A Novel Mutation in the Deoxyguanosine Kinase Gene
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

HOXB6 Homeo Box B6 HOXB5 Homeo Box B5 WNT5A Wingless-Type
5 6 6 5 . 4 2 1 1 1 2 4 6 4 3 2 9 9 7 0 5 7 5 8 6 4 0 8 2 3 1 8 3 7 1 0 0 4 0 2 5 0 8 7 5 4 1 1 0 3 6 0 4 8 3 7 4 7 6 9 6 7 1 5 0 8 1 4 1 1 7 1 0 0 4 2 0 8 1 1 1 2 5 3 5 0 7 2 6 9 1 2 1 8 3 5 2 9 8 0 6 0 9 5 1 9 9 2 1 1 6 0 2 3 0 3 6 9 1 6 5 5 7 1 1 2 1 1 7 5 4 6 6 4 1 1 2 8 4 7 1 6 2 7 7 5 4 3 2 4 3 6 9 4 1 7 1 3 4 1 2 1 3 1 1 4 7 3 1 1 1 1 5 3 2 6 1 5 1 3 5 4 5 2 3 1 1 6 1 7 3 2 5 4 3 1 6 1 5 3 1 7 6 5 1 1 1 4 6 1 6 2 7 2 1 2 e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S HOXB6 homeo box B6 HOXB5 homeo box B5 WNT5A wingless-type MMTV integration site family, member 5A WNT5A wingless-type MMTV integration site family, member 5A FKBP11 FK506 binding protein 11, 19 kDa EPOR erythropoietin receptor SLC5A6 solute carrier family 5 sodium-dependent vitamin transporter, member 6 SLC5A6 solute carrier family 5 sodium-dependent vitamin transporter, member 6 RAD52 RAD52 homolog S. -

Mutations in Mitochondrial DNA Polymerase-&Gamma
Journal of Human Genetics (2009) 54, 516–524 & 2009 The Japan Society of Human Genetics All rights reserved 1434-5161/09 $32.00 www.nature.com/jhg ORIGINAL ARTICLE Mutations in mitochondrial DNA polymerase-c promote breast tumorigenesis Keshav K Singh1, Vanniarajan Ayyasamy1, Kjerstin M Owens1, Manika Sapru Koul2 and Marija Vujcic1 Decreased mitochondrial oxidative phosphorylation (OXPHOS) is one of the hallmarks of cancer. To date, the identity of nuclear gene(s) responsible for decreased OXPHOS in tumors remains unknown. It is also unclear whether mutations in nuclear gene(s) responsible for decreased OXPHOS affect tumorigenesis. Polymerase-c (POLG) is the only DNA polymerase known to function in human mitochondria. Mutations in POLG are known to cause mitochondrial DNA (mtDNA) depletion and decreased OXPHOS, resulting in mtDNA depletion syndrome in humans. We therefore sequenced all coding exons (2–23) and flanking intron/splice junctions of POLG in breast tumors. We found that the POLG gene was mutated in 63% of breast tumors. We identified a total of 17 mutations across the POLG gene. Mutations were found in all three domains of the POLG protein, including T251I (the exonuclease domain), P587L (the linker region) and E1143G (the polymerase domain). We identified two novel mutations that include one silent (A703A) and one missense (R628Q) mutation in the evolutionarily conserved POLG linker region. In addition, we identified three novel mutations in the intronic region. Our study also revealed that mtDNA was depleted in breast tumors. Consistently, mutant POLG, when expressed in breast cancer cells, induced a depletion of mtDNA, decreased mitochondrial activity, decreased mitochondrial membrane potential, increased levels of reactive oxygen species and increased Matrigel invasion. -

Mitochondrial Hepatopathies Etiology and Genetics the Hepatocyte Mitochondrion Can Function Both As a Cause and As a Target of Liver Injury
Mitochondrial Hepatopathies Etiology and Genetics The hepatocyte mitochondrion can function both as a cause and as a target of liver injury. Most mitochondrial hepatopathies involve defects in the mitochondrial respiratory chain enzyme complexes (Figure 1). Resultant dysfunction of mitochondria yields deficient oxidative phosphorylation (OXPHOS), increased generation of reactive oxygen species (ROS), accumulation of hepatocyte lipid, impairment of other metabolic pathways and activation of both apoptotic and necrotic pathways of cellular death. Figure 1: Since the mitochondria are under dual control of nuclear DNA and mitochondrial DNA (mtDNA), mutations in genes of both classes have been associated with inherited mitochondrial myopathies, encephalopathies, and hepatopathies. Autosomal nuclear gene defects affect a variety of mitochondrial processes such as protein assembly, mtDNA synthesis and replication (e.g., deoxyguanosine kinase [dGUOK]) and DNA polymerase gamma [POLG]), and transport of nucleotides or metals. MPV17 (function unknown) and RRM2B (encoding the cytosolic p53-inducible ribonucleotide reductase small subunit) are two genes recently identified as also causing mtDNA depletion syndrome and liver failure, as has TWINKLE, TRMU, and SUCLG1. Most children with mitochondrial hepatopathies have identified or presumed mutations in these nuclear genes, rather than mtDNA genes. A classification of primary mitochondrial hepatopathies involving energy metabolism is presented in Table 1. Drug interference with mtDNA replication is now recognized as a cause of acquired mtDNA depletion that can result in liver failure, lactic acidosis, and myopathy in human immunodeficiency virus infected patients and, previously, in hepatitis B virus patients treated with nucleoside reverse transcriptase inhibitors. Current estimates suggest a minimum prevalence of all mitochondrial diseases of 11.5 cases per 100,000 individuals, or 1 in 8500 of the general population. -
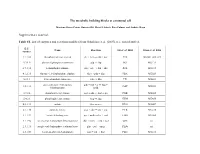
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

The Presence of Rntps Decreases the Speed of Mitochondrial DNA Replication Plos Genetics, 14(3): E1007315
http://www.diva-portal.org This is the published version of a paper published in PLoS Genetics. Citation for the original published paper (version of record): Forslund, J M., Pfeiffer, A., Stojkovič, G., Wanrooij, P H., Wanrooij, S. (2018) The presence of rNTPs decreases the speed of mitochondrial DNA replication PLoS Genetics, 14(3): e1007315 https://doi.org/10.1371/journal.pgen.1007315 Access to the published version may require subscription. N.B. When citing this work, cite the original published paper. Permanent link to this version: http://urn.kb.se/resolve?urn=urn:nbn:se:umu:diva-146802 RESEARCH ARTICLE The presence of rNTPs decreases the speed of mitochondrial DNA replication Josefin M. E. Forslund, Annika Pfeiffer☯, Gorazd Stojkovič☯, Paulina H. Wanrooij, Sjoerd Wanrooij* Department of Medical Biochemistry and Biophysics, Umeå University, Umeå, Sweden ☯ These authors contributed equally to this work. * [email protected] a1111111111 a1111111111 Abstract a1111111111 a1111111111 Ribonucleotides (rNMPs) are frequently incorporated during replication or repair by DNA a1111111111 polymerases and failure to remove them leads to instability of nuclear DNA (nDNA). Con- versely, rNMPs appear to be relatively well-tolerated in mitochondrial DNA (mtDNA), although the mechanisms behind the tolerance remain unclear. We here show that the human mitochondrial DNA polymerase gamma (Pol γ) bypasses single rNMPs with an OPEN ACCESS unprecedentedly high fidelity and efficiency. In addition, Pol γ exhibits a strikingly low fre- quency of rNMP incorporation, a property, which we find is independent of its exonuclease Citation: Forslund JME, Pfeiffer A, Stojkovič G, Wanrooij PH, Wanrooij S (2018) The presence of activity. -

Combined Gene Expression Profiling and Rnai Screening in Clear Cell Renal Cell Carcinoma Identify PLK1 and Other Therapeutic Kinase Targets
Published OnlineFirst June 3, 2011; DOI: 10.1158/0008-5472.CAN-11-0076 Cancer Therapeutics, Targets, and Chemical Biology Research Combined Gene Expression Profiling and RNAi Screening in Clear Cell Renal Cell Carcinoma Identify PLK1 and Other Therapeutic Kinase Targets Yan Ding1, Dan Huang1, Zhongfa Zhang1, Josh Smith1, David Petillo1, Brendan D. Looyenga2, Kristin Feenstra3, Jeffrey P. MacKeigan2, Kyle A. Furge1,4, and Bin T. Teh1,5 Abstract In recent years, several molecularly targeted therapies have been approved for clear cell renal cell carcinoma (ccRCC), a highly aggressive cancer. Although these therapies significantly extend overall survival, nearly all patients with advanced ccRCC eventually succumb to the disease. To identify other molecular targets, we profiled gene expression in 90 ccRCC patient specimens for which tumor grade information was available. Gene set enrichment analysis indicated that cell-cycle–related genes, in particular, Polo-like kinase 1 (PLK1), were associated with disease aggressiveness. We also carried out RNAi screening to identify kinases and phosphatases that when inhibited could prevent cell proliferation. As expected, RNAi-mediated knockdown of PLK1 and other cell-cycle kinases was sufficient to suppress ccRCC cell proliferation. The association of PLK1 in both disease aggression and in vitro growth prompted us to examine the effects of a small-molecule inhibitor of PLK1, BI 2536, in ccRCC cell lines. BI 2536 inhibited the proliferation of ccRCC cell lines at concentrations required to inhibit PLK1 kinase activity, and sustained inhibition of PLK1 by BI 2536 led to dramatic regression of ccRCC xenograft tumors in vivo. Taken together, these findings highlight PLK1 as a rational therapeutic target for ccRCC. -

Supplemental Table S1: Comparison of the Deleted Genes in the Genome-Reduced Strains
Supplemental Table S1: Comparison of the deleted genes in the genome-reduced strains Legend 1 Locus tag according to the reference genome sequence of B. subtilis 168 (NC_000964) Genes highlighted in blue have been deleted from the respective strains Genes highlighted in green have been inserted into the indicated strain, they are present in all following strains Regions highlighted in red could not be deleted as a unit Regions highlighted in orange were not deleted in the genome-reduced strains since their deletion resulted in severe growth defects Gene BSU_number 1 Function ∆6 IIG-Bs27-47-24 PG10 PS38 dnaA BSU00010 replication initiation protein dnaN BSU00020 DNA polymerase III (beta subunit), beta clamp yaaA BSU00030 unknown recF BSU00040 repair, recombination remB BSU00050 involved in the activation of biofilm matrix biosynthetic operons gyrB BSU00060 DNA-Gyrase (subunit B) gyrA BSU00070 DNA-Gyrase (subunit A) rrnO-16S- trnO-Ala- trnO-Ile- rrnO-23S- rrnO-5S yaaC BSU00080 unknown guaB BSU00090 IMP dehydrogenase dacA BSU00100 penicillin-binding protein 5*, D-alanyl-D-alanine carboxypeptidase pdxS BSU00110 pyridoxal-5'-phosphate synthase (synthase domain) pdxT BSU00120 pyridoxal-5'-phosphate synthase (glutaminase domain) serS BSU00130 seryl-tRNA-synthetase trnSL-Ser1 dck BSU00140 deoxyadenosin/deoxycytidine kinase dgk BSU00150 deoxyguanosine kinase yaaH BSU00160 general stress protein, survival of ethanol stress, SafA-dependent spore coat yaaI BSU00170 general stress protein, similar to isochorismatase yaaJ BSU00180 tRNA specific adenosine -

Dema and Faust Et Al., Suppl. Material 2020.02.03
Supplementary Materials Cyclin-dependent kinase 18 controls trafficking of aquaporin-2 and its abundance through ubiquitin ligase STUB1, which functions as an AKAP Dema Alessandro1,2¶, Dörte Faust1¶, Katina Lazarow3, Marc Wippich3, Martin Neuenschwander3, Kerstin Zühlke1, Andrea Geelhaar1, Tamara Pallien1, Eileen Hallscheidt1, Jenny Eichhorst3, Burkhard Wiesner3, Hana Černecká1, Oliver Popp1, Philipp Mertins1, Gunnar Dittmar1, Jens Peter von Kries3, Enno Klussmann1,4* ¶These authors contributed equally to this work 1Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC), Robert- Rössle-Strasse 10, 13125 Berlin, Germany 2current address: University of California, San Francisco, 513 Parnassus Avenue, CA 94122 USA 3Leibniz-Forschungsinstitut für Molekulare Pharmakologie (FMP), Robert-Rössle-Strasse 10, 13125 Berlin, Germany 4DZHK (German Centre for Cardiovascular Research), Partner Site Berlin, Oudenarder Strasse 16, 13347 Berlin, Germany *Corresponding author Enno Klussmann Max Delbrück Center for Molecular Medicine Berlin in the Helmholtz Association (MDC) Robert-Rössle-Str. 10, 13125 Berlin Germany Tel. +49-30-9406 2596 FAX +49-30-9406 2593 E-mail: [email protected] 1 Content 1. CELL-BASED SCREENING BY AUTOMATED IMMUNOFLUORESCENCE MICROSCOPY 3 1.1 Screening plates 3 1.2 Image analysis using CellProfiler 17 1.4 Identification of siRNA affecting cell viability 18 1.7 Hits 18 2. SUPPLEMENTARY TABLE S4, FIGURES S2-S4 20 2 1. Cell-based screening by automated immunofluorescence microscopy 1.1 Screening plates Table S1. Genes targeted with the Mouse Protein Kinases siRNA sub-library. Genes are sorted by plate and well. Accessions refer to National Center for Biotechnology Information (NCBI, BLA) entries. The siRNAs were arranged on three 384-well microtitre platres. -

030626 Mitochondrial Respiratory-Chain Diseases
The new england journal of medicine review article mechanisms of disease Mitochondrial Respiratory-Chain Diseases Salvatore DiMauro, M.D., and Eric A. Schon, Ph.D. From the Departments of Neurology (S.D., ore than a billion years ago, aerobic bacteria colonized E.A.S.) and Genetics and Development primordial eukaryotic cells that lacked the ability to use oxygen metabolical- (E.A.S.), Columbia University College of m Physicians and Surgeons, New York. Ad- ly. A symbiotic relationship developed and became permanent. The bacteria dress reprint requests to Dr. DiMauro at evolved into mitochondria, thus endowing the host cells with aerobic metabolism, a 4-420 College of Physicians and Surgeons, much more efficient way to produce energy than anaerobic glycolysis. Structurally, mito- 630 W. 168th St., New York, NY 10032, or at [email protected]. chondria have four compartments: the outer membrane, the inner membrane, the inter- membrane space, and the matrix (the region inside the inner membrane). They perform N Engl J Med 2003;348:2656-68. numerous tasks, such as pyruvate oxidation, the Krebs cycle, and metabolism of amino Copyright © 2003 Massachusetts Medical Society. acids, fatty acids, and steroids, but the most crucial is probably the generation of energy as adenosine triphosphate (ATP), by means of the electron-transport chain and the ox- idative-phosphorylation system (the “respiratory chain”) (Fig. 1). The respiratory chain, located in the inner mitochondrial membrane, consists of five multimeric protein complexes (Fig. 2B): reduced nicotinamide adenine dinucleotide (NADH) dehydrogenase–ubiquinone oxidoreductase (complex I, approximately 46 sub- units), succinate dehydrogenase–ubiquinone oxidoreductase (complex II, 4 subunits), ubiquinone–cytochrome c oxidoreductase (complex III, 11 subunits), cytochrome c oxi- dase (complex IV, 13 subunits), and ATP synthase (complex V, approximately 16 sub- units). -

Dntps and the Maintenance of Genome Stability
Review A Critical Balance: dNTPs and the Maintenance of Genome Stability Chen‐Chun Pai 1 and Stephen E. Kearsey 2,* 1 CRUK‐MRC Oxford Institute for Radiation Oncology, Department of Oncology, University of Oxford, ORCRB, Roosevelt Drive, Oxford OX3 7DQ, UK; chen‐[email protected] 2 Department of Zoology, University of Oxford, South Parks Road, Oxford OX1 3PS, UK * Correspondence: [email protected] Academic Editor: Eishi Noguchi Received: 6 December 2016; Accepted: 24 January 2017; Published: 31 January 2017 Abstract: A crucial factor in maintaining genome stability is establishing deoxynucleoside triphosphate (dNTP) levels within a range that is optimal for chromosomal replication. Since DNA replication is relevant to a wide range of other chromosomal activities, these may all be directly or indirectly affected when dNTP concentrations deviate from a physiologically normal range. The importance of understanding these consequences is relevant to genetic disorders that disturb dNTP levels, and strategies that inhibit dNTP synthesis in cancer chemotherapy and for treatment of other disorders. We review here how abnormal dNTP levels affect DNA replication and discuss the consequences for genome stability. Keywords: dNTP; DNA polymerase; genome instability; ribonucleotide reductase; replication fidelity; proofreading 1. Introduction Defects in genome maintenance, recognised as an enabling characteristic in cancer, contribute to the development of some neurodegenerative conditions and may be a significant factor in normal ageing. Genome instability may result from a wide range of defects affecting DNA replication, repair, checkpoint pathway function, and chromosome segregation (summarised in recent reviews [1–9]) and this review focuses on recent developments regarding one specific aspect, namely how abnormal levels of dNTPs may compromise genome stability. -

The Whole Vindaloo
FROM THE END TO THE MIDDLE: REGULATION OF TELOMERE LENGTH AND KINETOCHORE ASSEMBLY BY THE RNR INHIBITOR SML1 Amitabha Gupta Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2012 © 2012 Amitabha Gupta all rights reserved ABSTRACT FROM THE END TO THE MIDDLE: REGULATION OF TELOMERE LENGTH AND KINETOCHORE ASSEMBLY BY THE RNR INHIBITOR SML1 Amitabha Gupta Accurate DNA replication is essential for proper cellular growth and requires an adequate and balanced supply of dNTPs. In Saccharomyces cerevisiae, de novo dNTP synthesis through nucleotide reduction by the Ribonucleotide reductase (RNR) enzyme is the sole method of production. Hence, RNR activitity is highly regulated via allosteric control, transcriptional control, differential localization of subunits, and direct inhibition of the large subunit, Rnr1, by Sml1. Loss of RNR regulation results in increased mutation due to an imbalance or an absolute change in the dNTP levels in cells. In this study, I describe how mutants in dNTP regulation, including sml1∆, play a role in telomere length homeostasis. Reduction in total dNTP concentration results in a modest decrease in telomere length, while altering the ratios between the four dNTPs has a much more pronounced effect. The altered telomere lengths correlate with the relative amount of dGTP and are dependent on telomerase. At reduced levels of relative dGTP, telomerase repeatedly stalls and dissociates from telomeres, thereby leading to shorter telomeres. Conversely, with elevated relative dGTP levels, telomerase is able to processively add nucleotides and even shows low levels of repeat addition processivity.