Early ACCESS Diagnosed Conditions List
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Inherited Cerebellar Ataxia in Childhood: a Pattern-Recognition Approach Using Brain MRI
REVIEW ARTICLE Inherited Cerebellar Ataxia in Childhood: A Pattern-Recognition Approach Using Brain MRI L. Vedolin, G. Gonzalez, C.F. Souza, C. Lourenc¸o, and A.J. Barkovich ABSTRACT SUMMARY: Ataxia is the principal symptom of many common neurologic diseases in childhood. Ataxias caused by dysfunction of the cerebellum occur in acute, intermittent, and progressive disorders. Most of the chronic progressive processes are secondary to degen- erative and metabolic diseases. In addition, congenital malformation of the midbrain and hindbrain can also be present, with posterior fossa symptoms related to ataxia. Brain MR imaging is the most accurate imaging technique to investigate these patients, and imaging abnormalities include size, shape, and/or signal of the brain stem and/or cerebellum. Supratentorial and cord lesions are also common. This review will discuss a pattern-recognition approach to inherited cerebellar ataxia in childhood. The purpose is to provide a comprehensive discussion that ultimately could help neuroradiologists better manage this important topic in pediatric neurology. ABBREVIATIONS: AR ϭ autosomal recessive; CAC ϭ cerebellar ataxia in childhood; 4H ϭ hypomyelination with hypogonadotropic hypogonadism and hypodon- tia; JSRD ϭ Joubert syndrome and related disorders; OPHN1 ϭ oligophrenin-1 taxia is an inability to coordinate voluntary muscle move- plastic/paraneoplastic disorders, immune-mediated/demyelinat- Aments that cannot be attributed to weakness or involuntary ing disorders, and drugs/toxins (antiepileptic medications, -

A Brazilian Cohort of Individuals with Phelan-Mcdermid Syndrome
Samogy-Costa et al. Journal of Neurodevelopmental Disorders (2019) 11:13 https://doi.org/10.1186/s11689-019-9273-1 RESEARCH Open Access A Brazilian cohort of individuals with Phelan-McDermid syndrome: genotype- phenotype correlation and identification of an atypical case Claudia Ismania Samogy-Costa1†, Elisa Varella-Branco1†, Frederico Monfardini1, Helen Ferraz2, Rodrigo Ambrósio Fock3, Ricardo Henrique Almeida Barbosa3, André Luiz Santos Pessoa4,5, Ana Beatriz Alvarez Perez3, Naila Lourenço1, Maria Vibranovski1, Ana Krepischi1, Carla Rosenberg1 and Maria Rita Passos-Bueno1* Abstract Background: Phelan-McDermid syndrome (PMS) is a rare genetic disorder characterized by global developmental delay, intellectual disability (ID), autism spectrum disorder (ASD), and mild dysmorphisms associated with several comorbidities caused by SHANK3 loss-of-function mutations. Although SHANK3 haploinsufficiency has been associated with the major neurological symptoms of PMS, it cannot explain the clinical variability seen among individuals. Our goals were to characterize a Brazilian cohort of PMS individuals, explore the genotype-phenotype correlation underlying this syndrome, and describe an atypical individual with mild phenotype. Methodology: A total of 34 PMS individuals were clinically and genetically evaluated. Data were obtained by a questionnaire answered by parents, and dysmorphic features were assessed via photographic evaluation. We analyzed 22q13.3 deletions and other potentially pathogenic copy number variants (CNVs) and also performed genotype-phenotype correlation analysis to determine whether comorbidities, speech status, and ASD correlate to deletion size. Finally, a Brazilian cohort of 829 ASD individuals and another independent cohort of 2297 ID individuals was used to determine the frequency of PMS in these disorders. Results: Our data showed that 21% (6/29) of the PMS individuals presented an additional rare CNV, which may contribute to clinical variability in PMS. -

New Microdeletion and Microduplication Syndromes: a Comprehensive Review
Genetics and Molecular Biology, 37, 1 (suppl), 210-219 (2014) Copyright © 2014, Sociedade Brasileira de Genética. Printed in Brazil www.sbg.org.br Review Article New microdeletion and microduplication syndromes: A comprehensive review Julián Nevado1,2*, Rafaella Mergener3*, María Palomares-Bralo1,2, Karen Regina Souza3, Elena Vallespín1,2, Rocío Mena1,2, Víctor Martínez-Glez1,2, María Ángeles Mori1,2, Fernando Santos1,4, Sixto García-Miñaur1,4, Fé García-Santiago1,5, Elena Mansilla1,5, Luis Fernández1,6, María Luisa de Torres1,5, Mariluce Riegel3,7$ and Pablo Lapunzina1,4,8$ 1Centro de Investigación Biomédica en Red de Enfermedades Raras, Instituto de Salud Carlos III, Madrid, Spain. 2Section of Functional and Structural Genomics, Instituto de Genética Médica y Molecular, Hospital Universitario la Paz, Madrid, Spain. 3Programa de Pós-graduação em Genética e Biologia Molecular, Universidade Federal do Rio Grande do Sul, Porto Alegre,RS, Brazil. 4Section of Clinical Genetics, Instituto de Genética Médica y Molecular, Hospital Universitario la Paz, Madrid, Spain. 5Section of Cytogenetics, Instituto de Genética Médica y Molecular, Hospital Universitario la Paz, Madrid, Spain. 6Section of Preanalytics, Instituto de Genética Médica y Molecular, Hospital Universitario la Paz, Madrid, Spain. 7Serviço de Genética Médica, Hospital de Clínicas de Porto Alegre, Porto Alegre ,RS, Brazil. 8Section of Molecular Endocrinology, Overgrowth Disordes Laboratory, Instituto de Genética Médica y Molecular, Hospital Universitario la Paz, Madrid, Spain. Abstract Several new microdeletion and microduplication syndromes are emerging as disorders that have been proven to cause multisystem pathologies frequently associated with intellectual disability (ID), multiple congenital anomalies (MCA), autistic spectrum disorders (ASD) and other phenotypic findings. In this paper, we review the “new” and emergent microdeletion and microduplication syndromes that have been described and recognized in recent years with the aim of summarizing their main characteristics and chromosomal regions involved. -

A Randomized Controlled Trial of Intranasal Oxytocin in Phelan-Mcdermid Syndrome
A Randomized Controlled Trial of Intranasal Oxytocin in Phelan-McDermid Syndrome Jarrett Fastman Icahn School of Medicine at Mount Sinai Jennifer Foss-Feig Icahn School of Medicine at Mount Sinai Yitzchak Frank Icahn School of Medicine at Mount Sinai Danielle Halpern Icahn School of Medicine at Mount Sinai Hala Harony-Nicolas Icahn School of Medicine at Mount Sinai Christina Layton Icahn School of Medicine at Mount Sinai Sven Sandin Icahn School of Medicine at Mount Sinai Paige Siper Icahn School of Medicine at Mount Sinai Lara Tang Icahn School of Medicine at Mount Sinai Pilar Trelles Icahn School of Medicine at Mount Sinai Jessica Zweifach Icahn School of Medicine at Mount Sinai Joseph D. Buxbaum Icahn School of Medicine at Mount Sinai Alexander Kolevzon ( [email protected] ) Icahn School of Medicine at Mount Sinai https://orcid.org/0000-0001-8129-2671 Research Article Keywords: Phelan-McDermid syndrome, PMS, shank3, autism spectrum disorder, ASD, oxytocin Posted Date: March 5th, 2021 Page 1/24 DOI: https://doi.org/10.21203/rs.3.rs-268151/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 2/24 Abstract Background Phelan-McDermid syndrome (PMS) is a rare neurodevelopmental disorder caused by haploinsuciency of the SHANK3 gene and characterized by global developmental delays, decits in speech and motor function, and autism spectrum disorder (ASD). Monogenic causes of ASD such as PMS are well suited to investigations with novel therapeutics, as interventions can be targeted based on established genetic etiology. While preclinical studies have demonstrated that the neuropeptide oxytocin can reverse electrophysiological, attentional, and social recognition memory decits in Shank3-decient rats, there have been no trials in individuals with PMS. -

22Q13.3 Deletion Syndrome
22q13.3 deletion syndrome Description 22q13.3 deletion syndrome, which is also known as Phelan-McDermid syndrome, is a disorder caused by the loss of a small piece of chromosome 22. The deletion occurs near the end of the chromosome at a location designated q13.3. The features of 22q13.3 deletion syndrome vary widely and involve many parts of the body. Characteristic signs and symptoms include developmental delay, moderate to profound intellectual disability, decreased muscle tone (hypotonia), and absent or delayed speech. Some people with this condition have autism or autistic-like behavior that affects communication and social interaction, such as poor eye contact, sensitivity to touch, and aggressive behaviors. They may also chew on non-food items such as clothing. Less frequently, people with this condition have seizures or lose skills they had already acquired (developmental regression). Individuals with 22q13.3 deletion syndrome tend to have a decreased sensitivity to pain. Many also have a reduced ability to sweat, which can lead to a greater risk of overheating and dehydration. Some people with this condition have episodes of frequent vomiting and nausea (cyclic vomiting) and backflow of stomach acids into the esophagus (gastroesophageal reflux). People with 22q13.3 deletion syndrome typically have distinctive facial features, including a long, narrow head; prominent ears; a pointed chin; droopy eyelids (ptosis); and deep-set eyes. Other physical features seen with this condition include large and fleshy hands and/or feet, a fusion of the second and third toes (syndactyly), and small or abnormal toenails. Some affected individuals have rapid (accelerated) growth. -
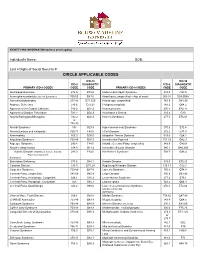
Circle Applicable Codes
IDENTIFYING INFORMATION (please print legibly) Individual’s Name: DOB: Last 4 Digits of Social Security #: CIRCLE APPLICABLE CODES ICD-10 ICD-10 ICD-9 DIAGNOSTIC ICD-9 DIAGNOSTIC PRIMARY ICD-9 CODES CODE CODE PRIMARY ICD-9 CODES CODE CODE Abetalipoproteinemia 272.5 E78.6 Hallervorden-Spatz Syndrome 333.0 G23.0 Acrocephalosyndactyly (Apert’s Syndrome) 755.55 Q87.0 Head Injury, unspecified – Age of onset: 959.01 S09.90XA Adrenaleukodystrophy 277.86 E71.529 Hemiplegia, unspecified 342.9 G81.90 Arginase Deficiency 270.6 E72.21 Holoprosencephaly 742.2 Q04.2 Agenesis of the Corpus Callosum 742.2 Q04.3 Homocystinuria 270.4 E72.11 Agenesis of Septum Pellucidum 742.2 Q04.3 Huntington’s Chorea 333.4 G10 Argyria/Pachygyria/Microgyria 742.2 Q04.3 Hurler’s Syndrome 277.5 E76.01 or 758.33 Aicardi Syndrome 333 G23.8 Hyperammonemia Syndrome 270.6 E72.4 Alcohol Embryo and Fetopathy 760.71 F84.5 I-Cell Disease 272.2 E77.0 Anencephaly 655.0 Q00.0 Idiopathic Torsion Dystonia 333.6 G24.1 Angelman Syndrome 759.89 Q93.5 Incontinentia Pigmenti 757.33 Q82.3 Asperger Syndrome 299.8 F84.5 Infantile Cerebral Palsy, unspecified 343.9 G80.9 Ataxia-Telangiectasia 334.8 G11.3 Intractable Seizure Disorder 345.1 G40.309 Autistic Disorder (Childhood Autism, Infantile 299.0 F84.0 Klinefelter’s Syndrome 758.7 Q98.4 Psychosis, Kanner’s Syndrome) Biotinidase Deficiency 277.6 D84.1 Krabbe Disease 333.0 E75.23 Canavan Disease 330.0 E75.29 Kugelberg-Welander Disease 335.11 G12.1 Carpenter Syndrome 759.89 Q87.0 Larsen’s Syndrome 755.8 Q74.8 Cerebral Palsy, unspecified 343.69 G80.9 -

2018 Etiologies by Frequencies
2018 Etiologies in Order of Frequency by Category Hereditary Syndromes and Disorders Count CHARGE Syndrome 958 Down syndrome (Trisomy 21 syndrome) 308 Usher I syndrome 252 Stickler syndrome 130 Dandy Walker syndrome 119 Cornelia de Lange 102 Goldenhar syndrome 98 Usher II syndrome 83 Wolf-Hirschhorn syndrome (Trisomy 4p) 68 Trisomy 13 (Trisomy 13-15, Patau syndrome) 60 Pierre-Robin syndrome 57 Moebius syndrome 55 Trisomy 18 (Edwards syndrome) 52 Norrie disease 38 Leber congenital amaurosis 35 Chromosome 18, Ring 18 31 Aicardi syndrome 29 Alstrom syndrome 27 Pfieffer syndrome 27 Treacher Collins syndrome 27 Waardenburg syndrome 27 Marshall syndrome 25 Refsum syndrome 21 Cri du chat syndrome (Chromosome 5p- synd) 16 Bardet-Biedl syndrome (Laurence Moon-Biedl) 15 Hurler syndrome (MPS I-H) 15 Crouzon syndrome (Craniofacial Dysotosis) 13 NF1 - Neurofibromatosis (von Recklinghausen dis) 13 Kniest Dysplasia 12 Turner syndrome 11 Usher III syndrome 10 Cockayne syndrome 9 Apert syndrome/Acrocephalosyndactyly, Type 1 8 Leigh Disease 8 Alport syndrome 6 Monosomy 10p 6 NF2 - Bilateral Acoustic Neurofibromatosis 6 Batten disease 5 Kearns-Sayre syndrome 5 Klippel-Feil sequence 5 Hereditary Syndromes and Disorders Count Prader-Willi 5 Sturge-Weber syndrome 5 Marfan syndrome 3 Hand-Schuller-Christian (Histiocytosis X) 2 Hunter Syndrome (MPS II) 2 Maroteaux-Lamy syndrome (MPS VI) 2 Morquio syndrome (MPS IV-B) 2 Optico-Cochleo-Dentate Degeneration 2 Smith-Lemli-Opitz (SLO) syndrome 2 Wildervanck syndrome 2 Herpes-Zoster (or Hunt) 1 Vogt-Koyanagi-Harada -

Megalencephaly and Macrocephaly
277 Megalencephaly and Macrocephaly KellenD.Winden,MD,PhD1 Christopher J. Yuskaitis, MD, PhD1 Annapurna Poduri, MD, MPH2 1 Department of Neurology, Boston Children’s Hospital, Boston, Address for correspondence Annapurna Poduri, Epilepsy Genetics Massachusetts Program, Division of Epilepsy and Clinical Electrophysiology, 2 Epilepsy Genetics Program, Division of Epilepsy and Clinical Department of Neurology, Fegan 9, Boston Children’s Hospital, 300 Electrophysiology, Department of Neurology, Boston Children’s Longwood Avenue, Boston, MA 02115 Hospital, Boston, Massachusetts (e-mail: [email protected]). Semin Neurol 2015;35:277–287. Abstract Megalencephaly is a developmental disorder characterized by brain overgrowth secondary to increased size and/or numbers of neurons and glia. These disorders can be divided into metabolic and developmental categories based on their molecular etiologies. Metabolic megalencephalies are mostly caused by genetic defects in cellular metabolism, whereas developmental megalencephalies have recently been shown to be caused by alterations in signaling pathways that regulate neuronal replication, growth, and migration. These disorders often lead to epilepsy, developmental disabilities, and Keywords behavioral problems; specific disorders have associations with overgrowth or abnor- ► megalencephaly malities in other tissues. The molecular underpinnings of many of these disorders are ► hemimegalencephaly now understood, providing insight into how dysregulation of critical pathways leads to ► -

Genes in Eyecare Geneseyedoc 3 W.M
Genes in Eyecare geneseyedoc 3 W.M. Lyle and T.D. Williams 15 Mar 04 This information has been gathered from several sources; however, the principal source is V. A. McKusick’s Mendelian Inheritance in Man on CD-ROM. Baltimore, Johns Hopkins University Press, 1998. Other sources include McKusick’s, Mendelian Inheritance in Man. Catalogs of Human Genes and Genetic Disorders. Baltimore. Johns Hopkins University Press 1998 (12th edition). http://www.ncbi.nlm.nih.gov/Omim See also S.P.Daiger, L.S. Sullivan, and B.J.F. Rossiter Ret Net http://www.sph.uth.tmc.edu/Retnet disease.htm/. Also E.I. Traboulsi’s, Genetic Diseases of the Eye, New York, Oxford University Press, 1998. And Genetics in Primary Eyecare and Clinical Medicine by M.R. Seashore and R.S.Wappner, Appleton and Lange 1996. M. Ridley’s book Genome published in 2000 by Perennial provides additional information. Ridley estimates that we have 60,000 to 80,000 genes. See also R.M. Henig’s book The Monk in the Garden: The Lost and Found Genius of Gregor Mendel, published by Houghton Mifflin in 2001 which tells about the Father of Genetics. The 3rd edition of F. H. Roy’s book Ocular Syndromes and Systemic Diseases published by Lippincott Williams & Wilkins in 2002 facilitates differential diagnosis. Additional information is provided in D. Pavan-Langston’s Manual of Ocular Diagnosis and Therapy (5th edition) published by Lippincott Williams & Wilkins in 2002. M.A. Foote wrote Basic Human Genetics for Medical Writers in the AMWA Journal 2002;17:7-17. A compilation such as this might suggest that one gene = one disease. -

Level Estimates of Maternal Smoking and Nicotine Replacement Therapy During Pregnancy
Using primary care data to assess population- level estimates of maternal smoking and nicotine replacement therapy during pregnancy Nafeesa Nooruddin Dhalwani BSc MSc Thesis submitted to the University of Nottingham for the degree of Doctor of Philosophy November 2014 ABSTRACT Background: Smoking in pregnancy is the most significant preventable cause of poor health outcomes for women and their babies and, therefore, is a major public health concern. In the UK there is a wide range of interventions and support for pregnant women who want to quit. One of these is nicotine replacement therapy (NRT) which has been widely available for retail purchase and prescribing to pregnant women since 2005. However, measures of NRT prescribing in pregnant women are scarce. These measures are vital to assess its usefulness in smoking cessation during pregnancy at a population level. Furthermore, evidence of NRT safety in pregnancy for the mother and child’s health so far is nebulous, with existing studies being small or using retrospectively reported exposures. Aims and Objectives: The main aim of this work was to assess population- level estimates of maternal smoking and NRT prescribing in pregnancy and the safety of NRT for both the mother and the child in the UK. Currently, the only population-level data on UK maternal smoking are from repeated cross-sectional surveys or routinely collected maternity data during pregnancy or at delivery. These obtain information at one point in time, and there are no population-level data on NRT use available. As a novel approach, therefore, this thesis used the routinely collected primary care data that are currently available for approximately 6% of the UK population and provide longitudinal/prospectively recorded information throughout pregnancy. -

Ophthalmology
Ophthalmology Information for health professionals MEDICAL GENETIC TESTING FOR OPHTHALMOLOGY Recent technologies, in particularly Next Generation Sequencing (NGS), allows fast, accurate and valuable diagnostic tests. For Ophthalmology, CGC Genetics has an extensive list of medical genetic tests with clinical integration of results by our Medical Geneticists. 1. EXOME SEQUENCING: Exome Sequencing is a very efficient strategy to study most exons of a patient’s genome, unraveling mutations associated with specific disorders or phenotypes. With this diagnostic strategy, patients can be studied with a significantly reduced turnaround time and cost. CGC Genetics has available 2 options for Exome Sequencing: • Whole Exome Sequencing (WES), which analyzes the entire exome (about 20 000 genes); • Disease Exome by CGC Genetics, which analyzes about 6 000 clinically-relevant genes. Any of these can be performed in the index case or in a Trio. 2. NGS PANELS For NGS panels, several genes associated with the same phenotype are simultaneously sequenced. These panels provide increased diagnostic capability with a significantly reduced turnaround time and cost. CGC Genetics has several NGS panels for Ophthalmology that are constantly updated (www.cgcgenetics.com). Any gene studied in exome or NGS panel can also be individually sequenced and analyzed for deletion/duplication events. 3. EXPERTISE IN MEDICAL GENETICS CGC Genetics has Medical Geneticists specialized in genetic counseling for ophthalmological diseases who may advice in choosing the most appropriate -
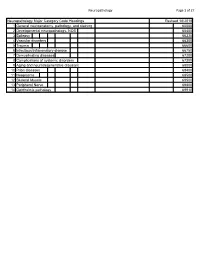
Neuropathology Category Code List
Neuropathology Page 1 of 27 Neuropathology Major Category Code Headings Revised 10/2018 1 General neuroanatomy, pathology, and staining 65000 2 Developmental neuropathology, NOS 65400 3 Epilepsy 66230 4 Vascular disorders 66300 5 Trauma 66600 6 Infectious/inflammatory disease 66750 7 Demyelinating diseases 67200 8 Complications of systemic disorders 67300 9 Aging and neurodegenerative diseases 68000 10 Prion diseases 68400 11 Neoplasms 68500 12 Skeletal Muscle 69500 13 Peripheral Nerve 69800 14 Ophthalmic pathology 69910 Neuropathology Page 2 of 27 Neuropathology 1 General neuroanatomy, pathology, and staining 65000 A Neuroanatomy, NOS 65010 1 Neocortex 65011 2 White matter 65012 3 Entorhinal cortex/hippocampus 65013 4 Deep (basal) nuclei 65014 5 Brain stem 65015 6 Cerebellum 65016 7 Spinal cord 65017 8 Pituitary 65018 9 Pineal 65019 10 Tracts 65020 11 Vascular supply 65021 12 Notochord 65022 B Cell types 65030 1 Neurons 65031 2 Astrocytes 65032 3 Oligodendroglia 65033 4 Ependyma 65034 5 Microglia and mononuclear cells 65035 6 Choroid plexus 65036 7 Meninges 65037 8 Blood vessels 65038 C Cerebrospinal fluid 65045 D Pathologic responses in neurons and axons 65050 1 Axonal degeneration/spheroid/reaction 65051 2 Central chromatolysis 65052 3 Tract degeneration 65053 4 Swollen/ballooned neurons 65054 5 Trans-synaptic neuronal degeneration 65055 6 Olivary hypertrophy 65056 7 Acute ischemic (hypoxic) cell change 65057 8 Apoptosis 65058 9 Protein aggregation 65059 10 Protein degradation/ubiquitin pathway 65060 E Neuronal nuclear inclusions 65100