Ribosomal Protein L3 Polyclonal Antibody Product Information
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Reprogramming of Trna Modifications Controls the Oxidative Stress Response by Codon-Biased Translation of Proteins
Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Chan, Clement T.Y. et al. “Reprogramming of tRNA Modifications Controls the Oxidative Stress Response by Codon-biased Translation of Proteins.” Nature Communications 3 (2012): 937. As Published http://dx.doi.org/10.1038/ncomms1938 Publisher Nature Publishing Group Version Author's final manuscript Citable link http://hdl.handle.net/1721.1/76775 Terms of Use Article is made available in accordance with the publisher's policy and may be subject to US copyright law. Please refer to the publisher's site for terms of use. Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins Clement T.Y. Chan,1,2 Yan Ling Joy Pang,1 Wenjun Deng,1 I. Ramesh Babu,1 Madhu Dyavaiah,3 Thomas J. Begley3 and Peter C. Dedon1,4* 1Department of Biological Engineering, 2Department of Chemistry and 4Center for Environmental Health Sciences, Massachusetts Institute of Technology, Cambridge, MA 02139; 3College of Nanoscale Science and Engineering, University at Albany, SUNY, Albany, NY 12203 * Corresponding author: PCD, Department of Biological Engineering, NE47-277, Massachusetts Institute of Technology, 77 Massachusetts Avenue, Cambridge, MA 02139; tel 617-253-8017; fax 617-324-7554; email [email protected] 2 ABSTRACT Selective translation of survival proteins is an important facet of cellular stress response. We recently demonstrated that this translational control involves a stress-specific reprogramming of modified ribonucleosides in tRNA. -

Allele-Specific Expression of Ribosomal Protein Genes in Interspecific Hybrid Catfish
Allele-specific Expression of Ribosomal Protein Genes in Interspecific Hybrid Catfish by Ailu Chen A dissertation submitted to the Graduate Faculty of Auburn University in partial fulfillment of the requirements for the Degree of Doctor of Philosophy Auburn, Alabama August 1, 2015 Keywords: catfish, interspecific hybrids, allele-specific expression, ribosomal protein Copyright 2015 by Ailu Chen Approved by Zhanjiang Liu, Chair, Professor, School of Fisheries, Aquaculture and Aquatic Sciences Nannan Liu, Professor, Entomology and Plant Pathology Eric Peatman, Associate Professor, School of Fisheries, Aquaculture and Aquatic Sciences Aaron M. Rashotte, Associate Professor, Biological Sciences Abstract Interspecific hybridization results in a vast reservoir of allelic variations, which may potentially contribute to phenotypical enhancement in the hybrids. Whether the allelic variations are related to the downstream phenotypic differences of interspecific hybrid is still an open question. The recently developed genome-wide allele-specific approaches that harness high- throughput sequencing technology allow direct quantification of allelic variations and gene expression patterns. In this work, I investigated allele-specific expression (ASE) pattern using RNA-Seq datasets generated from interspecific catfish hybrids. The objective of the study is to determine the ASE genes and pathways in which they are involved. Specifically, my study investigated ASE-SNPs, ASE-genes, parent-of-origins of ASE allele and how ASE would possibly contribute to heterosis. My data showed that ASE was operating in the interspecific catfish system. Of the 66,251 and 177,841 SNPs identified from the datasets of the liver and gill, 5,420 (8.2%) and 13,390 (7.5%) SNPs were identified as significant ASE-SNPs, respectively. -

The VE-Cadherin/Amotl2 Mechanosensory Pathway Suppresses Aortic In�Ammation and the Formation of Abdominal Aortic Aneurysms
The VE-cadherin/AmotL2 mechanosensory pathway suppresses aortic inammation and the formation of abdominal aortic aneurysms Yuanyuan Zhang Karolinska Institute Evelyn Hutterer Karolinska Institute Sara Hultin Karolinska Institute Otto Bergman Karolinska Institute Maria Forteza Karolinska Institute Zorana Andonovic Karolinska Institute Daniel Ketelhuth Karolinska University Hospital, Stockholm, Sweden Joy Roy Karolinska Institute Per Eriksson Karolinska Institute Lars Holmgren ( [email protected] ) Karolinska Institute Article Keywords: arterial endothelial cells (ECs), vascular disease, abdominal aortic aneurysms Posted Date: June 15th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-600069/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License The VE-cadherin/AmotL2 mechanosensory pathway suppresses aortic inflammation and the formation of abdominal aortic aneurysms Yuanyuan Zhang1, Evelyn Hutterer1, Sara Hultin1, Otto Bergman2, Maria J. Forteza2, Zorana Andonovic1, Daniel F.J. Ketelhuth2,3, Joy Roy4, Per Eriksson2 and Lars Holmgren1*. 1Department of Oncology-Pathology, BioClinicum, Karolinska Institutet, Stockholm, Sweden. 2Department of Medicine Solna, BioClinicum, Karolinska Institutet, Karolinska University Hospital, Stockholm, Sweden. 3Department of Cardiovascular and Renal Research, Institutet of Molecular Medicine, Univ. of Southern Denmark, Odense, Denmark 4Department of Molecular Medicine and Surgery, Karolinska Institutet, Karolinska University Hospital, Stockholm, -

Inflammation Leads to Distinct Populations of Extracellular Vesicles from Microglia Yiyi Yang1* , Antonio Boza-Serrano1, Christopher J
Yang et al. Journal of Neuroinflammation (2018) 15:168 https://doi.org/10.1186/s12974-018-1204-7 RESEARCH Open Access Inflammation leads to distinct populations of extracellular vesicles from microglia Yiyi Yang1* , Antonio Boza-Serrano1, Christopher J. R. Dunning2, Bettina Hjelm Clausen3,4, Kate Lykke Lambertsen3,4,5 and Tomas Deierborg1* Abstract Background: Activated microglia play an essential role in inflammatory responses elicited in the central nervous system (CNS). Microglia-derived extracellular vesicles (EVs) are suggested to be involved in propagation of inflammatory signals and in the modulation of cell-to-cell communication. However, there is a lack of knowledge on the regulation of EVs and how this in turn facilitates the communication between cells in the brain. Here, we characterized microglial EVs under inflammatory conditions and investigated the effects of inflammation on the EV size, quantity, and protein content. Methods: We have utilized western blot, nanoparticle tracking analysis (NTA), and mass spectrometry to characterize EVs and examine the alterations of secreted EVs from a microglial cell line (BV2) following lipopolysaccharide (LPS) and tumor necrosis factor (TNF) inhibitor (etanercept) treatments, or either alone. The inflammatory responses were measured with multiplex cytokine ELISA and western blot. We also subjected TNF knockout mice to experimental stroke (permanent middle cerebral artery occlusion) and validated the effect of TNF inhibition on EV release. Results: Our analysis of EVs originating from activated BV2 microglia revealed a significant increase in the intravesicular levels of TNF and interleukin (IL)-6. We also observed that the number of EVs released was reduced both in vitro and in vivo when inflammation was inhibited via the TNF pathway. -
Supplementary Table 1. the List of Proteins with at Least 2 Unique
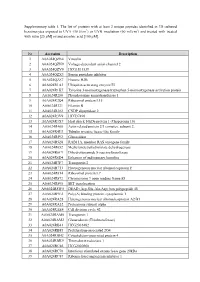
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31 -

Inhibition of the MID1 Protein Complex
Matthes et al. Cell Death Discovery (2018) 4:4 DOI 10.1038/s41420-017-0003-8 Cell Death Discovery ARTICLE Open Access Inhibition of the MID1 protein complex: a novel approach targeting APP protein synthesis Frank Matthes1,MoritzM.Hettich1, Judith Schilling1, Diana Flores-Dominguez1, Nelli Blank1, Thomas Wiglenda2, Alexander Buntru2,HannaWolf1, Stephanie Weber1,InaVorberg 1, Alina Dagane2, Gunnar Dittmar2,3,ErichWanker2, Dan Ehninger1 and Sybille Krauss1 Abstract Alzheimer’s disease (AD) is characterized by two neuropathological hallmarks: senile plaques, which are composed of amyloid-β (Aβ) peptides, and neurofibrillary tangles, which are composed of hyperphosphorylated tau protein. Aβ peptides are derived from sequential proteolytic cleavage of the amyloid precursor protein (APP). In this study, we identified a so far unknown mode of regulation of APP protein synthesis involving the MID1 protein complex: MID1 binds to and regulates the translation of APP mRNA. The underlying mode of action of MID1 involves the mTOR pathway. Thus, inhibition of the MID1 complex reduces the APP protein level in cultures of primary neurons. Based on this, we used one compound that we discovered previously to interfere with the MID1 complex, metformin, for in vivo experiments. Indeed, long-term treatment with metformin decreased APP protein expression levels and consequently Aβ in an AD mouse model. Importantly, we have initiated the metformin treatment late in life, at a time-point where mice were in an already progressed state of the disease, and could observe an improved behavioral phenotype. These 1234567890 1234567890 findings together with our previous observation, showing that inhibition of the MID1 complex by metformin also decreases tau phosphorylation, make the MID1 complex a particularly interesting drug target for treating AD. -

PNAS 07-04849-SI Table 3. 6-18-2007
Table 5. β-arrestin 2-interacting proteins under nonstimulated (-) condition IPI accession Swiss- Number of Prot Gene symbol Protein name experiments accession number number detected Signal transduction Adaptor proteins IPI00027355 P32121 ARRB2 β-arrestin 2 6 IPI00293857 P49407 ARRB1 β-arrestin 1 6 IPI00021353 P10523 SAG S-arrestin (Retinal S-antigen) (48 kDa protein) (S-AG) (Rod photoreceptor arrestin) 2 IPI00003917 P36575 ARR3 X-arrestin (Arrestin-C) (Cone arrestin) (cArr) (Retinal cone arrestin-3) (C-arrestin) 2 IPI00216318 P31946 YWHAB 14-3-3 β/α (14-3-3 protein beta/alpha) (Protein kinase C inhibitor protein 1) (KCIP-1) (Protein 1054) 2 IPI00220642 P61981 YWHAG 14-3-3 γ (14-3-3 protein gamma) (Protein kinase C inhibitor protein 1) (KCIP-1) 2 IPI00018146 P27348 YWHAQ 14-3-3 θ (14-3-3 protein tau) (14-3-3 protein theta) (14-3-3 protein T-cell) (HS1 protein) 2 IPI00216319 Q04917 YWHAH 14-3-3 η (14-3-3 protein eta) (Protein AS1) 2 IPI00000816 P62258 YWHAE 14-3-3 ε (14-3-3 protein epsilon) (14-3-3E) 2 Protein kinases IPI00027251 Q15208 STK38 STK38 (Serine/threonine-protein kinase 38) (NDR1 protein kinase) (Nuclear Dbf2-related kinase 1) 4 SCY1-like 2 (SCY1-like 2 protein) (coated vesicle-associated kinase of 104 kDa) (Eukaryotic protein IPI00396218 Q6P3W7 SCYL2 2 kinase family protein) (CDNA FLJ10074 fis, clone HEMBA1001744, weakly similar to SCY1 IPI00013835 Q13574 DGKZ DGK ζ (Diacylglycerol kinase zeta) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta) 2 DGK ε (Diacylglycerol kinase epsilon) (Diglyceride kinase epsilon) (DGK-epsilon) -

(HBV) Infection of the Liver Ahmed Mohamed Abdel-B Diab Purdue University
Purdue University Purdue e-Pubs Open Access Dissertations Theses and Dissertations January 2016 The oler of PLK1 in Hepatitis B Virus (HBV) infection of the liver Ahmed Mohamed Abdel-B Diab Purdue University Follow this and additional works at: https://docs.lib.purdue.edu/open_access_dissertations Recommended Citation Diab, Ahmed Mohamed Abdel-B, "The or le of PLK1 in Hepatitis B Virus (HBV) infection of the liver" (2016). Open Access Dissertations. 1214. https://docs.lib.purdue.edu/open_access_dissertations/1214 This document has been made available through Purdue e-Pubs, a service of the Purdue University Libraries. Please contact [email protected] for additional information. Graduate School Form 30 Updated 12/26/2015 PURDUE UNIVERSITY GRADUATE SCHOOL Thesis/Dissertation Acceptance This is to certify that the thesis/dissertation prepared By Ahmed Mohamed Abdel-B Diab Entitled The role of PLK1 in Hepatitis B Virus (HBV) Infection of the Liver For the degree of Doctor of Philosophy Is approved by the final examining committee: Ourania Andrisani Andy W. Tao Co-chair Fabien Zoulim Co-chair Robert Geahlen Xiaoqi Liu To the best of my knowledge and as understood by the student in the Thesis/Dissertation Agreement, Publication Delay, and Certification Disclaimer (Graduate School Form 32), this thesis/dissertation adheres to the provisions of Purdue University’s “Policy of Integrity in Research” and the use of copyright material. Approved by Major Professor(s): Ourania Andrisani Laurie Jaeger 10/6/2016 Approved by: Head of the Departmental Graduate Program Date i THE ROLE OF PLK1 IN HEPATITIS B VIRUS (HBV) INFECTION OF THE LIVER A Dissertation Submitted to the Faculty of Purdue University by Ahmed M Diab In Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy December 2016 Purdue University West Lafayette, Indiana ii ACKNOWLEDGEMENTS I would like to begin by extending my gratitude to my committee members Robert Geahlen, Andy W. -

Supplementary Table S1. Protein Expression Changes in the Functionally Annotated Categories Within Cluster from Figure 8A
Supplementary Table S1. Protein expression changes in the functionally annotated categories within cluster from Figure 8A. Acute phase response signaling Human equivalent Zq (log2 FC) FDR Accession No. Ac. No. No. Protein 120 1 4 7 120 1 4 7 peptides Q13233 F1SLK9 PIG Uncharacterized protein MAP3K1 0.09 0.00 0.00 0.55 1 - I3LQR9 PIG Fibrinogen alpha chain 0.89 0.23 0.03 0.11 11 - F1RX36 PIG Fibrinogen alpha chain 0.97 0.09 0.00 0.09 4 P02675 I3L651 PIG Uncharacterized protein 0.91 0.24 0.05 0.23 27 - Q8SPS7 PIG Haptoglobin 0.11 0.02 0.00 0.36 14 P45985 F1SS51 PIG Uncharacterized protein (Fragment) MAP2K4 0.26 0.06 0.09 0.33 1 - P00747 HUMAN Plasminogen 0.93 0.00 0.02 0.67 1 - P18648 PIG Apolipoprotein A-I 0.36 0.19 0.64 0.86 34 P02679 F1RX35 PIG Uncharacterized protein LOC100627396 0.99 0.40 0.02 0.11 23 P04196 F1SFI5 PIG Uncharacterized protein HRG 0.44 0.04 0.55 0.82 12 - P27485 PIG Retinol-binding protein 4 0.34 0.30 0.55 0.81 2 - F1S0J2 PIG Uncharacterized protein C4BPA 0.58 0.17 0.15 0.65 4 - P08835 PIG Serum albumin 0.04 0.09 0.39 0.75 27 - P06867 PIG Plasminogen 0.94 0.17 0.08 0.82 13 - O19063 PIG Serum amyloid P-component 0.93 0.16 0.09 0.85 6 - O19062 PIG C-reactive protein 0.93 0.01 0.00 0.78 7 - P00450 HUMAN Ceruloplasmin 0.32 0.61 0.36 0.41 1 - F1SB81 PIG Plasminogen 0.99 0.17 0.14 0.52 2 - Q19AZ8 PIG Prothrombin 0.36 0.08 0.54 0.90 6 - I3L818 PIG Uncharacterized protein (Fragment) SERPINF2 0.91 0.59 0.12 0.65 4 P05546 F1RKY2 PIG Uncharacterized protein SERPIND1 0.38 0.44 0.72 0.86 9 P02749 I3LGN5 PIG Uncharacterized protein -

Protein List
Protein Accession Protein Id Protein Name P11171 41 Protein 4. -

Deficiency of MMP17/MT4-MMP Proteolytic Activity Predisposes to Aortic Aneurysm in Mice
Deficiency of MMP17/MT4-MMP Proteolytic Activity Predisposes to Aortic Aneurysm in Mice Mara Martín-Alonso1, Ana B. García-Redondo2, Dongchuan Guo3, Emilio Camafeita4, Fernando Martínez5, Arántzazu Alfranca1, Nerea Méndez-Barbero1, Ángela Pollán1, Cristina Sánchez-Camacho6, David T. Denhardt7, Motoharu Seiki8, Jesús Vázquez1,4, Mercedes Salaices2, Juan Miguel Redondo1, Dianna Milewicz3 and Alicia G. Arroyo1 1Vascular Biology and Inflammation, Centro Nacional de Investigaciones Cardiovasculares Carlos III (CNIC), 28029 Madrid Spain; 2Department of Pharmacology/Nephrology, Faculty of Medicine, Universidad Autónoma de Madrid, 28029 Madrid, Spain; 3Department of Internal Medicine, University of Texas Health Science Center at Houston, Houston, 77030 Texas, USA;4Proteomics Unit; 5Bioinformatics Unit, Centro Nacional de Investigaciones Cardiovasculares Carlos III (CNIC), 28029 Madrid Spain; 6Universidad Europea de Madrid, Villaviciosa de Odón, 28670 Madrid, Spain; 7Department of Cell Biology and Neuroscience, Rutgers University, Piscataway, New Jersey, USA, and; 8Division of Cancer Cell Research, Institute of Medical Science, University of Tokyo, Minato-ku, 108-8639 Tokyo, Japan. Running title: MMP17 in Aortic Pathology Subject codes: [162] Smooth muscle proliferation and differentiation [130] Animal models of human disease Address correspondence to: Dr. Alicia G. Arroyo Matrix Metalloproteinases Lab Centro Nacional de Investigaciones Cardiovasculares (CNIC) Melchor Fernández Almagro 3 28029 Madrid Spain Tel 34 91 4531241 ext 1159 Fax 34 914531265 [email protected] In April 2015, the average time from submission to first decision for all original research papers submitted to Circulation Research was 13.84 days. DOI: 10.1161/CIRCRESAHA.117.305108 1 Downloaded from http://circres.ahajournals.org/ at CNIC -F.C.Nac.Inv.Cardiovasculares Carlos III on May 25, 2016 ABSTRACT Rationale: Aortic dissection or rupture resulting from aneurysm causes 1-2% of deaths in developed countries. -
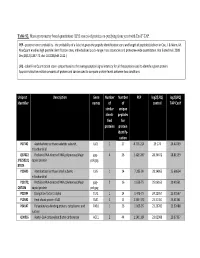
Table S2. Mass Spectrometry Based Quantitation (LFQ Scores) of Proteins Co-Purifying from Yeast with Exoy-TAP
Table S2. Mass spectrometry based quantitation (LFQ scores) of proteins co-purifying from yeast with ExoY-TAP. PEP - posterior error probability - the probability of a false hit given the peptide identification score and length of peptides (defined in Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide quantitation. Nat Biotechnol. 2008 Dec;26(12):1367-72. doi: 10.1038/nbt.1511.) LFQ - Label Free Quantitation score - proportional to the average peptide signal intensity for all the peptides used to identify a given protein. Approximates the relative amounts of protein and can be used to compare protein levels between two conditions. Uniprot Description Gene Number Number PEP log2(LFQ) log2(LFQ) identifier names of of control TAP-ExoY similar unique identi- peptides fied for proteins protein identify- cation P07342 Acetolactate synthase catalytic subunit, ILV2 1 27 8.77E-214 29.174 26.40759 mitochondrial Q87022 Probable RNA-directed RNA polymerase;Major gag- 4 26 1.42E-267 28.34672 28.81559 ;P32503;Q capsid protein pol;gag 87024 P25605 Acetolactate synthase small subunit, ILV6 1 14 7.28E-99 28.14865 25.68654 mitochondrial P23172; Probable RNA-directed RNA polymerase;Major gag- 216 1.63E-75 25.06163 20.40561 Q87026 capsid protein pol;gag P02994 Elongation factor 1-alpha TEF1 1 14 1.47E-75 24.22957 25.30567 P10591 Heat shock protein SSA1 SSA1 2 8 2.54E-173 23.31451 25.41361 P04147 Polyadenylate-binding protein, cytoplasmic and PAB1 126 1.06E-95 23.29782 23.92488 nuclear Q00955 Acetyl-CoA carboxylase;Biotin carboxylase ACC1 2 44 1.54E-164 23.02989 20.97337 P0C2I6;P0 Transposon Ty1-LR3 Gag-Pol polyprotein;Capsid TY1B- 57 3 1.88E-88 22.48395 23.64713 C2I9;P0C2 protein;Ty1 protease;Integrase; Reverse LR3;TY1B- J1;P47098 transcriptase/ribonuclease H (Rev.