1 a Pan Genome Reverse Vaccinology Approach to Prevent Streptococcus
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Shrimp Fishing in Mexico
235 Shrimp fishing in Mexico Based on the work of D. Aguilar and J. Grande-Vidal AN OVERVIEW Mexico has coastlines of 8 475 km along the Pacific and 3 294 km along the Atlantic Oceans. Shrimp fishing in Mexico takes place in the Pacific, Gulf of Mexico and Caribbean, both by artisanal and industrial fleets. A large number of small fishing vessels use many types of gear to catch shrimp. The larger offshore shrimp vessels, numbering about 2 212, trawl using either two nets (Pacific side) or four nets (Atlantic). In 2003, shrimp production in Mexico of 123 905 tonnes came from three sources: 21.26 percent from artisanal fisheries, 28.41 percent from industrial fisheries and 50.33 percent from aquaculture activities. Shrimp is the most important fishery commodity produced in Mexico in terms of value, exports and employment. Catches of Mexican Pacific shrimp appear to have reached their maximum. There is general recognition that overcapacity is a problem in the various shrimp fleets. DEVELOPMENT AND STRUCTURE Although trawling for shrimp started in the late 1920s, shrimp has been captured in inshore areas since pre-Columbian times. Magallón-Barajas (1987) describes the lagoon shrimp fishery, developed in the pre-Hispanic era by natives of the southeastern Gulf of California, which used barriers built with mangrove sticks across the channels and mouths of estuaries and lagoons. The National Fisheries Institute (INP, 2000) and Magallón-Barajas (1987) reviewed the history of shrimp fishing on the Pacific coast of Mexico. It began in 1921 at Guaymas with two United States boats. -

Shrimp: Oceana Reveals Misrepresentation of America's
Shrimp: Oceana Reveals Misrepresentation of America’s Favorite Seafood October 2014 Authors: Kimberly Warner, Ph.D., Rachel Golden, Beth Lowell, Carlos Disla, Jacqueline Savitz and Michael Hirshfield, Ph.D. Executive Summary With shrimp, it is almost impossible to know what you are getting. Shrimp is the most commonly consumed seafood in the United States and the most highly traded seafood in the world. However, this high demand has led to many environmental and human rights abuses in the fishing, farming and processing of shrimp. Despite the popularity of shrimp, as well as the associated sustainability, human rights and environmental concerns, U.S. consumers are routinely given little information about the shrimp they purchase, making it nearly impossible to find and follow sound sustainability recommendations. Oceana’s previous studies have shown that species substitution, a form of seafood fraud, is common in the U.S. Last year, Oceana found that one-third of the more than 1,200 fish samples it tested nationwide were mislabeled, according to Food and Drug Administration guidelines. We have now turned our attention to shrimp, American’s most popular seafood, to investigate mislabeling as well as the information that consumers are given about the products they purchase. Consumers may wish to choose their shrimp more carefully for many important social and ecological reasons. For instance, consumers may wish to avoid shrimp caught in fisheries that are not responsibly managed, that have high rates of waste or discards, or that are associated with human rights abuses. At the same time, consumers may wish to avoid farmed shrimp due to health and environmental impacts. -

Beaver Street Fisheries, Inc
Why Participate? How ODP Works What's Included? About Us News Beaver Street Fisheries, Inc. Beaver Street Fisheries is a leading importer, manufacturer and distributor of quality frozen seafood products from the USA and around the world. With headquarters in Jacksonville, Florida, a vertically integrated supply chain, and the advantage of both on-site and off-shore processing capabilities, Beaver Street Fisheries offers a wide variety of products, competitive pricing, and can satisfy the diverse needs of wholesale, retail, institutional and foodservice operators. The success and reputation that Beaver Street Fisheries enjoys is attributed to its dedication to undeniable quality, efficient, and attentive service and the disciplined exercise of a single principle, "Treat the customer as you would a friend and all else will follow.” 2019 Number of Wild Caught Number of Certified Number of Fisheries in a Number of Farmed Species Used Fisheries FIP Species Used 21 16 11 3 Production Methods Used · Bottom trawl · Purse seine · Longlines · Rake / hand gathered / · Dredge · Handlines and pole-lines hand netted · Pots and traps · Farmed Summary For over seventy year, Beaver Street Fisheries has always been a leader in the seafood industry, and we understand that we have a global responsibility to support and sustain the earth and its ecosystems. As part of our commitment to sustainability and responsible sourcing, we work closely with our supply chain partners to embrace strategies to support the ever-growing need for responsible seafood from around the world. We do this by working with standard-setting organizations for wild caught and aquaculture seafood. Additionally, we have partnered with Sustainable Fisheries Partnership (SFP) to help us develop and implement fishery improvement projects for both wild and farmed raised species. -

Presence of Pacific White Shrimp Litopenaeus Vannamei (Boone, 1931) in the Southern Gulf of Mexico
Aquatic Invasions (2011) Volume 6, Supplement 1: S139–S142 doi: 10.3391/ai.2011.6.S1.031 Open Access © 2011 The Author(s). Journal compilation © 2011 REABIC Aquatic Invasions Records Presence of Pacific white shrimp Litopenaeus vannamei (Boone, 1931) in the Southern Gulf of Mexico Armando T. Wakida-Kusunoki1*, Luis Enrique Amador-del Angel2, Patricia Carrillo Alejandro1 and Cecilia Quiroga Brahms1 1Instituto Nacional de Pesca, Ave. Héroes del 21 de Abril s/n. Col Playa Norte, Ciudad del Carmen Campeche, México 2Universidad Autónoma del Carmen, Centro de Investigación de Ciencias Ambientales (CICA), Ave. Laguna de Términos s/n Col. Renovación 2da Sección, C.P. 24155, Ciudad del Carmen, Campeche, México E-mail: [email protected] (ATWK), [email protected] (LEAA), [email protected] (PCA), [email protected] (CQB) *Corresponding author Received: 12 July 2011 / Accepted: 12 October 2011 / Published online: 27 October 2011 Abstract This is the first report of the presence of Pacific white shrimp Litopenaeus vannamei in the Southern Gulf of Mexico coast. Seven specimens were collected in the Carmen-Pajonal-Machona lagoons near La Azucena and Sanchez Magallanes in Tabasco, Mexico, during a shrimp monitoring program survey conducted in this area. Further sampling and monitoring are required to find evidence that confirms the establishment of a population of Pacific white shrimp L. vannamei in Southern Gulf of Mexico. Key words: Litopenaeus vannamei, Pacific white shrimp, invasive species, Tabasco, Mexico Introduction covering 319.6 ha (Diario Oficial de la Federacion 2011). Almost all of these farms are Litopenaeus vannamei (Boone, 1931) is native to located in the Southern part of the Machona the Eastern Pacific coast from the Gulf of Lagoon. -

Assessment and Modeling of Whiteleg Shrimp Production in a Low-Salinity Recirculating Aquaculture System
ASSESSMENT AND MODELING OF WHITELEG SHRIMP PRODUCTION IN A LOW-SALINITY RECIRCULATING AQUACULTURE SYSTEM by Si Qi (Cindy) Yao A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science School of Environment and Sustainability University of Michigan August 2018 Thesis Committee: Professor James Diana Professor Lutgarde Raskin ii Acknowledgments There are many people to thank for their contributions and support that made this thesis research possible. I am deeply grateful to have been co-advised by Prof. Jim Diana and Prof. Lut Raskin, whose combined insights on aquaculture and water treatment guided my research in purposeful directions. Being part of two research groups was a great opportunity to share ideas across a diverse range of projects and learn from the experiences of fellow students with different perspectives. Operation of the pilot aquaculture system was possible with the help of undergraduate research assistant Lindsay Rasmussen, who was instrumental in assisting with system maintenance, shrimp handling, and water analysis. Two Doris Duke Conservation Scholar fellows, Citralina Haruo and Jannice Newson, made valuable contributions to setting up the pilot system early on. The undergraduate senior design team consisting of Harrison Addy, Michael Graves, Katherine Rouen, and Jeremy Solomon used their mechanical engineering experience to design and construct the pilot system. I am also thankful for the valuable support with lab operations offered by Sucila Fernandes, whose dedication to ensuring that proper conditions are maintained made her a great resource. Timely processing of shrimp deliveries was facilitated by the reliable communications from Jane Parish. Additional thanks to Tom Yavaraski and Nivea Vydiswaran for their help in the environmental engineering labs. -

Asc Shrimp Standard Revision
ASC SHRIMP STANDARD REVISION Revision of Current Metrics Background Analysis Document March 2020 Revision of current metrics – Background analysis document Shrimp Standard Revision Purpose The purpose of this document is to present the acquired data for the revision of the ASC Shrimp Standard v.1.1 and propose changes to the metric requirements where relevant. This document will be used for the decision-making process within the revision. Background The ASC Shrimp Standard v.1.1 is based on the anterior work of the Shrimp Aquaculture Dialogue (ShAD) and sets requirements that define what has been deemed ‘acceptable’ levels as regards the major social and environmental impacts of saltwater shrimp farming. The purpose of the ASC Shrimp Standard was and is to provide means to measurably improve the environmental and social performance of shrimp aquaculture operations worldwide. The Standard currently covers species under the genus Penaeus (previously Litopenaeus)1 and is oriented towards the production of P. vannamei2 and P. monodon. A Rationale document3 was produced as part of the ASC Shrimp Standard revision to evaluate the necessity to specifically include Penaeus stylirostris (Blue Shrimp), Penaeus merguiensis (Banana Prawn), Penaeus japonicus (Kuruma Prawn) and Penaeus ensis (Greasyback Shrimp) within the ASC Shrimp Standard. It was concluded that specific metrics for these species are not necessary and certification can remain on the basis of the metrics already contained therein for P. vannamei and P. monodon. Corresponding Metrics The ASC Shrimp Standard covers seven principles regarding legal regulations, environmentally suitable sighting and operation, community interactions, responsible operation practices, shrimp health management, stock management and resources use. -

Sensory Systems and Feeding Behaviour of the Giant Freshwater Prawn, Macrobrachium Rosenbergii, and the Marine Whiteleg Shrimp, Litopenaeus Vannamei
Borneo Journal of Marine Science and Aquaculture Volume: 01 | December 2017, 80 - 91 Sensory systems and feeding behaviour of the giant freshwater prawn, Macrobrachium rosenbergii, and the marine whiteleg shrimp, Litopenaeus vannamei Gunzo Kawamura1*, Teodora Uy Bagarinao2 and Annita Seok Kian Yong1 1Borneo Marine Research Institute, Universiti Malaysia Sabah, 88400 Kota Kinabalu, Sabah, Malaysia 2Aquaculture Department, Southeast Asian Fisheries Development Center, Tigbauan, Iloilo, Philippines *Corresponding author: [email protected] Abstract Information on the sensory basis of shrimp feeding provides the means for assessment of the effectiveness of food items in terms of smell, taste, size, and colour. This chapter summarizes information about the sensory basis of the feeding behaviour of the giant freshwater prawn (Macrobrachium rosenbergii) and the marine whiteleg shrimp (Litopenaeus vannamei). Existing literature on these shrimp species and other decapod crustaceans is reviewed, and unpublished experiments using the selective sensory ablation technique to determine the involvement of vision, chemoreception, and touch sense in the feeding behavior of the juveniles of M. rosenbergii and L. vannamei are also described. To determine the role of vision in feeding, the eyes of the juveniles were painted over (deprived of vision) with white manicure and their feeding response to commercial pellets was compared with those with untreated eyes. The untreated eyed juveniles detected and approached a feed pellet right away, but the specimens blinded by the coating detected a pellet only after random accidental touch with the walking legs while roaming on the aquarium bottom. Juveniles that had learned to feed on pellets showed food search and manipulation responses to a pellet-like pebble without smell and taste. -

Shrimp Welfare (2020 Recommended Idea)
CE Research Report: Animal Welfare - Shrimp Welfare, 2020 Page 2 Research Report: Animal Welfare – Shrimp Welfare (2020 Recommended Idea) Primary author: Vicky Cox Review: Karolina Sarek & Erik Hausen Date of publication: August 2020 Research period: 2020 This is a summary report about improving whiteleg shrimp (Litopenaeus vannamei) welfare. In our five-step research process this report corresponds to step 4, the drafting of an in-depth, 80-hour report on a potential intervention. All the ideas considered for animal advocacy are listed in this spreadsheet. Other reports on animal welfare can be found here. Thanks to Michael St. Jules, Karolina Sarek, Erik Hausen, and Daniela Waldhorn for reviewing the research, and to Antonia Shann, Nicoleta Faina, Bella Forristal, Patrick Stadler, and Urszula Zarosa for their contributions to this report. We are also grateful to the seven experts who took the time to offer their thoughts on this research. For questions about the content of this research please contact Vicky Cox at [email protected]. For questions about the research process, charity recommendations, and intervention comparisons please contact Karolina Sarek at [email protected]. Charity Entrepreneurship is a research and training program that incubates multiple high-impact charities annually. Our mission is to cause more effective charities to exist in the world by connecting talented individuals with high-impact intervention opportunities. We achieve this through an extensive research process and through our Incubation Program. CE Research Report: Animal Welfare - Shrimp Welfare, 2020 Page 3 Research Process Before opening the report, we think it is important to introduce our research process. Knowing the principles of the process helps readers understand how we formed our conclusions and enables greater reasoning transparency. -

Whiteleg Shrimp Criterion 1
Whiteleg Shrimp, Giant Tiger Prawn Litopenaeus vannamei, Penaeus monodon Image © Monterey Bay Aquarium India Ponds September 21, 2015 Matthew Thompson, Independent Research Analyst Disclaimer Seafood Watch® strives to have all Seafood Reports reviewed for accuracy and completeness by external scientists with expertise in ecology, fisheries science and aquaculture. Scientific review, however, does not constitute an endorsement of the Seafood Watch® program or its recommendations on the part of the reviewing scientists. Seafood Watch® is solely responsible for the conclusions reached in this report. 2 About Seafood Watch® Monterey Bay Aquarium’s Seafood Watch® program evaluates the ecological sustainability of wild-caught and farmed seafood commonly found in the United States marketplace. Seafood Watch® defines sustainable seafood as originating from sources, whether wild-caught or farmed, which can maintain or increase production in the long-term without jeopardizing the structure or function of affected ecosystems. Seafood Watch® makes its science-based recommendations available to the public in the form of regional pocket guides that can be downloaded from www.seafoodwatch.org. The program’s goals are to raise awareness of important ocean conservation issues and empower seafood consumers and businesses to make choices for healthy oceans. Each sustainability recommendation on the regional pocket guides is supported by a Seafood Report. Each report synthesizes and analyzes the most current ecological, fisheries and ecosystem science on a species, then evaluates this information against the program’s conservation ethic to arrive at a recommendation of “Best Choices,” “Good Alternatives” or “Avoid.” The detailed evaluation methodology is available upon request. In producing the Seafood Reports, Seafood Watch® seeks out research published in academic, peer-reviewed journals whenever possible. -

Laemson Marine National Park - Kraburi Estuary Wetlands, Thailand Bampen Chaiyarak and Kittama Khunthong
Climate Change Vulnerability Assessment Kaper Estuary - Laemson Marine National Park - Kraburi Estuary Wetlands, Thailand Bampen Chaiyarak and Kittama Khunthong Mekong WET: Building Resilience of Wetlands in the Lower Mekong Region Climate Change Vulnerability Assessment Kaper Estuary - Laemson Marine National Park - Kraburi Estuary Wetlands, Thailand Bampen Chaiyarak and Kittama Khunthong The designation of geographical entities in this report, and the presentation of the material, do not imply the expression of any opinion whatsoever on the part of IUCN or the German Federal Ministry for the Environment, Nature Conservation, Building and Nuclear Safety. The views expressed in this publication do not necessarily reflect those of IUCN or the German Federal Ministry for the Environment, Nature Conservation, Building and Nuclear Safety. Special acknowledgement to the International Climate Initiative of the German Federal Ministry for the Environment, Nature Conservation, Building and Nuclear Safety for supporting Mekong WET. Published by: IUCN Asia Regional Office (ARO), Bangkok, Thailand Copyright: © 2019 IUCN, International Union for Conservation of Nature and Natural Resources Reproduction of this publication for educational or other non-commercial purposes is authorised without prior written permission from the copyright holder provided the source is fully acknowledged. Reproduction of this publication for resale or other commercial purposes is prohibited without prior written permission of the copyright holder. Citation: Chaiyarah, -

Shrimp Aquaculture Landscape
Shrimp Aquaculture Landscape January 25, 2018 Table of contents • Introduction Slides 3-8 1 Prologue • Executive Summary • Key takeaways Slides 9-31 Production • Trends in production and productivity 2 • Environmental concerns trends • Country deep-dives • Supply chain structures and market-based initiatives Slides 32-48 3 Supply chain • Deep-dive: Southeast Asian supply chains • OSMI objective prioritization framework and results chain assessment • Context for investing in shrimp Slides 49-59 4 Financing • Risk, return, and access to finance along the value chain • Value proposition and impact investment considerations 2 • Introduction 1 Prologue • Executive Summary • Key takeaways 3 Prologue Introduction In late October 2017, the Gordon and Betty Moore Foundation’s (Moore) Ocean and Seafood Markets Initiative (OSMI) asked CEA to aggregate information on a series of key questions relating to farmed shrimp to inform their Advisory Committee’s (AC) December 12th and 13th meetings. As we understand it, our research and insights will inform the AC’s future farmed shrimp work planning that will take place in early 2018. CEA was asked to provide detailed responses to the following topics: • Production: Document global production trends by geography over the past 20 years including countries of origin, intensity of production (land use vs. total production), key markets, key differences in production systems, and their impacts and other issue (e.g. feed, disease, efficiency, mangroves, cumulative impacts). • Value Chain: Lay out/revisit the value chain (as represented in the Shrimp Results Chain) from production to local and global markets. How does shrimp move from the producer in 5 key production geographies to the consumer in 5 key consumption markets? • Financing: What types of finance (investment vs. -
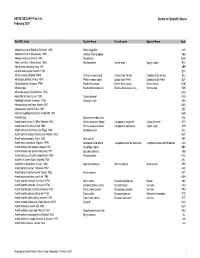
ASFIS ISSCAAP Fish List February 2007 Sorted on Scientific Name
ASFIS ISSCAAP Fish List Sorted on Scientific Name February 2007 Scientific name English Name French name Spanish Name Code Abalistes stellaris (Bloch & Schneider 1801) Starry triggerfish AJS Abbottina rivularis (Basilewsky 1855) Chinese false gudgeon ABB Ablabys binotatus (Peters 1855) Redskinfish ABW Ablennes hians (Valenciennes 1846) Flat needlefish Orphie plate Agujón sable BAF Aborichthys elongatus Hora 1921 ABE Abralia andamanika Goodrich 1898 BLK Abralia veranyi (Rüppell 1844) Verany's enope squid Encornet de Verany Enoploluria de Verany BLJ Abraliopsis pfefferi (Verany 1837) Pfeffer's enope squid Encornet de Pfeffer Enoploluria de Pfeffer BJF Abramis brama (Linnaeus 1758) Freshwater bream Brème d'eau douce Brema común FBM Abramis spp Freshwater breams nei Brèmes d'eau douce nca Bremas nep FBR Abramites eques (Steindachner 1878) ABQ Abudefduf luridus (Cuvier 1830) Canary damsel AUU Abudefduf saxatilis (Linnaeus 1758) Sergeant-major ABU Abyssobrotula galatheae Nielsen 1977 OAG Abyssocottus elochini Taliev 1955 AEZ Abythites lepidogenys (Smith & Radcliffe 1913) AHD Acanella spp Branched bamboo coral KQL Acanthacaris caeca (A. Milne Edwards 1881) Atlantic deep-sea lobster Langoustine arganelle Cigala de fondo NTK Acanthacaris tenuimana Bate 1888 Prickly deep-sea lobster Langoustine spinuleuse Cigala raspa NHI Acanthalburnus microlepis (De Filippi 1861) Blackbrow bleak AHL Acanthaphritis barbata (Okamura & Kishida 1963) NHT Acantharchus pomotis (Baird 1855) Mud sunfish AKP Acanthaxius caespitosa (Squires 1979) Deepwater mud lobster Langouste