Mouse Scfd1 Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Identification of Differentially Expressed Genes in Human Bladder Cancer Through Genome-Wide Gene Expression Profiling
521-531 24/7/06 18:28 Page 521 ONCOLOGY REPORTS 16: 521-531, 2006 521 Identification of differentially expressed genes in human bladder cancer through genome-wide gene expression profiling KAZUMORI KAWAKAMI1,3, HIDEKI ENOKIDA1, TOKUSHI TACHIWADA1, TAKENARI GOTANDA1, KENGO TSUNEYOSHI1, HIROYUKI KUBO1, KENRYU NISHIYAMA1, MASAKI TAKIGUCHI2, MASAYUKI NAKAGAWA1 and NAOHIKO SEKI3 1Department of Urology, Graduate School of Medical and Dental Sciences, Kagoshima University, 8-35-1 Sakuragaoka, Kagoshima 890-8520; Departments of 2Biochemistry and Genetics, and 3Functional Genomics, Graduate School of Medicine, Chiba University, 1-8-1 Inohana, Chuo-ku, Chiba 260-8670, Japan Received February 15, 2006; Accepted April 27, 2006 Abstract. Large-scale gene expression profiling is an effective CKS2 gene not only as a potential biomarker for diagnosing, strategy for understanding the progression of bladder cancer but also for staging human BC. This is the first report (BC). The aim of this study was to identify genes that are demonstrating that CKS2 expression is strongly correlated expressed differently in the course of BC progression and to with the progression of human BC. establish new biomarkers for BC. Specimens from 21 patients with pathologically confirmed superficial (n=10) or Introduction invasive (n=11) BC and 4 normal bladder samples were studied; samples from 14 of the 21 BC samples were subjected Bladder cancer (BC) is among the 5 most common to microarray analysis. The validity of the microarray results malignancies worldwide, and the 2nd most common tumor of was verified by real-time RT-PCR. Of the 136 up-regulated the genitourinary tract and the 2nd most common cause of genes we detected, 21 were present in all 14 BCs examined death in patients with cancer of the urinary tract (1-7). -

Developing Specific Molecular Biomarkers for Thermal Stress In
Akbarzadeh et al. BMC Genomics (2018) 19:749 https://doi.org/10.1186/s12864-018-5108-9 RESEARCHARTICLE Open Access Developing specific molecular biomarkers for thermal stress in salmonids Arash Akbarzadeh1,2* , Oliver P Günther3, Aimee Lee Houde1, Shaorong Li1, Tobi J Ming1, Kenneth M Jeffries4, Scott G Hinch5 and Kristina M Miller1 Abstract Background: Pacific salmon (Oncorhynchus spp.) serve as good biological indicators of the breadth of climate warming effects on fish because their anadromous life cycle exposes them to environmental challenges in both marine and freshwater environments. Our study sought to mine the extensive functional genomic studies in fishes to identify robust thermally-responsive biomarkers that could monitor molecular physiological signatures of chronic thermal stress in fish using non-lethal sampling of gill tissue. Results: Candidate thermal stress biomarkers for gill tissue were identified using comparisons among microarray datasets produced in the Molecular Genetics Laboratory, Pacific Biological Station, Nanaimo, BC, six external, published microarray studies on chronic and acute temperature stress in salmon, and a comparison of significant genes across published studies in multiple fishes using deep literature mining. Eighty-two microarray features related to 39 unique gene IDs were selected as candidate chronic thermal stress biomarkers. Most of these genes were identified both in the meta-analysis of salmon microarray data and in the literature mining for thermal stress markers in salmonids and other fishes. Quantitative reverse transcription PCR (qRT-PCR) assays for 32 unique genes with good efficiencies across salmon species were developed, and their activity in response to thermally challenged sockeye salmon (O. nerka)and Chinook salmon (O. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

Characterization of a Novel Monogenic Form of Liver Failure
FAKULTÄT FÜR MEDIZIN INSTITUT FÜR HUMANGENETIK Lehrstuhl: Prof. Dr. Thomas Meitinger Characterization of a Novel Monogenic Form of Liver Failure Marlies Gwendolyn Köpke Vollständiger Abdruck der von der Fakultät für Medizin der Technischen Universität München zur Erlangung des akademischen Grades eines Doktors der Medizin (Dr. med.) genehmigten Dissertation. Vorsitzender: Prof. Dr. Ernst J. Rummeny Prüfer der DissertAtion: 1. Prof. Dr. Thomas Meitinger 2. Prof. Dr. Percy A. Knolle Die Dissertation wurde am 07.08.2017 bei der Technischen Universität München eingereicht und durch die Fakultät für Medizin am 04.07.2018 angenommen. Marlies G. Köpke ChArActerizAtion of A Novel Monogenic Form of Liver Failure Table of Contents Table of contents List of abbreviations ................................................................................................. V List of prefixes ........................................................................................................ VIII Introduction ............................................................................................................... 1 Acute liver failure in children ............................................................................................ 1 Definition .......................................................................................................................... 1 Epidemiology ................................................................................................................... 2 Pathophysiology .............................................................................................................. -

Genetic Analysis of Amyotrophic Lateral Sclerosis Identifies Contributing Pathways and Cell Types
University of Massachusetts Medical School eScholarship@UMMS Open Access Publications by UMMS Authors 2021-01-15 Genetic analysis of amyotrophic lateral sclerosis identifies contributing pathways and cell types Sara Saez-Atienzar National Institutes of Health Et al. Let us know how access to this document benefits ou.y Follow this and additional works at: https://escholarship.umassmed.edu/oapubs Part of the Genetics and Genomics Commons, Molecular and Cellular Neuroscience Commons, Nervous System Diseases Commons, and the Neurology Commons Repository Citation Saez-Atienzar S, Landers JE. (2021). Genetic analysis of amyotrophic lateral sclerosis identifies contributing pathways and cell types. Open Access Publications by UMMS Authors. https://doi.org/ 10.1126/sciadv.abd9036. Retrieved from https://escholarship.umassmed.edu/oapubs/4570 Creative Commons License This work is licensed under a Creative Commons Attribution-Noncommercial 4.0 License This material is brought to you by eScholarship@UMMS. It has been accepted for inclusion in Open Access Publications by UMMS Authors by an authorized administrator of eScholarship@UMMS. For more information, please contact [email protected]. SCIENCE ADVANCES | RESEARCH ARTICLE GENETICS Copyright © 2021 The Authors, some rights reserved; Genetic analysis of amyotrophic lateral sclerosis exclusive licensee American Association identifies contributing pathways and cell types for the Advancement Sara Saez-Atienzar1*†, Sara Bandres-Ciga2,3†, Rebekah G. Langston4, Jonggeol J. Kim2, of Science. No claim to 5 6,7,8 original U.S. Government Shing Wan Choi , Regina H. Reynolds , the International ALS Genomics Consortium, Works. Distributed 1,9 1 10 ITALSGEN, Yevgeniya Abramzon , Ramita Dewan , Sarah Ahmed , under a Creative 11 1 7,8 4 2,12 John E. -

Tepzz 8Z6z54a T
(19) TZZ ZZ_T (11) EP 2 806 054 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 26.11.2014 Bulletin 2014/48 C40B 40/06 (2006.01) C12Q 1/68 (2006.01) C40B 30/04 (2006.01) C07H 21/00 (2006.01) (21) Application number: 14175049.7 (22) Date of filing: 28.05.2009 (84) Designated Contracting States: (74) Representative: Irvine, Jonquil Claire AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HGF Limited HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL 140 London Wall PT RO SE SI SK TR London EC2Y 5DN (GB) (30) Priority: 28.05.2008 US 56827 P Remarks: •Thecomplete document including Reference Tables (62) Document number(s) of the earlier application(s) in and the Sequence Listing can be downloaded from accordance with Art. 76 EPC: the EPO website 09753364.0 / 2 291 553 •This application was filed on 30-06-2014 as a divisional application to the application mentioned (71) Applicant: Genomedx Biosciences Inc. under INID code 62. Vancouver, British Columbia V6J 1J8 (CA) •Claims filed after the date of filing of the application/ after the date of receipt of the divisional application (72) Inventor: Davicioni, Elai R.68(4) EPC). Vancouver British Columbia V6J 1J8 (CA) (54) Systems and methods for expression- based discrimination of distinct clinical disease states in prostate cancer (57) A system for expression-based discrimination of distinct clinical disease states in prostate cancer is provided that is based on the identification of sets of gene transcripts, which are characterized in that changes in expression of each gene transcript within a set of gene transcripts can be correlated with recurrent or non- recur- rent prostate cancer. -

The SARS-Cov-2 RNA–Protein Interactome in Infected Human Cells
The SARS-CoV-2 RNA–protein interactome in infected human cells The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Schmidt, Nora et al. "The SARS-CoV-2 RNA–protein interactome in infected human cells." Nature Microbiology (December 2020): doi.org/10.1038/s41564-020-00846-z. © 2020 The Author(s) As Published http://dx.doi.org/10.1038/s41564-020-00846-z Publisher Springer Science and Business Media LLC Version Final published version Citable link https://hdl.handle.net/1721.1/128950 Terms of Use Creative Commons Attribution 4.0 International license Detailed Terms https://creativecommons.org/licenses/by/4.0/ ARTICLES https://doi.org/10.1038/s41564-020-00846-z The SARS-CoV-2 RNA–protein interactome in infected human cells Nora Schmidt 1,10, Caleb A. Lareau 2,10, Hasmik Keshishian3,10, Sabina Ganskih 1, Cornelius Schneider4,5, Thomas Hennig6, Randy Melanson3, Simone Werner1, Yuanjie Wei1, Matthias Zimmer1, Jens Ade 1, Luisa Kirschner6, Sebastian Zielinski 1, Lars Dölken1,6, Eric S. Lander3,7,8, Neva Caliskan1,9, Utz Fischer1,5, Jörg Vogel 1,4, Steven A. Carr3, Jochen Bodem 6 ✉ and Mathias Munschauer 1 ✉ Characterizing the interactions that SARS-CoV-2 viral RNAs make with host cell proteins during infection can improve our understanding of viral RNA functions and the host innate immune response. Using RNA antisense purification and mass spec- trometry, we identified up to 104 human proteins that directly and specifically bind to SARS-CoV-2 RNAs in infected human cells. We integrated the SARS-CoV-2 RNA interactome with changes in proteome abundance induced by viral infection and linked interactome proteins to cellular pathways relevant to SARS-CoV-2 infections. -

An Integrative Transcriptome-Wide Analysis of Amyotrophic Lateral Sclerosis for the Identification of Potential Genetic Markers and Drug Candidates
International Journal of Molecular Sciences Article An Integrative Transcriptome-Wide Analysis of Amyotrophic Lateral Sclerosis for the Identification of Potential Genetic Markers and Drug Candidates Sungmin Park 1 , Daeun Kim 2 , Jaeseung Song 2 and Jong Wha J. Joo 1,* 1 Department of Computer Engineering, Dongguk University, Seoul 04620, Korea; [email protected] 2 Department of Life Science, Dongguk University, Seoul 04620, Korea; [email protected] (D.K.); [email protected] (J.S.) * Correspondence: [email protected] Abstract: Amyotrophic lateral sclerosis (ALS) is a neurodegenerative neuromuscular disease. Al- though genome-wide association studies (GWAS) have successfully identified many variants signifi- cantly associated with ALS, it is still difficult to characterize the underlying biological mechanisms inducing ALS. In this study, we performed a transcriptome-wide association study (TWAS) to iden- tify disease-specific genes in ALS. Using the largest ALS GWAS summary statistic (n = 80,610), we identified seven novel genes using 19 tissue reference panels. We conducted a conditional analysis to verify the genes’ independence and to confirm that they are driven by genetically regulated expres- sions. Furthermore, we performed a TWAS-based enrichment analysis to highlight the association of important biological pathways, one in each of the four tissue reference panels. Finally, utilizing a connectivity map, a database of human cell expression profiles cultured with bioactive small Citation: Park, S.; Kim, D.; Song, J.; molecules, we discovered functional associations between genes and drugs to identify 15 bioactive Joo, J.W.J. An Integrative small molecules as potential drug candidates for ALS. We believe that, by integrating the largest ALS Transcriptome-Wide Analysis of GWAS summary statistic with gene expression to identify new risk loci and causal genes, our study Amyotrophic Lateral Sclerosis for the Identification of Potential Genetic provides strong candidates for molecular basis experiments in ALS. -

Inbred Mouse Strains Expression in Primary Immunocytes Across
Downloaded from http://www.jimmunol.org/ by guest on September 28, 2021 Daphne is online at: average * The Journal of Immunology published online 29 September 2014 from submission to initial decision 4 weeks from acceptance to publication Sara Mostafavi, Adriana Ortiz-Lopez, Molly A. Bogue, Kimie Hattori, Cristina Pop, Daphne Koller, Diane Mathis, Christophe Benoist, The Immunological Genome Consortium, David A. Blair, Michael L. Dustin, Susan A. Shinton, Richard R. Hardy, Tal Shay, Aviv Regev, Nadia Cohen, Patrick Brennan, Michael Brenner, Francis Kim, Tata Nageswara Rao, Amy Wagers, Tracy Heng, Jeffrey Ericson, Katherine Rothamel, Adriana Ortiz-Lopez, Diane Mathis, Christophe Benoist, Taras Kreslavsky, Anne Fletcher, Kutlu Elpek, Angelique Bellemare-Pelletier, Deepali Malhotra, Shannon Turley, Jennifer Miller, Brian Brown, Miriam Merad, Emmanuel L. Gautier, Claudia Jakubzick, Gwendalyn J. Randolph, Paul Monach, Adam J. Best, Jamie Knell, Ananda Goldrath, Vladimir Jojic, J Immunol http://www.jimmunol.org/content/early/2014/09/28/jimmun ol.1401280 Koller, David Laidlaw, Jim Collins, Roi Gazit, Derrick J. Rossi, Nidhi Malhotra, Katelyn Sylvia, Joonsoo Kang, Natalie A. Bezman, Joseph C. Sun, Gundula Min-Oo, Charlie C. Kim and Lewis L. Lanier Variation and Genetic Control of Gene Expression in Primary Immunocytes across Inbred Mouse Strains Submit online. Every submission reviewed by practicing scientists ? is published twice each month by http://jimmunol.org/subscription http://www.jimmunol.org/content/suppl/2014/09/28/jimmunol.140128 0.DCSupplemental Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2014 by The American Association of Immunologists, Inc. -
Downloaded from the Mouse Lysosome Gene Database, Mlgdb
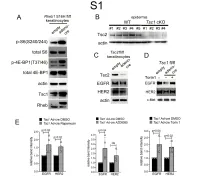
1 Supplemental Figure Legends 2 3 Supplemental Figure S1: Epidermal-specific mTORC1 gain-of-function models show 4 increased mTORC1 activation and down-regulate EGFR and HER2 protein expression in a 5 mTORC1-sensitive manner. (A) Immunoblotting of Rheb1 S16H flox/flox keratinocyte cultures 6 infected with empty or adenoviral cre recombinase for markers of mTORC1 (p-S6, p-4E-BP1) 7 activity. (B) Tsc1 cKO epidermal lysates also show decreased expression of TSC2 by 8 immunoblotting of the same experiment as in Figure 2A. (C) Immunoblotting of Tsc2 flox/flox 9 keratinocyte cultures infected with empty or adenoviral cre recombinase showing decreased EGFR 10 and HER2 protein expression. (D) Expression of EGFR and HER2 was decreased in Tsc1 cre 11 keratinocytes compared to empty controls, and up-regulated in response to Torin1 (1µM, 24 hrs), 12 by immunoblot analyses. Immunoblots are contemporaneous and parallel from the same biological 13 replicate and represent the same experiment as depicted in Figure 7B. (E) Densitometry 14 quantification of representative immunoblot experiments shown in Figures 2E and S1D (r≥3; error 15 bars represent STDEV; p-values by Student’s T-test). 16 17 18 19 20 21 22 23 Supplemental Figure S2: EGFR and HER2 transcription are unchanged with epidermal/ 24 keratinocyte Tsc1 or Rptor loss. Egfr and Her2 mRNA levels in (A) Tsc1 cKO epidermal lysates, 25 (B) Tsc1 cKO keratinocyte lysates and(C) Tsc1 cre keratinocyte lysates are minimally altered 26 compared to their respective controls. (r≥3; error bars represent STDEV; p-values by Student’s T- 27 test). -

S41467-019-13965-X.Pdf
ARTICLE https://doi.org/10.1038/s41467-019-13965-x OPEN Genome-wide CRISPR screen identifies host dependency factors for influenza A virus infection Bo Li1,2, Sara M. Clohisey 3, Bing Shao Chia1,2, Bo Wang 3, Ang Cui2,4, Thomas Eisenhaure2, Lawrence D. Schweitzer2, Paul Hoover2, Nicholas J. Parkinson3, Aharon Nachshon 5, Nikki Smith3, Tim Regan 3, David Farr3, Michael U. Gutmann6, Syed Irfan Bukhari7, Andrew Law 3, Maya Sangesland8, Irit Gat-Viks2,5, Paul Digard 3, Shobha Vasudevan7, Daniel Lingwood8, David H. Dockrell9, John G. Doench 2, J. Kenneth Baillie 3,10* & Nir Hacohen 2,11* 1234567890():,; Host dependency factors that are required for influenza A virus infection may serve as therapeutic targets as the virus is less likely to bypass them under drug-mediated selection pressure. Previous attempts to identify host factors have produced largely divergent results, with few overlapping hits across different studies. Here, we perform a genome-wide CRISPR/ Cas9 screen and devise a new approach, meta-analysis by information content (MAIC) to systematically combine our results with prior evidence for influenza host factors. MAIC out- performs other meta-analysis methods when using our CRISPR screen as validation data. We validate the host factors, WDR7, CCDC115 and TMEM199, demonstrating that these genes are essential for viral entry and regulation of V-type ATPase assembly. We also find that CMTR1, a human mRNA cap methyltransferase, is required for efficient viral cap snatching and regulation of a cell autonomous immune response, and provides synergistic protection with the influenza endonuclease inhibitor Xofluza. 1 Harvard University Virology Program, Harvfvard Medical School, Boston MA02142, USA. -

Pituitary Cell Translation and Secretory Capacities Are Enhanced Cell Autonomously by the Transcription Factor Creb3l2
ARTICLE https://doi.org/10.1038/s41467-019-11894-3 OPEN Pituitary cell translation and secretory capacities are enhanced cell autonomously by the transcription factor Creb3l2 Konstantin Khetchoumian1, Aurélio Balsalobre1, Alexandre Mayran1, Helen Christian2, Valérie Chénard3, Julie St-Pierre3 & Jacques Drouin1,3 1234567890():,; Translation is a basic cellular process and its capacity is adapted to cell function. In particular, secretory cells achieve high protein synthesis levels without triggering the protein stress response. It is unknown how and when translation capacity is increased during differentiation. Here, we show that the transcription factor Creb3l2 is a scaling factor for translation capacity in pituitary secretory cells and that it directly binds ~75% of regulatory and effector genes for translation. In parallel with this cell-autonomous mechanism, implementation of the phy- siological UPR pathway prevents triggering the protein stress response. Knockout mice for Tpit, a pituitary differentiation factor, show that Creb3l2 expression and its downstream regulatory network are dependent on Tpit. Further, Creb3l2 acts by direct targeting of translation effector genes in parallel with signaling pathways that otherwise regulate protein synthesis. Expression of Creb3l2 may be a useful means to enhance production of therapeutic proteins. 1 Laboratoire de génétique moléculaire, Institut de recherches cliniques de Montréal (IRCM), Montréal, QC H2W 1R7, Canada. 2 Departments of Physiology, Anatomy and Genetics, University of Oxford, Oxford OX2 6HS, UK. 3 Department of Biochemistry, McGill University, Rosalind and Morris Goodman Research Centre, Montréal, QC H3A 1A3, Canada. Correspondence and requests for materials should be addressed to K.K. (email: [email protected]) or to J.D.