Assessing Equivalent and Inverse Change in Genes and Biological Pathways Between Diverse Experiments
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Functional Genomics Atlas of Synovial Fibroblasts Defining Rheumatoid Arthritis
medRxiv preprint doi: https://doi.org/10.1101/2020.12.16.20248230; this version posted December 18, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Functional genomics atlas of synovial fibroblasts defining rheumatoid arthritis heritability Xiangyu Ge1*, Mojca Frank-Bertoncelj2*, Kerstin Klein2, Amanda Mcgovern1, Tadeja Kuret2,3, Miranda Houtman2, Blaž Burja2,3, Raphael Micheroli2, Miriam Marks4, Andrew Filer5,6, Christopher D. Buckley5,6,7, Gisela Orozco1, Oliver Distler2, Andrew P Morris1, Paul Martin1, Stephen Eyre1* & Caroline Ospelt2*,# 1Versus Arthritis Centre for Genetics and Genomics, School of Biological Sciences, Faculty of Biology, Medicine and Health, The University of Manchester, Manchester, UK 2Department of Rheumatology, Center of Experimental Rheumatology, University Hospital Zurich, University of Zurich, Zurich, Switzerland 3Department of Rheumatology, University Medical Centre, Ljubljana, Slovenia 4Schulthess Klinik, Zurich, Switzerland 5Institute of Inflammation and Ageing, University of Birmingham, Birmingham, UK 6NIHR Birmingham Biomedical Research Centre, University Hospitals Birmingham NHS Foundation Trust, University of Birmingham, Birmingham, UK 7Kennedy Institute of Rheumatology, University of Oxford Roosevelt Drive Headington Oxford UK *These authors contributed equally #corresponding author: [email protected] NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. 1 medRxiv preprint doi: https://doi.org/10.1101/2020.12.16.20248230; this version posted December 18, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line -

1 Single-Cell Mrna Profiling Reveals Heterogeneous
bioRxiv preprint doi: https://doi.org/10.1101/327619; this version posted May 22, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Single-cell mRNA profiling reveals heterogeneous combinatorial expression of Hoxd genes during limb development Short title: Single-cell Hox combinations in developing limbs Authors : P. J. Fabre1,4,*, M. Leleu1, B. Mascrez2, Q. Lo Giudice4, J. Cobb3 and D. Duboule1,2,* Affiliations: 1School of Life Sciences, Ecole Polytechnique Fédérale, Lausanne, 1015 Lausanne, Switzerland. 2Department of Genetics and Evolution, University of Geneva, 1211 Geneva 4, Switzerland. 3Department of Biological Sciences, University of Calgary, Calgary, Canada. 4Department of Basic Neurosciences, University of Geneva, 1205 Geneva, Switzerland. KEYWORDS: Hox genes, digits, limb, development, enhancers, single-cell, transcriptome, differentiation, gene expression. *Corresponding authors: Pierre Fabre ([email protected]) and Denis Duboule ([email protected]) HIGHLIGHTS • Collinear expression of Hox genes is only weaved at the tissue scale • Enhancer-sharing to specific target genes is reduced at the single-cell level • Hoxd gene combinatorial expression is linked to distinct transcriptional signatures • In presumptive digits, Hoxd combinations follow a pseudotime trajectory 1 bioRxiv preprint doi: https://doi.org/10.1101/327619; this version posted May 22, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. -

Comprehensive Epigenome Characterization Reveals Diverse Transcriptional Regulation Across Human Vascular Endothelial Cells
Nakato et al. Epigenetics & Chromatin (2019) 12:77 https://doi.org/10.1186/s13072-019-0319-0 Epigenetics & Chromatin RESEARCH Open Access Comprehensive epigenome characterization reveals diverse transcriptional regulation across human vascular endothelial cells Ryuichiro Nakato1,2† , Youichiro Wada2,3*†, Ryo Nakaki4, Genta Nagae2,4, Yuki Katou5, Shuichi Tsutsumi4, Natsu Nakajima1, Hiroshi Fukuhara6, Atsushi Iguchi7, Takahide Kohro8, Yasuharu Kanki2,3, Yutaka Saito2,9,10, Mika Kobayashi3, Akashi Izumi‑Taguchi3, Naoki Osato2,4, Kenji Tatsuno4, Asuka Kamio4, Yoko Hayashi‑Takanaka2,11, Hiromi Wada3,12, Shinzo Ohta12, Masanori Aikawa13, Hiroyuki Nakajima7, Masaki Nakamura6, Rebecca C. McGee14, Kyle W. Heppner14, Tatsuo Kawakatsu15, Michiru Genno15, Hiroshi Yanase15, Haruki Kume6, Takaaki Senbonmatsu16, Yukio Homma6, Shigeyuki Nishimura16, Toutai Mitsuyama2,9, Hiroyuki Aburatani2,4, Hiroshi Kimura2,11,17* and Katsuhiko Shirahige2,5* Abstract Background: Endothelial cells (ECs) make up the innermost layer throughout the entire vasculature. Their phe‑ notypes and physiological functions are initially regulated by developmental signals and extracellular stimuli. The underlying molecular mechanisms responsible for the diverse phenotypes of ECs from diferent organs are not well understood. Results: To characterize the transcriptomic and epigenomic landscape in the vascular system, we cataloged gene expression and active histone marks in nine types of human ECs (generating 148 genome‑wide datasets) and carried out a comprehensive analysis with chromatin interaction data. We developed a robust procedure for comparative epigenome analysis that circumvents variations at the level of the individual and technical noise derived from sample preparation under various conditions. Through this approach, we identifed 3765 EC‑specifc enhancers, some of which were associated with disease‑associated genetic variations. -
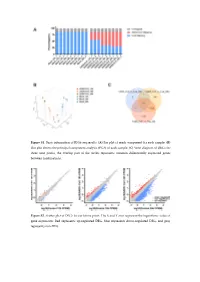
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

View Conveyed by the Global Labelling
Fabre et al. BMC Biology (2018) 16:101 https://doi.org/10.1186/s12915-018-0570-z RESEARCHARTICLE Open Access Heterogeneous combinatorial expression of Hoxd genes in single cells during limb development P. J. Fabre1,4* , M. Leleu1, B. Mascrez2, Q. Lo Giudice4, J. Cobb3 and D. Duboule1,2* Abstract Background: Global analyses of gene expression during development reveal specific transcription patterns associated with the emergence of various cell types, tissues, and organs. These heterogeneous patterns are instrumental to ensure the proper formation of the different parts of our body, as shown by the phenotypic effects generated by functional genetic approaches. However, variations at the cellular level can be observed within each structure or organ. In the developing mammalian limbs, expression of Hox genes from the HoxD cluster is differentially controlled in space and time, in cells that will pattern the digits and the forearms. While the Hoxd genes broadly share a common regulatory landscape and large-scale analyses have suggested a homogenous Hox gene transcriptional program, it has not previously been clear whether Hoxd genes are expressed together at the same levels in the same cells. Results: We report a high degree of heterogeneity in the expression of the Hoxd11 and Hoxd13 genes. We analyzed single-limb bud cell transcriptomes and show that Hox genes are expressed in specific combinations that appear to match particular cell types. In cells giving rise to digits, we find that the expression of the five relevant Hoxd genes (Hoxd9 to Hoxd13) is unbalanced, despite their control by known global enhancers. We also report that specific combinatorial expression follows a pseudo-time sequence, which is established based on the transcriptional diversity of limb progenitors. -

Functional Dissection of HOXD Cluster Genes in Regulation of Neuroblastoma Cell Proliferation and Differentiation
Functional Dissection of HOXD Cluster Genes in Regulation of Neuroblastoma Cell Proliferation and Differentiation Yunhong Zha1,4., Emily Ding3., Liqun Yang1, Ling Mao1,5, Xiangwei Wang1,6, Brian A. McCarthy1, Shuang Huang2, Han-Fei Ding1* 1 Cancer Center and Department of Pathology, Medical College of Georgia, Georgia Health Sciences University, Augusta, Georgia, United States of America, 2 Department of Biochemistry and Molecular Biology, Medical College of Georgia, Georgia Health Sciences University, Augusta, Georgia, United States of America, 3 Lakeside High School, Evans, Georgia, United States of America, 4 Department of Neurology, the First People’s Hospital of Yichang, Three Gorges University College of Medicine, Yichang, Hubei, China, 5 Department of Neurology, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, Hubei, China, 6 Department of Urology, Second Affiliated Hospital, Third Military Medical University, Chongqing, China Abstract Retinoic acid (RA) can induce growth arrest and neuronal differentiation of neuroblastoma cells and has been used in clinic for treatment of neuroblastoma. It has been reported that RA induces the expression of several HOXD genes in human neuroblastoma cell lines, but their roles in RA action are largely unknown. The HOXD cluster contains nine genes (HOXD1, HOXD3, HOXD4, and HOXD8-13) that are positioned sequentially from 39 to 59, with HOXD1 at the 39 end and HOXD13 the 59 end. Here we show that all HOXD genes are induced by RA in the human neuroblastoma BE(2)-C cells, with the genes located at the 39 end being activated generally earlier than those positioned more 59 within the cluster. Individual induction of HOXD8, HOXD9, HOXD10 or HOXD12 is sufficient to induce both growth arrest and neuronal differentiation, which is associated with downregulation of cell cycle-promoting genes and upregulation of neuronal differentiation genes. -

BMC Biology Biomed Central
BMC Biology BioMed Central Research article Open Access Classification and nomenclature of all human homeobox genes PeterWHHolland*†1, H Anne F Booth†1 and Elspeth A Bruford2 Address: 1Department of Zoology, University of Oxford, South Parks Road, Oxford, OX1 3PS, UK and 2HUGO Gene Nomenclature Committee, European Bioinformatics Institute (EMBL-EBI), Wellcome Trust Genome Campus, Hinxton, Cambridgeshire, CB10 1SA, UK Email: Peter WH Holland* - [email protected]; H Anne F Booth - [email protected]; Elspeth A Bruford - [email protected] * Corresponding author †Equal contributors Published: 26 October 2007 Received: 30 March 2007 Accepted: 26 October 2007 BMC Biology 2007, 5:47 doi:10.1186/1741-7007-5-47 This article is available from: http://www.biomedcentral.com/1741-7007/5/47 © 2007 Holland et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: The homeobox genes are a large and diverse group of genes, many of which play important roles in the embryonic development of animals. Increasingly, homeobox genes are being compared between genomes in an attempt to understand the evolution of animal development. Despite their importance, the full diversity of human homeobox genes has not previously been described. Results: We have identified all homeobox genes and pseudogenes in the euchromatic regions of the human genome, finding many unannotated, incorrectly annotated, unnamed, misnamed or misclassified genes and pseudogenes. -

Hox5 Interacts with Plzf to Restrict Shh Expression in the Developing Forelimb
Hox5 interacts with Plzf to restrict Shh expression in the developing forelimb Ben Xua,1, Steven M. Hrycaja, Daniel C. McIntyrea,2, Nicholas C. Bakera, Jun K. Takeuchib, Lucie Jeannottec, Zachary B. Gaberd, Bennett G. Novitchd, and Deneen M. Wellika,3 aDepartment of Internal Medicine, Division of Molecular Medicine and Genetics, University of Michigan, Ann Arbor, MI 48109; bCardiovascular Regeneration Institute of Molecular and Cellular Biosciences, University of Tokyo, Tokyo 113-0032, Japan; cCentre de Recherche en Cancérologie de l’Université Laval, Centre Hospitalier Universitaire de Québec, Québec, Canada G1R 2J6; and dDepartment of Neurobiology, Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095 Edited by Clifford J. Tabin, Harvard Medical School, Boston, MA, and approved October 18, 2013 (received for review August 8, 2013) To date, only the five most posterior groups of Hox genes, Hox9– expression in limb AP patterning (12, 13). Although misexpression Hox13, have demonstrated loss-of-function roles in limb pattern- of more anterior Hox genes in mice reportedly affects limb patterning ing. Individual paralog groups control proximodistal patterning of (16), no loss-of-function mutants of anterior, non–abdominal B the limb skeletal elements. Hox9 genes also initiate the onset of (AbdB)-related genes have demonstrated defects in the patterning of Hand2 expression in the posterior forelimb compartment, and col- limb skeletal elements. Moreover, no HoxB or HoxC group genes had lectively, the posterior HoxA/D genes maintain posterior Sonic been shown to play a role in forelimb development until a report by Hedgehog (Shh) expression. -

Control of Growth and Gut Maturation by Hoxd Genes and the Associated
Control of growth and gut maturation by HoxD genes PNAS PLUS and the associated lncRNA Haglr Jozsef Zakanya, Fabrice Darbellayb, Bénédicte Mascreza, Anamaria Necsuleab,1, and Denis Duboulea,b,2 aDepartment of Genetics and Evolution, University of Geneva, 1211 Geneva 4, Switzerland; and bSchool of Life Sciences, Federal Institute of Technology, Lausanne (EPFL), 1015 Lausanne, Switzerland Contributed by Denis Duboule, September 21, 2017 (sent for review July 13, 2017; reviewed by Nicoletta Bobola and Natasza Kurpios) During embryonic development, Hox genes participate in the build- side (9), emphasizing a potential link between growth retardation ing of a functional digestive system in metazoans, and genetic con- and alterations in gut development. ditions involving these genes lead to important, sometimes lethal, In this study, we tried to find a common explanatory frame- growth retardation. Recently, this phenotype was obtained after work for the various cases described above. We decided to deletion of Haglr,theHoxd antisense growth-associated long non- complete the allelic series with a few more mutant chromosomes coding RNA (lncRNA) located between Hoxd1 and Hoxd3. In this necessary to unambiguously interpret the previous observations study, we have analyzed the function of Hoxd genes in delayed relating phenotypes to particular genes. By using CRISPR/Cas9 growth trajectories by looking at several nested targeted defi- mutagenesis in embryo, we demonstrate that the growth-retardation – ciencies of the mouse HoxD cluster. Mutant pups were severely phenotype observed in HoxDDel(1 9) mice is caused by the dominant- stunted during the suckling period, but many recovered after wean- negative effect of the HOXD10 protein. -

Control of Hoxd Gene Transcription in the Mammary Bud by Hijacking a Preexisting Regulatory Landscape
Control of Hoxd gene transcription in the mammary bud by hijacking a preexisting regulatory landscape Ruben Schepa, Anamaria Necsuleab, Eddie Rodríguez-Carballoa, Isabel Guerreiroa, Guillaume Andreyb,1, Thi Hanh Nguyen Huynha, Virginie Marceta, Jozsef Zákánya, Denis Duboulea,b,2, and Leonardo Beccaria aDepartment of Genetics and Evolution, University of Geneva,1211 Geneva, Switzerland; and bSchool of Life Sciences, Ecole Polytechnique Fédérale de Lausanne, 1015 Lausanne, Switzerland Contributed by Denis Duboule, October 23, 2016 (sent for review July 31, 2016; reviewed by Marcelo A. Nobrega and Gunter P. Wagner) Vertebrate Hox genes encode transcription factors operating dur- that these two large regulatory landscapes match in their extent ing the development of multiple organs and structures. However, two topologically associating domains (TADs) [i.e., chromosome the evolutionary mechanism underlying this remarkable pleiotropy domains where specific and constitutive physical interactions are remains to be fully understood. Here, we show that Hoxd8 and privileged (23, 24)]. These two TADs are separated by a tight Hoxd9, two genes of the HoxD complex, are transcribed during mam- boundary localized within the HoxD cluster itself, isolating those mary bud (MB) development. However, unlike in other developmen- genes controlled by the telomeric TAD (T-DOM) from those tal contexts, their coexpression does not rely on the same regulatory genes responding to the centromeric regulation domain (C-DOM) mechanism. Hoxd8 is regulated by the combined activity of closely (17) (see Fig. 5A). Within the T-DOM, series of enhancers are located sequences and the most distant telomeric gene desert. On the found, which regulate groups of genes lying in the central part of other hand, Hoxd9 is controlled by an enhancer-rich region that is also the cluster, as if the regulatory sequences would contact a chro- located within the telomeric gene desert but has no impact on Hoxd8 matin pocket encompassing Hoxd8 to Hoxd11 (18). -

Supplementary File Table S1
Supplementary file Table S1 Grouping of Homeobox genes according to their main known function. Anatomical Structure Morphogenesis EN1, HOXC10, HOXC13, HOXD3, LBX1, SIX2, SIX4 Organ Morphogenesis CDX1, CDX2, HOXA11, HOXA13, ISL1, LHX1, PAX3, PDHX, PITX2, PITX3, PROX1, SIX6 Body Pattern Formation ALX3, EMX2, HHEX, HOXA11, HOXA2, HOXA4, HOXA5, HOXA6, HOXB1, HOXB5, HOXB6, HOXC5, HOXD10, HOXD8, LMX1B, PITX2 Ectoderm Development PROX1, VAX2 Endoderm Development HOXC11 Brain & Nervous System Development Brain Development ALX1, DLX2, EMX2 Nervous System Development: ARX, DLX5, DLX6, HOXD10, LBX1, LHX1, OTP, PAX3, PHOX2A, PHOX2B Skeletal Development: ALX3, ALX4, DLX3, DLX5, DLX6, EN1, HOXA11, HOXA13, HOXA2, HOXB6, HOXD10, HOXD13, MSX2 Muscle Development: BARX2, MKX, SIRT1, SIRT2, SIX1 Other Homeobox Genes Involved In BARX1, CDX4, CUX1, DLX1, EMX1, EN2, Multicellular Organismal HOXA1, HOXA7, HOXA9, HOXB13, HOXB2, Development: HOXB3, HOXB4, HOXB7, HOXB8, HOXB9, HOXC12, HOXC8, HOXC9, HOXD1, HOXD11, HOXD12, HOXD9, ISL2, LBX2, LMX1A, MEIS1, NKX3-1, OTX1, TLX1, VAX1, VSX1, VSX2 Homeobox Genes Involved In Cell ARX, EMX2, HHEX, HLX, HOPX, LBX1, LHX1, Differentiation: LMX1B, MIXL1, OTP, PHOX2A, SIRT1, VSX2 Other Genes: PHTF1, SIRT3, SIRT6, SIRT7, ZHX1, ZHX2 Homeobox genes include two subsets of genes coding for transcription factors involved in multiple functions. The clustered HOX genes are indicated in bold. Supplementary file Figure S2 5’ Spatial collinearity 3’ HOXA Chr. 7p15.3 HOXB Chr. 17q21.3 HOXC Chr. 12q13.3 HOXD Chr. 2q31 13 12 11 10 9 8 7 6 5 4 3 2 1 Paralogous HOX groups Distribution of the 39 human HOX genes in four clusters located in different chromosomal regions*. Blue indicates anterior HOX genes. Yellow, paralogy group 3 Hox genes, green and purple indicatete central HOX genes and Red the posterior HOX genes.