Disease-Associated Mutation in SRSF2 Misregulates Splicing By
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Family Member in Teleost Fish Identification of a Novel IL-1 Cytokine
The Journal of Immunology Identification of a Novel IL-1 Cytokine Family Member in Teleost Fish1 Tiehui Wang,* Steve Bird,* Antonis Koussounadis,† Jason W. Holland,* Allison Carrington,* Jun Zou,* and Christopher J. Secombes2* A novel IL-1 family member (nIL-1F) has been discovered in fish, adding a further member to this cytokine family. The unique gene organization of nIL-1F, together with its location in the genome and low homology to known family members, suggests that this molecule is not homologous to known IL-1F. Nevertheless, it contains a predicted C-terminal -trefoil structure, an IL-1F signature region within the final exon, a potential IL-1 converting enzyme cut site, and its expression level is clearly increased following infection, or stimulation of macrophages with LPS or IL-1. A thrombin cut site is also present and may have functional relevance. The C-terminal recombinant protein antagonized the effects of rainbow trout rIL-1 on inflammatory gene expression in a trout macrophage cell line, suggesting it is an IL-1 antagonist. Modeling studies confirmed that nIL-1F has the potential to bind to the trout IL-1RI receptor protein, and may be a novel IL-1 receptor antagonist. The Journal of Immunology, 2009, 183: 962–974. he IL-1 family (IL-1F)3 of cytokines is characterized by of transcription factors such as NF-B and MAPK-regulated tran- their common secondary structure of an all- fold, the scription factors, leading to IL-1F-regulated gene transcription in T -trefoil, which they have in common with another cyto- the target cells (1). -

The Dawn of Next Generation DNA Sequencing in Myelodysplastic Syndromes- Experience from Pakistan
The Dawn Of Next Generation DNA Sequencing In Myelodysplastic Syndromes- Experience From Pakistan. Nida Anwar ( [email protected] ) National Institute of Blood Diseases and Bone Marrow Transplantation Faheem Ahmed Memon Liaquat University of Medical & Health Sciences Saba Shahid National Institute of Blood Diseases and Bone Marrow Transplantation Muhammad Shakeel Jamil-ur-Rahman Center for Genome Research, University of Karachi Muhammad Irfan Jamil-ur-Rahman Center for Genome Research, University of Karachi Aisha Arshad National Institute of Blood Diseases and Bone Marrow Transplantation Arshi Naz Liaquat University of Medical & Health Sciences Ikram Din Ujjan Liaquat University of Medical & Health Sciences Tahir Shamsi National Institute of Blood Diseases and Bone Marrow Transplantation Research Article Keywords: Myelodysplastic Syndromes, Next generation sequencing, Gene analysis, Mutation, Pakistan. Posted Date: June 8th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-571430/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/16 Abstract Background: Myelodysplastic syndromes (MDS) are clonal disorders of hematopoietic stem cells exhibiting ineffective hematopoiesis and tendency for transformation into acute myeloid leukemia (AML). The available karyotyping and uorescent in situ hybridization provide limited information on molecular abnormalities for diagnosis/prognosis of MDS. Next generation DNA sequencing (NGS), providing deep insights into molecular mechanisms being involved in pathophysiology, was employed to study MDS in Pakistani cohort. Patients and Methods: It was a descriptive cross-sectional study carried out at National institute of blood diseases and bone marrow transplant from 2016 to 2019. Total of 22 cases of MDS were included. Complete blood counts, bone marrow assessment and cytogenetic analysis was done. -

Minor Intron Retention Drives Clonal Hematopoietic Disorders and Diverse Cancer Predisposition
ARTICLES https://doi.org/10.1038/s41588-021-00828-9 Minor intron retention drives clonal hematopoietic disorders and diverse cancer predisposition Daichi Inoue1,2,12, Jacob T. Polaski 3,12, Justin Taylor 4,12, Pau Castel 5, Sisi Chen2, Susumu Kobayashi1,6, Simon J. Hogg2, Yasutaka Hayashi1, Jose Mario Bello Pineda3,7,8, Ettaib El Marabti2, Caroline Erickson2, Katherine Knorr2, Miki Fukumoto1, Hiromi Yamazaki1, Atsushi Tanaka1,9, Chie Fukui1, Sydney X. Lu2, Benjamin H. Durham 2, Bo Liu2, Eric Wang2, Sanjoy Mehta10, Daniel Zakheim10, Ralph Garippa10, Alex Penson2, Guo-Liang Chew 11, Frank McCormick 5, Robert K. Bradley 3,7 ✉ and Omar Abdel-Wahab 2 ✉ Most eukaryotes harbor two distinct pre-mRNA splicing machineries: the major spliceosome, which removes >99% of introns, and the minor spliceosome, which removes rare, evolutionarily conserved introns. Although hypothesized to serve important regulatory functions, physiologic roles of the minor spliceosome are not well understood. For example, the minor spliceosome component ZRSR2 is subject to recurrent, leukemia-associated mutations, yet functional connections among minor introns, hematopoiesis and cancers are unclear. Here, we identify that impaired minor intron excision via ZRSR2 loss enhances hemato- poietic stem cell self-renewal. CRISPR screens mimicking nonsense-mediated decay of minor intron-containing mRNA species converged on LZTR1, a regulator of RAS-related GTPases. LZTR1 minor intron retention was also discovered in the RASopathy Noonan syndrome, due to intronic mutations disrupting splicing and diverse solid tumors. These data uncover minor intron rec- ognition as a regulator of hematopoiesis, noncoding mutations within minor introns as potential cancer drivers and links among ZRSR2 mutations, LZTR1 regulation and leukemias. -

HOXB6 Homeo Box B6 HOXB5 Homeo Box B5 WNT5A Wingless-Type
5 6 6 5 . 4 2 1 1 1 2 4 6 4 3 2 9 9 7 0 5 7 5 8 6 4 0 8 2 3 1 8 3 7 1 0 0 4 0 2 5 0 8 7 5 4 1 1 0 3 6 0 4 8 3 7 4 7 6 9 6 7 1 5 0 8 1 4 1 1 7 1 0 0 4 2 0 8 1 1 1 2 5 3 5 0 7 2 6 9 1 2 1 8 3 5 2 9 8 0 6 0 9 5 1 9 9 2 1 1 6 0 2 3 0 3 6 9 1 6 5 5 7 1 1 2 1 1 7 5 4 6 6 4 1 1 2 8 4 7 1 6 2 7 7 5 4 3 2 4 3 6 9 4 1 7 1 3 4 1 2 1 3 1 1 4 7 3 1 1 1 1 5 3 2 6 1 5 1 3 5 4 5 2 3 1 1 6 1 7 3 2 5 4 3 1 6 1 5 3 1 7 6 5 1 1 1 4 6 1 6 2 7 2 1 2 e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a a S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S S HOXB6 homeo box B6 HOXB5 homeo box B5 WNT5A wingless-type MMTV integration site family, member 5A WNT5A wingless-type MMTV integration site family, member 5A FKBP11 FK506 binding protein 11, 19 kDa EPOR erythropoietin receptor SLC5A6 solute carrier family 5 sodium-dependent vitamin transporter, member 6 SLC5A6 solute carrier family 5 sodium-dependent vitamin transporter, member 6 RAD52 RAD52 homolog S. -

Mutations in Mitochondrial DNA Polymerase-&Gamma
Journal of Human Genetics (2009) 54, 516–524 & 2009 The Japan Society of Human Genetics All rights reserved 1434-5161/09 $32.00 www.nature.com/jhg ORIGINAL ARTICLE Mutations in mitochondrial DNA polymerase-c promote breast tumorigenesis Keshav K Singh1, Vanniarajan Ayyasamy1, Kjerstin M Owens1, Manika Sapru Koul2 and Marija Vujcic1 Decreased mitochondrial oxidative phosphorylation (OXPHOS) is one of the hallmarks of cancer. To date, the identity of nuclear gene(s) responsible for decreased OXPHOS in tumors remains unknown. It is also unclear whether mutations in nuclear gene(s) responsible for decreased OXPHOS affect tumorigenesis. Polymerase-c (POLG) is the only DNA polymerase known to function in human mitochondria. Mutations in POLG are known to cause mitochondrial DNA (mtDNA) depletion and decreased OXPHOS, resulting in mtDNA depletion syndrome in humans. We therefore sequenced all coding exons (2–23) and flanking intron/splice junctions of POLG in breast tumors. We found that the POLG gene was mutated in 63% of breast tumors. We identified a total of 17 mutations across the POLG gene. Mutations were found in all three domains of the POLG protein, including T251I (the exonuclease domain), P587L (the linker region) and E1143G (the polymerase domain). We identified two novel mutations that include one silent (A703A) and one missense (R628Q) mutation in the evolutionarily conserved POLG linker region. In addition, we identified three novel mutations in the intronic region. Our study also revealed that mtDNA was depleted in breast tumors. Consistently, mutant POLG, when expressed in breast cancer cells, induced a depletion of mtDNA, decreased mitochondrial activity, decreased mitochondrial membrane potential, increased levels of reactive oxygen species and increased Matrigel invasion. -

Mitochondrial Hepatopathies Etiology and Genetics the Hepatocyte Mitochondrion Can Function Both As a Cause and As a Target of Liver Injury
Mitochondrial Hepatopathies Etiology and Genetics The hepatocyte mitochondrion can function both as a cause and as a target of liver injury. Most mitochondrial hepatopathies involve defects in the mitochondrial respiratory chain enzyme complexes (Figure 1). Resultant dysfunction of mitochondria yields deficient oxidative phosphorylation (OXPHOS), increased generation of reactive oxygen species (ROS), accumulation of hepatocyte lipid, impairment of other metabolic pathways and activation of both apoptotic and necrotic pathways of cellular death. Figure 1: Since the mitochondria are under dual control of nuclear DNA and mitochondrial DNA (mtDNA), mutations in genes of both classes have been associated with inherited mitochondrial myopathies, encephalopathies, and hepatopathies. Autosomal nuclear gene defects affect a variety of mitochondrial processes such as protein assembly, mtDNA synthesis and replication (e.g., deoxyguanosine kinase [dGUOK]) and DNA polymerase gamma [POLG]), and transport of nucleotides or metals. MPV17 (function unknown) and RRM2B (encoding the cytosolic p53-inducible ribonucleotide reductase small subunit) are two genes recently identified as also causing mtDNA depletion syndrome and liver failure, as has TWINKLE, TRMU, and SUCLG1. Most children with mitochondrial hepatopathies have identified or presumed mutations in these nuclear genes, rather than mtDNA genes. A classification of primary mitochondrial hepatopathies involving energy metabolism is presented in Table 1. Drug interference with mtDNA replication is now recognized as a cause of acquired mtDNA depletion that can result in liver failure, lactic acidosis, and myopathy in human immunodeficiency virus infected patients and, previously, in hepatitis B virus patients treated with nucleoside reverse transcriptase inhibitors. Current estimates suggest a minimum prevalence of all mitochondrial diseases of 11.5 cases per 100,000 individuals, or 1 in 8500 of the general population. -

Mutations in Splicing Factor Genes in Myeloid Malignancies: Significance and Impact on Clinical Features
cancers Review Mutations in Splicing Factor Genes in Myeloid Malignancies: Significance and Impact on Clinical Features Valeria Visconte 1, Megan O. Nakashima 2 and Heesun J. Rogers 2,* 1 Department of Translational Hematology and Oncology Research, Taussig Cancer Institute, Cleveland Clinic, Cleveland, OH 44195, USA; [email protected] 2 Department of Laboratory Medicine, Cleveland Clinic, Cleveland, OH 44195, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-216-445-2719 Received: 15 October 2019; Accepted: 19 November 2019; Published: 22 November 2019 Abstract: Components of the pre-messenger RNA splicing machinery are frequently mutated in myeloid malignancies. Mutations in LUC7L2, PRPF8, SF3B1, SRSF2, U2AF1, and ZRSR2 genes occur at various frequencies ranging between 40% and 85% in different subtypes of myelodysplastic syndrome (MDS) and 5% and 10% of acute myeloid leukemia (AML) and myeloproliferative neoplasms (MPNs). In some instances, splicing factor (SF) mutations have provided diagnostic utility and information on clinical outcomes as exemplified by SF3B1 mutations associated with increased ring sideroblasts (RS) in MDS-RS or MDS/MPN-RS with thrombocytosis. SF3B1 mutations are associated with better survival outcomes, while SRSF2 mutations are associated with a shorter survival time and increased AML progression, and U2AF1 mutations with a lower remission rate and shorter survival time. Beside the presence of mutations, transcriptomics technologies have shown that one third of genes in AML patients are differentially expressed, leading to altered transcript stability, interruption of protein function, and improper translation compared to those of healthy individuals. The detection of SF mutations demonstrates the importance of splicing abnormalities in the hematopoiesis of MDS and AML patients given the fact that abnormal splicing regulates the function of several transcriptional factors (PU.1, RUNX1, etc.) crucial in hematopoietic function. -
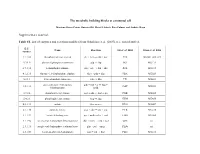
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

Based on Network Pharmacology and RNA Sequencing Techniques to Explore the Molecular Mechanism of Huatan Jiangzhuo Decoction for Treating Hyperlipidemia
Hindawi Evidence-Based Complementary and Alternative Medicine Volume 2021, Article ID 9863714, 16 pages https://doi.org/10.1155/2021/9863714 Research Article Based on Network Pharmacology and RNA Sequencing Techniques to Explore the Molecular Mechanism of Huatan Jiangzhuo Decoction for Treating Hyperlipidemia XiaowenZhou ,1 ZhenqianYan ,1 YaxinWang ,1 QiRen ,1 XiaoqiLiu ,2 GeFang ,3 Bin Wang ,4 and Xiantao Li 1 1Laboratory of TCM Syndrome Essence and Objectification, School of Basic Medical Sciences, Guangzhou University of Chinese Medicine, Guangzhou 510006, China 2Guangzhou Sagene Tech Co., Ltd., Guangzhou 510006, China 3College of Traditional Chinese Medicine, Hunan University of Chinese Medicine, Changsha 410208, China 4Shenzhen Traditional Chinese Medicine Hospital, Shenzhen 518000, China Correspondence should be addressed to Xiantao Li; [email protected] Received 22 May 2020; Revised 12 March 2021; Accepted 18 March 2021; Published 12 April 2021 Academic Editor: Jianbo Wan Copyright © 2021 Xiaowen Zhou et al. +is is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Background. Hyperlipidemia, due to the practice of unhealthy lifestyles of modern people, has been a disturbance to a large portion of population worldwide. Recently, several scholars have turned their attention to Chinese medicine (CM) to seek out a lipid-lowering approach with high efficiency and low toxicity. +is study aimed to explore the mechanism of Huatan Jiangzhuo decoction (HTJZD, a prescription of CM) in the treatment of hyperlipidemia and to determine the major regulation pathways and potential key targets involved in the treatment process. -

A Novel Mutation in the Deoxyguanosine Kinase Gene
BRIEFCOMMUNICATIONS decreasedinnervation.13 Thelatterfindingaffordedan ProgressiveLossofCardiac opportunitytodeterminewhetherthelossofcardiac SympatheticInnervationin sympathetic innervation in Parkinson’s disease progressesovertimeand,ifso,withwhattiming,pat- Parkinson’sDisease tern,andconsistencyacrosspatients.Thisreportde- scribestheresultsofretestingsuchpatientswith Sheng-TingLi,MD,PhD,RaghuveerDendi,MD, 6-[18F]fluorodopaminepositronemissiontomography CourtneyHolmes,CMT, scanningafteranaverageof2years. andDavidS.Goldstein,MD,PhD PatientsandMethods Thisstudyaddressedwhethercardiacsympathetic denervationprogressesovertimeinParkinson’sdis- ThestudyprotocolwasapprovedbytheIntramuralResearch ease.In9patientswithoutorthostatichypotension, BoardoftheNationalInstituteofNeurologicalDisorders 6-[18F]fluorodopaminepositronemissiontomography andStroke.Eachpatientgaveinformed,writtenconsent. scanningwasrepeatedafterameanof2yearsfromthe firstscan.6-[18F]fluorodopamine-derivedradioactivity waslessinthesecondscanthaninthefirstscan,by Patients 31%intheleftventricularfreewalland16%inthe Thoracicpositronemissiontomographyscanningwasper- septum.InParkinson’sdisease,lossofcardiacsympa- formedafterintravenousinjectionof6-[18F]fluoro- theticdenervationprogressesinapatternoflosssug- dopaminein9patientswithParkinson’sdisease(age: gestingadying-backmechanism. mean,60years;SEM,3years).Noneofthepatientshad orthostatichypotension,whichwasdefinedasadecreasein AnnNeurol2002;52:220–223 systolicbloodpressuregreaterthan20mmHganddecrease indiastolicpressuregreaterthan5mmHgbetweenthesu- -

The Presence of Rntps Decreases the Speed of Mitochondrial DNA Replication Plos Genetics, 14(3): E1007315
http://www.diva-portal.org This is the published version of a paper published in PLoS Genetics. Citation for the original published paper (version of record): Forslund, J M., Pfeiffer, A., Stojkovič, G., Wanrooij, P H., Wanrooij, S. (2018) The presence of rNTPs decreases the speed of mitochondrial DNA replication PLoS Genetics, 14(3): e1007315 https://doi.org/10.1371/journal.pgen.1007315 Access to the published version may require subscription. N.B. When citing this work, cite the original published paper. Permanent link to this version: http://urn.kb.se/resolve?urn=urn:nbn:se:umu:diva-146802 RESEARCH ARTICLE The presence of rNTPs decreases the speed of mitochondrial DNA replication Josefin M. E. Forslund, Annika Pfeiffer☯, Gorazd Stojkovič☯, Paulina H. Wanrooij, Sjoerd Wanrooij* Department of Medical Biochemistry and Biophysics, Umeå University, Umeå, Sweden ☯ These authors contributed equally to this work. * [email protected] a1111111111 a1111111111 Abstract a1111111111 a1111111111 Ribonucleotides (rNMPs) are frequently incorporated during replication or repair by DNA a1111111111 polymerases and failure to remove them leads to instability of nuclear DNA (nDNA). Con- versely, rNMPs appear to be relatively well-tolerated in mitochondrial DNA (mtDNA), although the mechanisms behind the tolerance remain unclear. We here show that the human mitochondrial DNA polymerase gamma (Pol γ) bypasses single rNMPs with an OPEN ACCESS unprecedentedly high fidelity and efficiency. In addition, Pol γ exhibits a strikingly low fre- quency of rNMP incorporation, a property, which we find is independent of its exonuclease Citation: Forslund JME, Pfeiffer A, Stojkovič G, Wanrooij PH, Wanrooij S (2018) The presence of activity.