Pancreatic Progenitor Cells in Mice
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
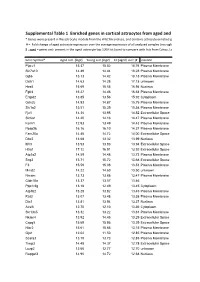
Supplemental Table 1 Enriched Genes in Cortical Astrocytes from Aged
Supplemental Table 1 Enriched genes in cortical astrocytes from aged and young-adult mice * Genes were present in the astrocyte module from the WGCNA analysis, and contains astrocyte enriched genes compared to microglia and oligodendrocytes # = Fold change of aged astrocyte expression over the average expression of all analyzed samples (microglia, astrocytes: young, old, with and without myelin contamination) $ ; aged = genes only present in the aged astrocyte top 1000 list (used to compare with lists from Cahoy, Lovatt, Doyle; see Fig. 4B), all = genes present in all astrocyte top 1000 lists Gene Symbol* Aged astr. (log2) Young astr.(log2) FC (aged/ aver.)# Location Ptprz1 15.37 15.02 18.76 Plasma Membrane Slc7a10 14.49 14.44 18.28 Plasma Membrane Gjb6 15.13 14.42 18.18 Plasma Membrane Dclk1 14.63 14.28 17.18 unknown Hes5 15.69 15.55 16.94 Nucleus Fgfr3 15.27 14.46 16.54 Plasma Membrane Entpd2 13.85 13.56 15.92 Cytoplasm Grin2c 14.93 14.87 15.75 Plasma Membrane Slc1a2 15.51 15.39 15.58 Plasma Membrane Fjx1 14.36 13.98 14.52 Extracellular Space Slc6a1 14.20 14.16 14.47 Plasma Membrane Kcnk1 12.93 13.49 14.43 Plasma Membrane Ppap2b 16.16 16.10 14.37 Plasma Membrane Fam20a 14.48 14.72 14.00 Extracellular Space Dbx2 13.68 13.32 13.99 Nucleus Itih3 13.93 13.93 13.94 Extracellular Space Htra1 17.12 16.91 13.92 Extracellular Space Atp1a2 14.59 14.48 13.73 Plasma Membrane Scg3 15.71 15.72 13.68 Extracellular Space F3 15.59 15.08 13.51 Plasma Membrane Mmd2 14.22 14.60 13.50 unknown Nrcam 13.73 13.88 13.47 Plasma Membrane Cldn10a 13.37 13.57 13.46 -

Nuclear Organization and the Epigenetic Landscape of the Mus Musculus X-Chromosome Alicia Liu University of Connecticut - Storrs, [email protected]
University of Connecticut OpenCommons@UConn Doctoral Dissertations University of Connecticut Graduate School 8-9-2019 Nuclear Organization and the Epigenetic Landscape of the Mus musculus X-Chromosome Alicia Liu University of Connecticut - Storrs, [email protected] Follow this and additional works at: https://opencommons.uconn.edu/dissertations Recommended Citation Liu, Alicia, "Nuclear Organization and the Epigenetic Landscape of the Mus musculus X-Chromosome" (2019). Doctoral Dissertations. 2273. https://opencommons.uconn.edu/dissertations/2273 Nuclear Organization and the Epigenetic Landscape of the Mus musculus X-Chromosome Alicia J. Liu, Ph.D. University of Connecticut, 2019 ABSTRACT X-linked imprinted genes have been hypothesized to contribute parent-of-origin influences on social cognition. A cluster of imprinted genes Xlr3b, Xlr4b, and Xlr4c, implicated in cognitive defects, are maternally expressed and paternally silent in the murine brain. These genes defy classic mechanisms of autosomal imprinting, suggesting a novel method of imprinted gene regulation. Using Xlr3b and Xlr4c as bait, this study uses 4C-Seq on neonatal whole brain of a 39,XO mouse model, to provide the first in-depth analysis of chromatin dynamics surrounding an imprinted locus on the X-chromosome. Significant differences in long-range contacts exist be- tween XM and XP monosomic samples. In addition, XM interaction profiles contact a greater number of genes linked to cognitive impairment, abnormality of the nervous system, and abnormality of higher mental function. This is not a pattern that is unique to the imprinted Xlr3/4 locus. Additional Alicia J. Liu - University of Connecticut - 2019 4C-Seq experiments show that other genes on the X-chromosome, implicated in intellectual disability and/or ASD, also produce more maternal contacts to other X-linked genes linked to cognitive impairment. -

Environmental Influences on Endothelial Gene Expression
ENDOTHELIAL CELL GENE EXPRESSION John Matthew Jeff Herbert Supervisors: Prof. Roy Bicknell and Dr. Victoria Heath PhD thesis University of Birmingham August 2012 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT Tumour angiogenesis is a vital process in the pathology of tumour development and metastasis. Targeting markers of tumour endothelium provide a means of targeted destruction of a tumours oxygen and nutrient supply via destruction of tumour vasculature, which in turn ultimately leads to beneficial consequences to patients. Although current anti -angiogenic and vascular targeting strategies help patients, more potently in combination with chemo therapy, there is still a need for more tumour endothelial marker discoveries as current treatments have cardiovascular and other side effects. For the first time, the analyses of in-vivo biotinylation of an embryonic system is performed to obtain putative vascular targets. Also for the first time, deep sequencing is applied to freshly isolated tumour and normal endothelial cells from lung, colon and bladder tissues for the identification of pan-vascular-targets. Integration of the proteomic, deep sequencing, public cDNA libraries and microarrays, delivers 5,892 putative vascular targets to the science community. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

140503 IPF Signatures Supplement Withfigs Thorax
Supplementary material for Heterogeneous gene expression signatures correspond to distinct lung pathologies and biomarkers of disease severity in idiopathic pulmonary fibrosis Daryle J. DePianto1*, Sanjay Chandriani1⌘*, Alexander R. Abbas1, Guiquan Jia1, Elsa N. N’Diaye1, Patrick Caplazi1, Steven E. Kauder1, Sabyasachi Biswas1, Satyajit K. Karnik1#, Connie Ha1, Zora Modrusan1, Michael A. Matthay2, Jasleen Kukreja3, Harold R. Collard2, Jackson G. Egen1, Paul J. Wolters2§, and Joseph R. Arron1§ 1Genentech Research and Early Development, South San Francisco, CA 2Department of Medicine, University of California, San Francisco, CA 3Department of Surgery, University of California, San Francisco, CA ⌘Current address: Novartis Institutes for Biomedical Research, Emeryville, CA. #Current address: Gilead Sciences, Foster City, CA. *DJD and SC contributed equally to this manuscript §PJW and JRA co-directed this project Address correspondence to Paul J. Wolters, MD University of California, San Francisco Department of Medicine Box 0111 San Francisco, CA 94143-0111 [email protected] or Joseph R. Arron, MD, PhD Genentech, Inc. MS 231C 1 DNA Way South San Francisco, CA 94080 [email protected] 1 METHODS Human lung tissue samples Tissues were obtained at UCSF from clinical samples from IPF patients at the time of biopsy or lung transplantation. All patients were seen at UCSF and the diagnosis of IPF was established through multidisciplinary review of clinical, radiological, and pathological data according to criteria established by the consensus classification of the American Thoracic Society (ATS) and European Respiratory Society (ERS), Japanese Respiratory Society (JRS), and the Latin American Thoracic Association (ALAT) (ref. 5 in main text). Non-diseased normal lung tissues were procured from lungs not used by the Northern California Transplant Donor Network. -

(WCPG): Poster Abstracts: Sunday
ARTICLE IN PRESS JID: NEUPSY [m6+; October 2, 2018;12:59 ] European Neuropsychopharmacology (2018) 000, 1–75 www.elsevier.com/locate/euroneuro Abstracts of the 26th World Congress of Psychiatric Genetics (WCPG): Poster Abstracts: Sunday Sunday, October 14, 2018 with a 2kb upstream and 1kb downstream region was consid- ered for each gene. Gene-set analysis was conducted using MAGMA, with a principal components regression model for gene-based analyses. Sex, age, the 10 first and otherwise as- Poster Session III sociated principal components were included as covariates 4:00 p.m. - 6:00 p.m. in this step. We retrieved a “Hallmark Androgen response” gene-set from MSigDB to be tested in a case-control asso- SU1 ciation study. This gene-set contains 98 curated genes in- ANDROGEN RECEPTOR SIGNALING PATHWAYS IN- volved in the response to androgen receptor signaling. Also, FLUENCE IN ATTENTION-DEFICIT/HYPERACTIVITY a list of 534 annotated genes with at least one occurrence DISORDER of potential transcription factor binding sites (TFBS) for AR was created to investigate potential gene targets related to Djenifer Kappel 1, Bruna da Silva 1, Renata B. Cupertino 1, ADHD susceptibility. Diana Müller 1, Vitor Breda 2, Stefania Pigatto Teche 2, Results: No genome-wide association was observed at the Rogério Margis 1, Luis Augusto Rohde 2, Nina Roth Mota 3, SNPs or gene level. In the gene-set analysis, we found evi- Diego L. Rovaris 1, Eugênio H. Grevet 1, Claiton Bau 1 dence that the “Hallmark Androgen response” gene-set was significantly associated with ADHD susceptibility in our sam- 1 Universidade Federal do Rio Grande do Sul ple (p = 0.039). -

WO 2012/174282 A2 20 December 2012 (20.12.2012) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2012/174282 A2 20 December 2012 (20.12.2012) P O P C T (51) International Patent Classification: David [US/US]; 13539 N . 95th Way, Scottsdale, AZ C12Q 1/68 (2006.01) 85260 (US). (21) International Application Number: (74) Agent: AKHAVAN, Ramin; Caris Science, Inc., 6655 N . PCT/US20 12/0425 19 Macarthur Blvd., Irving, TX 75039 (US). (22) International Filing Date: (81) Designated States (unless otherwise indicated, for every 14 June 2012 (14.06.2012) kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, English (25) Filing Language: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, Publication Language: English DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, KR, (30) Priority Data: KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, 61/497,895 16 June 201 1 (16.06.201 1) US MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, 61/499,138 20 June 201 1 (20.06.201 1) US OM, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SC, SD, 61/501,680 27 June 201 1 (27.06.201 1) u s SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, 61/506,019 8 July 201 1(08.07.201 1) u s TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. -

Regulation and Function of Ptf1a in the Developing Nervous System
REGULATION AND FUNCTION OF PTF1A IN THE DEVELOPING NERVOUS SYSTEM APPROVED BY SUPERVISORY COMMITTEE Jane E. Johnson, Ph.D. Raymond J. MacDonald, Ph.D. Ondine Cleaver, Ph.D. Steven L. McKnight, Ph.D. DEDICATION Work of this nature never occurs in a vacuum and would be utterly impossible without the contributions of a great many. I would like to thank my fellow lab members and collaborators, both past and present, for all of their help over the past several years, my thesis committee for their guidance and encouragement, and of course Jane for giving me every opportunity to succeed. To my friends, family, and my dear wife Maria, your love and support have made all of this possible. REGULATION AND FUNCTION OF PTF1A IN THE DEVELOPING NERVOUS SYSTEM by DAVID MILES MEREDITH DISSERTATION Presented to the Faculty of the Graduate School of Biomedical Sciences The University of Texas Southwestern Medical Center at Dallas In Partial Fulfillment of the Requirements For the Degree of DOCTOR OF PHILOSOPHY The University of Texas Southwestern Medical Center at Dallas Dallas, Texas May, 2012 REGULATION AND FUNCTION OF PTF1A IN THE DEVELOPING NERVOUS SYSTEM DAVID MILES MEREDITH The University of Texas Southwestern Medical Center at Dallas, 2012 JANE E. JOHNSON, Ph.D. Basic helix-loop-helix transcription factors serve many roles in development, including regulation of neurogenesis. Many of these factors are activated in naïve neural progenitors and function to promote neuronal differentiation and cell-type specification. Ptf1a is a basic helix-loop-helix protein that is required for proper inhibitory neuron formation in several regions of the developing nervous system, including the spinal cord, cerebellum, retina, and hypothalamus. -

Candidate Chromosome 1 Disease Susceptibility Genes for Sjogren's Syndrome Xerostomia Are Narrowed by Novel NOD.B10 Congenic Mice P
Donald and Barbara Zucker School of Medicine Journal Articles Academic Works 2014 Candidate chromosome 1 disease susceptibility genes for Sjogren's syndrome xerostomia are narrowed by novel NOD.B10 congenic mice P. K. A. Mongini Zucker School of Medicine at Hofstra/Northwell J. M. Kramer Northwell Health T. Ishikawa H. Herschman D. Esposito Follow this and additional works at: https://academicworks.medicine.hofstra.edu/articles Part of the Medical Molecular Biology Commons Recommended Citation Mongini P, Kramer J, Ishikawa T, Herschman H, Esposito D. Candidate chromosome 1 disease susceptibility genes for Sjogren's syndrome xerostomia are narrowed by novel NOD.B10 congenic mice. 2014 Jan 01; 153(1):Article 2861 [ p.]. Available from: https://academicworks.medicine.hofstra.edu/articles/2861. Free full text article. This Article is brought to you for free and open access by Donald and Barbara Zucker School of Medicine Academic Works. It has been accepted for inclusion in Journal Articles by an authorized administrator of Donald and Barbara Zucker School of Medicine Academic Works. For more information, please contact [email protected]. NIH Public Access Author Manuscript Clin Immunol. Author manuscript; available in PMC 2015 July 01. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Clin Immunol. 2014 July ; 153(1): 79–90. doi:10.1016/j.clim.2014.03.012. Candidate chromosome 1 disease susceptibility genes for Sjogren’s syndrome xerostomia are narrowed by novel NOD.B10 congenic mice Patricia K. A. Monginia and Jill M. Kramera,1 Tomo-o Ishikawab,2 and Harvey Herschmanb Donna Espositoc aThe Feinstein Institute for Medical Research North Shore-Long Island Jewish Health System 350 Community Drive Manhasset, NY 11030 bDavid Geffen School of Medicine at UCLA 341 Boyer Hall (MBI) 611 Charles E. -

The Application of Mouse and Human Embryonic Stem Cells with Transcriptomics in Alternative Developmental Toxicity Tests
The application of mouse and human embryonic stem cells with transcriptomics in alternative developmental toxicity tests A bridge from model species to man Sjors Schulpen © Copyright All rights reserved. No part of this publication may be reproduced or transmitted in any form by any means without permission of the author. ISBN: 978-94-6203-861-5 Cover and layout: Maud van Deursen, Utrecht the Netherlands Production: CPI/ Wöhrmann Print Service, Zutphen the Netherlands The application of mouse and human embryonic stem cells with transcriptomics in alternative developmental toxicity tests A bridge from model species to man De toepassing van embryonale stamcellen van muis en mens met transcriptomics in alternatieve testen voor ontwikkelingstoxiciteit Een brug van modelorganisme naar de mens (met een samenvatting in het Nederlands) Proefschrift ter verkrijging van de graad van doctor aan de Universiteit Utrecht op gezag van de rector magnificus, prof.dr. G.J. van der Zwaan, ingevolge het besluit van het college voor promoties in het openbaar te verdedigen op dinsdag 7 juli 2015 des middags te 2.30 uur door Sjors Hubertus Wilhelmina Schulpen geboren op 5 januari 1983 te Sittard Promotoren: Prof. dr. A.H. Piersma Prof. dr. M. van den Berg Dit proefschrift werd mogelijk gemaakt door financiële steun van: Institute for Risk Assessment Sciences van de Universiteit Utrecht, Laboratorium voor gezondheidsbeschermingsonderzoek van het Rijksinstituut voor Volksgezondheid en Mileu, Greiner Bio-One en Stichting Proefdiervrij. Contents Abbreviations 6 -

Notch Controls Multiple Pancreatic Cell Fate Regulators Through Direct Hes1-Mediated Repression
bioRxiv preprint doi: https://doi.org/10.1101/336305; this version posted June 1, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Notch Controls Multiple Pancreatic Cell Fate Regulators Through Direct Hes1-mediated Repression by Kristian H. de Lichtenberg1, Philip A. Seymour1, Mette C. Jørgensen1, Yung-Hae Kim1, Anne Grapin-Botton1, Mark A. Magnuson2, Nikolina Nakic3,4, Jorge Ferrer3, Palle Serup1* 1 Novo Nordisk Foundation Center for Stem Cell Biology (Danstem), University of Copenhagen, Denmark. 2 Center for Stem Cell Biology, Vanderbilt University, Tennessee, USA. 3 Section on Epigenetics and Disease, Imperial College London, UK. 4 Current Address: GSK, Stevenage, UK. *Corresponding Author: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/336305; this version posted June 1, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Notch signaling and its effector Hes1 regulate multiple cell fate choices in the developing pancreas, but few direct target genes are known. Here we use transcriptome analyses combined with chromatin immunoprecipitation with next-generation sequencing (ChIP-seq) to identify direct target genes of Hes1. ChIP-seq analysis of endogenous Hes1 in 266-6 cells, a model of multipotent pancreatic progenitor cells, revealed high-confidence peaks associated with 354 genes. Among these were genes important for tip/trunk segregation such as Ptf1a and Nkx6-1, genes involved in endocrine differentiation such as Insm1 and Dll4, and genes encoding non-pancreatic basic-Helic-Loop-Helix (bHLH) factors such as Neurog2 and Ascl1. -

Identification, by Systematic RNA Sequencing, of Novel Candidate
Laboratory Investigation (2015) 95, 1077–1088 © 2015 USCAP, Inc All rights reserved 0023-6837/15 Identification, by systematic RNA sequencing, of novel candidate biomarkers and therapeutic targets in human soft tissue tumors Anne E Sarver1, Aaron L Sarver2, Venugopal Thayanithy1 and Subbaya Subramanian1 Human sarcomas comprise a heterogeneous group of more than 50 subtypes broadly classified into two groups: bone and soft tissue sarcomas. Such heterogeneity and their relative rarity have made them challenging targets for classification, biomarker identification, and development of improved treatment strategies. In this study, we used RNA sequencing to analyze 35 primary human tissue samples representing 13 different sarcoma subtypes, along with benign schwannoma, and normal bone and muscle tissues. For each sarcoma subtype, we detected unique messenger RNA (mRNA) expression signatures, which we further subjected to bioinformatic functional analysis, upstream regulatory analysis, and microRNA (miRNA) targeting analysis. We found that, for each sarcoma subtype, significantly upregulated genes and their deduced upstream regulators included not only previously implicated known players but also novel candidates not previously reported to be associated with sarcoma. For example, the schwannoma samples were characterized by high expression of not only the known associated proteins GFAP and GAP43 but also the novel player GJB6. Further, when we integrated our expression profiles with miRNA expression data from each sarcoma subtype, we were able to deduce potential key miRNA–gene regulator relationships for each. In the Ewing’s sarcoma and fibromatosis samples, two sarcomas where miR-182-5p is significantly downregulated, multiple predicted targets were significantly upregulated, including HMCN1, NKX2-2, SCNN1G, and SOX2.