Loss of Full-Length Dystrophin Expression Results in Major Cell- Autonomous Abnormalities in Proliferating Myoblasts
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Histone Isoform H2A1H Promotes Attainment of Distinct Physiological
Bhattacharya et al. Epigenetics & Chromatin (2017) 10:48 DOI 10.1186/s13072-017-0155-z Epigenetics & Chromatin RESEARCH Open Access Histone isoform H2A1H promotes attainment of distinct physiological states by altering chromatin dynamics Saikat Bhattacharya1,4,6, Divya Reddy1,4, Vinod Jani5†, Nikhil Gadewal3†, Sanket Shah1,4, Raja Reddy2,4, Kakoli Bose2,4, Uddhavesh Sonavane5, Rajendra Joshi5 and Sanjay Gupta1,4* Abstract Background: The distinct functional efects of the replication-dependent histone H2A isoforms have been dem- onstrated; however, the mechanistic basis of the non-redundancy remains unclear. Here, we have investigated the specifc functional contribution of the histone H2A isoform H2A1H, which difers from another isoform H2A2A3 in the identity of only three amino acids. Results: H2A1H exhibits varied expression levels in diferent normal tissues and human cancer cell lines (H2A1C in humans). It also promotes cell proliferation in a context-dependent manner when exogenously overexpressed. To uncover the molecular basis of the non-redundancy, equilibrium unfolding of recombinant H2A1H-H2B dimer was performed. We found that the M51L alteration at the H2A–H2B dimer interface decreases the temperature of melting of H2A1H-H2B by ~ 3 °C as compared to the H2A2A3-H2B dimer. This diference in the dimer stability is also refected in the chromatin dynamics as H2A1H-containing nucleosomes are more stable owing to M51L and K99R substitu- tions. Molecular dynamic simulations suggest that these substitutions increase the number of hydrogen bonds and hydrophobic interactions of H2A1H, enabling it to form more stable nucleosomes. Conclusion: We show that the M51L and K99R substitutions, besides altering the stability of histone–histone and histone–DNA complexes, have the most prominent efect on cell proliferation, suggesting that the nucleosome sta- bility is intimately linked with the physiological efects observed. -

Analysis of Trans Esnps Infers Regulatory Network Architecture
Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2014 Anat Kreimer All rights reserved ABSTRACT Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer eSNPs are genetic variants associated with transcript expression levels. The characteristics of such variants highlight their importance and present a unique opportunity for studying gene regulation. eSNPs affect most genes and their cell type specificity can shed light on different processes that are activated in each cell. They can identify functional variants by connecting SNPs that are implicated in disease to a molecular mechanism. Examining eSNPs that are associated with distal genes can provide insights regarding the inference of regulatory networks but also presents challenges due to the high statistical burden of multiple testing. Such association studies allow: simultaneous investigation of many gene expression phenotypes without assuming any prior knowledge and identification of unknown regulators of gene expression while uncovering directionality. This thesis will focus on such distal eSNPs to map regulatory interactions between different loci and expose the architecture of the regulatory network defined by such interactions. We develop novel computational approaches and apply them to genetics-genomics data in human. We go beyond pairwise interactions to define network motifs, including regulatory modules and bi-fan structures, showing them to be prevalent in real data and exposing distinct attributes of such arrangements. We project eSNP associations onto a protein-protein interaction network to expose topological properties of eSNPs and their targets and highlight different modes of distal regulation. -
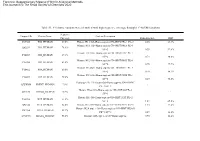
Table S1. 49 Histone Variants Were Identified with High Sequence Coverage Through LC-MS/MS Analysis Electronic Supplementary
Electronic Supplementary Material (ESI) for Analytical Methods. This journal is © The Royal Society of Chemistry 2020 Table S1. 49 histone variants were identified with high sequence coverage through LC-MS/MS analysis Sequence Uniprot IDs Protein Name Protein Description Coverage Ratio E2+/E2- RSD P07305 H10_HUMAN 67.5% Histone H1.0 OS=Homo sapiens GN=H1F0 PE=1 SV=3 4.85 23.3% Histone H1.1 OS=Homo sapiens GN=HIST1H1A PE=1 Q02539 H11_HUMAN 74.4% SV=3 0.35 92.6% Histone H1.2 OS=Homo sapiens GN=HIST1H1C PE=1 P16403 H12_HUMAN 67.1% SV=2 0.73 80.6% Histone H1.3 OS=Homo sapiens GN=HIST1H1D PE=1 P16402 H13_HUMAN 63.8% SV=2 0.75 77.7% Histone H1.4 OS=Homo sapiens GN=HIST1H1E PE=1 P10412 H14_HUMAN 69.0% SV=2 0.70 80.3% Histone H1.5 OS=Homo sapiens GN=HIST1H1B PE=1 P16401 H15_HUMAN 79.6% SV=3 0.29 98.3% Testis-specific H1 histone OS=Homo sapiens GN=H1FNT Q75WM6 H1FNT_HUMAN 7.8% \ \ PE=2 SV=3 Histone H1oo OS=Homo sapiens GN=H1FOO PE=2 Q8IZA3 H1FOO_HUMAN 5.2% \ \ SV=1 Histone H1t OS=Homo sapiens GN=HIST1H1T PE=2 P22492 H1T_HUMAN 31.4% SV=4 1.42 65.0% Q92522 H1X_HUMAN 82.6% Histone H1x OS=Homo sapiens GN=H1FX PE=1 SV=1 1.15 33.2% Histone H2A type 1 OS=Homo sapiens GN=HIST1H2AG P0C0S8 H2A1_HUMAN 99.2% PE=1 SV=2 0.57 26.8% Q96QV6 H2A1A_HUMAN 58.0% Histone H2A type 1-A OS=Homo sapiens 0.90 11.2% GN=HIST1H2AA PE=1 SV=3 Histone H2A type 1-B/E OS=Homo sapiens P04908 H2A1B_HUMAN 99.2% GN=HIST1H2AB PE=1 SV=2 0.92 30.2% Histone H2A type 1-C OS=Homo sapiens Q93077 H2A1C_HUMAN 100.0% GN=HIST1H2AC PE=1 SV=3 0.76 27.6% Histone H2A type 1-D OS=Homo sapiens P20671 -

Environmental Influences on Endothelial Gene Expression
ENDOTHELIAL CELL GENE EXPRESSION John Matthew Jeff Herbert Supervisors: Prof. Roy Bicknell and Dr. Victoria Heath PhD thesis University of Birmingham August 2012 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT Tumour angiogenesis is a vital process in the pathology of tumour development and metastasis. Targeting markers of tumour endothelium provide a means of targeted destruction of a tumours oxygen and nutrient supply via destruction of tumour vasculature, which in turn ultimately leads to beneficial consequences to patients. Although current anti -angiogenic and vascular targeting strategies help patients, more potently in combination with chemo therapy, there is still a need for more tumour endothelial marker discoveries as current treatments have cardiovascular and other side effects. For the first time, the analyses of in-vivo biotinylation of an embryonic system is performed to obtain putative vascular targets. Also for the first time, deep sequencing is applied to freshly isolated tumour and normal endothelial cells from lung, colon and bladder tissues for the identification of pan-vascular-targets. Integration of the proteomic, deep sequencing, public cDNA libraries and microarrays, delivers 5,892 putative vascular targets to the science community. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Research Article Microarray-Based Comparisons of Ion Channel Expression Patterns: Human Keratinocytes to Reprogrammed Hipscs To
Hindawi Publishing Corporation Stem Cells International Volume 2013, Article ID 784629, 25 pages http://dx.doi.org/10.1155/2013/784629 Research Article Microarray-Based Comparisons of Ion Channel Expression Patterns: Human Keratinocytes to Reprogrammed hiPSCs to Differentiated Neuronal and Cardiac Progeny Leonhard Linta,1 Marianne Stockmann,1 Qiong Lin,2 André Lechel,3 Christian Proepper,1 Tobias M. Boeckers,1 Alexander Kleger,3 and Stefan Liebau1 1 InstituteforAnatomyCellBiology,UlmUniversity,Albert-EinsteinAllee11,89081Ulm,Germany 2 Institute for Biomedical Engineering, Department of Cell Biology, RWTH Aachen, Pauwelstrasse 30, 52074 Aachen, Germany 3 Department of Internal Medicine I, Ulm University, Albert-Einstein Allee 11, 89081 Ulm, Germany Correspondence should be addressed to Alexander Kleger; [email protected] and Stefan Liebau; [email protected] Received 31 January 2013; Accepted 6 March 2013 Academic Editor: Michael Levin Copyright © 2013 Leonhard Linta et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Ion channels are involved in a large variety of cellular processes including stem cell differentiation. Numerous families of ion channels are present in the organism which can be distinguished by means of, for example, ion selectivity, gating mechanism, composition, or cell biological function. To characterize the distinct expression of this group of ion channels we have compared the mRNA expression levels of ion channel genes between human keratinocyte-derived induced pluripotent stem cells (hiPSCs) and their somatic cell source, keratinocytes from plucked human hair. This comparison revealed that 26% of the analyzed probes showed an upregulation of ion channels in hiPSCs while just 6% were downregulated. -

Genome-Wide Screen of Cell-Cycle Regulators in Normal and Tumor Cells
bioRxiv preprint doi: https://doi.org/10.1101/060350; this version posted June 23, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Genome-wide screen of cell-cycle regulators in normal and tumor cells identifies a differential response to nucleosome depletion Maria Sokolova1, Mikko Turunen1, Oliver Mortusewicz3, Teemu Kivioja1, Patrick Herr3, Anna Vähärautio1, Mikael Björklund1, Minna Taipale2, Thomas Helleday3 and Jussi Taipale1,2,* 1Genome-Scale Biology Program, P.O. Box 63, FI-00014 University of Helsinki, Finland. 2Science for Life laboratory, Department of Biosciences and Nutrition, Karolinska Institutet, SE- 141 83 Stockholm, Sweden. 3Science for Life laboratory, Division of Translational Medicine and Chemical Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 21 Stockholm, Sweden To identify cell cycle regulators that enable cancer cells to replicate DNA and divide in an unrestricted manner, we performed a parallel genome-wide RNAi screen in normal and cancer cell lines. In addition to many shared regulators, we found that tumor and normal cells are differentially sensitive to loss of the histone genes transcriptional regulator CASP8AP2. In cancer cells, loss of CASP8AP2 leads to a failure to synthesize sufficient amount of histones in the S-phase of the cell cycle, resulting in slowing of individual replication forks. Despite this, DNA replication fails to arrest, and tumor cells progress in an elongated S-phase that lasts several days, finally resulting in death of most of the affected cells. -

Stem Cells and Ion Channels
Stem Cells International Stem Cells and Ion Channels Guest Editors: Stefan Liebau, Alexander Kleger, Michael Levin, and Shan Ping Yu Stem Cells and Ion Channels Stem Cells International Stem Cells and Ion Channels Guest Editors: Stefan Liebau, Alexander Kleger, Michael Levin, and Shan Ping Yu Copyright © 2013 Hindawi Publishing Corporation. All rights reserved. This is a special issue published in “Stem Cells International.” All articles are open access articles distributed under the Creative Com- mons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Editorial Board Nadire N. Ali, UK Joseph Itskovitz-Eldor, Israel Pranela Rameshwar, USA Anthony Atala, USA Pavla Jendelova, Czech Republic Hannele T. Ruohola-Baker, USA Nissim Benvenisty, Israel Arne Jensen, Germany D. S. Sakaguchi, USA Kenneth Boheler, USA Sue Kimber, UK Paul R. Sanberg, USA Dominique Bonnet, UK Mark D. Kirk, USA Paul T. Sharpe, UK B. Bunnell, USA Gary E. Lyons, USA Ashok Shetty, USA Kevin D. Bunting, USA Athanasios Mantalaris, UK Igor Slukvin, USA Richard K. Burt, USA Pilar Martin-Duque, Spain Ann Steele, USA Gerald A. Colvin, USA EvaMezey,USA Alexander Storch, Germany Stephen Dalton, USA Karim Nayernia, UK Marc Turner, UK Leonard M. Eisenberg, USA K. Sue O’Shea, USA Su-Chun Zhang, USA Marina Emborg, USA J. Parent, USA Weian Zhao, USA Josef Fulka, Czech Republic Bruno Peault, USA Joel C. Glover, Norway Stefan Przyborski, UK Contents Stem Cells and Ion Channels, Stefan Liebau, -

Ion Channels
UC Davis UC Davis Previously Published Works Title THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels. Permalink https://escholarship.org/uc/item/1442g5hg Journal British journal of pharmacology, 176 Suppl 1(S1) ISSN 0007-1188 Authors Alexander, Stephen PH Mathie, Alistair Peters, John A et al. Publication Date 2019-12-01 DOI 10.1111/bph.14749 License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: Ion channels. British Journal of Pharmacology (2019) 176, S142–S228 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels Stephen PH Alexander1 , Alistair Mathie2 ,JohnAPeters3 , Emma L Veale2 , Jörg Striessnig4 , Eamonn Kelly5, Jane F Armstrong6 , Elena Faccenda6 ,SimonDHarding6 ,AdamJPawson6 , Joanna L Sharman6 , Christopher Southan6 , Jamie A Davies6 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 3Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 4Pharmacology and Toxicology, Institute of Pharmacy, University of Innsbruck, A-6020 Innsbruck, Austria 5School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 6Centre for Discovery Brain Science, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. -

Replicated Risk Nicotinic Cholinergic Receptor Genes for Nicotine Dependence
G C A T T A C G G C A T genes Article Replicated Risk Nicotinic Cholinergic Receptor Genes for Nicotine Dependence Lingjun Zuo 1, Rolando Garcia-Milian 2, Xiaoyun Guo 1,3,4,*, Chunlong Zhong 5,*, Yunlong Tan 6, Zhiren Wang 6, Jijun Wang 3, Xiaoping Wang 7, Longli Kang 8, Lu Lu 9,10, Xiangning Chen 11,12, Chiang-Shan R. Li 1 and Xingguang Luo 1,6,* 1 Department of Psychiatry, Yale University School of Medicine, New Haven, CT 06510, USA; [email protected] (L.Z.); [email protected] (C.-S.R.L.) 2 Curriculum & Research Support Department, Cushing/Whitney Medical Library, Yale University School of Medicine, New Haven, CT 06510, USA; [email protected] 3 Shanghai Mental Health Center, Shanghai 200030, China; [email protected] 4 Department of Cellular and Molecular Physiology, Yale University School of Medicine, New Haven, CT 06510, USA 5 Department of Neurosurgery, Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200127, China 6 Biological Psychiatry Research Center, Beijing Huilongguan Hospital, Beijing 100096, China; [email protected] (Y.T.); [email protected] (Z.W.) 7 Department of Neurology, Shanghai First People’s Hospital, Shanghai Jiao Tong University, Shanghai 200080, China; [email protected] 8 Key Laboratory for Molecular Genetic Mechanisms and Intervention Research on High Altitude Diseases of Tibet Autonomous Region, Xizang Minzu University School of Medicine, Xianyang, Shanxi 712082, China; [email protected] 9 Provincial Key Laboratory for Inflammation and Molecular Drug Target, Medical -

Rap1-Mediated Chromatin and Gene Expression Changes at Senescence
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2019 Rap1-Mediated Chromatin And Gene Expression Changes At Senescence Shufei Song University of Pennsylvania Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Biochemistry Commons, and the Cell Biology Commons Recommended Citation Song, Shufei, "Rap1-Mediated Chromatin And Gene Expression Changes At Senescence" (2019). Publicly Accessible Penn Dissertations. 3557. https://repository.upenn.edu/edissertations/3557 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/3557 For more information, please contact [email protected]. Rap1-Mediated Chromatin And Gene Expression Changes At Senescence Abstract ABSTRACT RAP1-MEDIATED CHROMATIN AND GENE EXPRESSION CHANGES AT SENESCENCE The telomeric protein Rap1 has been extensively studied for its roles as a transcriptional activator and repressor. Indeed, in both yeast and mammals, Rap1 is known to bind throughout the genome to reorganize chromatin and regulate gene transcription. Previously, our lab published evidence that Rap1 plays important roles in cellular senescence. In telomerase-deficient S. cerevisiae, Rap1 relocalizes from telomeres and subtelomeres to new Rap1 target at senescence (NRTS). This leads to two types of histone loss: Rap1 lowers global histone levels by repressing histone gene transcription and it also results in local nucleosome displacement at the promoters of the activated NRTS. Here, I examine mechanisms of site-specific histone loss by presenting evidence that Rap1 can directly interact with histone tetramers H3/H4, and map this interaction to a three-amino-acid-patch within the DNA binding domain. Functional studies are performed in vivo using a mutant form of Rap1 with weakened histone interactions, and deficient promoter clearance as well as blunted gene activation is observed, indicating that direct Rap1-H3/H4 interactions are involved in nucleosome displacement. -

A KMT2A-AFF1 Gene Regulatory Network Highlights the Role of Core Transcription Factors and Reveals the Regulatory Logic of Key Downstream Target Genes
Downloaded from genome.cshlp.org on October 7, 2021 - Published by Cold Spring Harbor Laboratory Press Research A KMT2A-AFF1 gene regulatory network highlights the role of core transcription factors and reveals the regulatory logic of key downstream target genes Joe R. Harman,1,7 Ross Thorne,1,7 Max Jamilly,2 Marta Tapia,1,8 Nicholas T. Crump,1 Siobhan Rice,1,3 Ryan Beveridge,1,4 Edward Morrissey,5 Marella F.T.R. de Bruijn,1 Irene Roberts,3,6 Anindita Roy,3,6 Tudor A. Fulga,2,9 and Thomas A. Milne1,6 1MRC Molecular Haematology Unit, MRC Weatherall Institute of Molecular Medicine, Radcliffe Department of Medicine, University of Oxford, Oxford, OX3 9DS, United Kingdom; 2MRC Weatherall Institute of Molecular Medicine, Radcliffe Department of Medicine, University of Oxford, Oxford, OX3 9DS, United Kingdom; 3MRC Molecular Haematology Unit, MRC Weatherall Institute of Molecular Medicine, Department of Paediatrics, University of Oxford, Oxford, OX3 9DS, United Kingdom; 4Virus Screening Facility, MRC Weatherall Institute of Molecular Medicine, John Radcliffe Hospital, University of Oxford, Oxford, OX3 9DS, United Kingdom; 5Center for Computational Biology, Weatherall Institute of Molecular Medicine, University of Oxford, John Radcliffe Hospital, Oxford OX3 9DS, United Kingdom; 6NIHR Oxford Biomedical Research Centre Haematology Theme, University of Oxford, Oxford, OX3 9DS, United Kingdom Regulatory interactions mediated by transcription factors (TFs) make up complex networks that control cellular behavior. Fully understanding these gene regulatory networks (GRNs) offers greater insight into the consequences of disease-causing perturbations than can be achieved by studying single TF binding events in isolation. Chromosomal translocations of the lysine methyltransferase 2A (KMT2A) gene produce KMT2A fusion proteins such as KMT2A-AFF1 (previously MLL-AF4), caus- ing poor prognosis acute lymphoblastic leukemias (ALLs) that sometimes relapse as acute myeloid leukemias (AMLs).