670-9113; Fax: +39-011-670-9138 Received: 4 April 2014; in revised form: 15 May 2014 / Accepted: 8 June 2014 / Published: 19 June 2014 Abstract: PA28 is a -interferon-induced 11S complex that associates with the ends of the 20S proteasome and stimulates in vitro breakdown of small peptide substrates, but not proteins or ubiquitin-conjugated proteins. In cells, PA28 also exists in larger complexes along with the 19S particle, which allows ATP-dependent degradation of proteins; although in vivo a large fraction of PA28 is present as PA28-20S particles whose exact biological functions are largely unknown. Although several lines of evidence strongly indicate that PA28 plays a role in MHC class I antigen presentation, the exact molecular mechanisms of this activity are still poorly understood. Herein, we review current knowledge about the biochemical and biological properties of PA28 and discuss recent findings concerning its role in modifying the spectrum of proteasome’s peptide products, which are important to better understand the molecular mechanisms and biological consequences of PA28 activity. Keywords: PA28; proteasomes; immunoproteasomes; protein degradation; MHC class I antigen presentation; epitopes; antigenic peptides 1. MHC Class I Antigen Presentation The continual presentation of intracellular proteins fragments on major histocompatibility complex (MHC) class I molecules is a process that allows cytotoxic CD8+ T lymphocytes (CTLs) to identify and selectively eliminate cells that synthesize foreign (e.g., viral) or abnormal (e.g., oncogene products) proteins [1,2].