This Thesis Has Been Submitted in Fulfilment of the Requirements for a Postgraduate Degree (E.G
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

An Order Estimation Based Approach to Identify Response Genes
AN ORDER ESTIMATION BASED APPROACH TO IDENTIFY RESPONSE GENES FOR MICRO ARRAY TIME COURSE DATA A Thesis Presented to The Faculty of Graduate Studies of The University of Guelph by ZHIHENG LU In partial fulfilment of requirements for the degree of Doctor of Philosophy September, 2008 © Zhiheng Lu, 2008 Library and Bibliotheque et 1*1 Archives Canada Archives Canada Published Heritage Direction du Branch Patrimoine de I'edition 395 Wellington Street 395, rue Wellington Ottawa ON K1A0N4 Ottawa ON K1A0N4 Canada Canada Your file Votre reference ISBN: 978-0-494-47605-5 Our file Notre reference ISBN: 978-0-494-47605-5 NOTICE: AVIS: The author has granted a non L'auteur a accorde une licence non exclusive exclusive license allowing Library permettant a la Bibliotheque et Archives and Archives Canada to reproduce, Canada de reproduire, publier, archiver, publish, archive, preserve, conserve, sauvegarder, conserver, transmettre au public communicate to the public by par telecommunication ou par Plntemet, prefer, telecommunication or on the Internet, distribuer et vendre des theses partout dans loan, distribute and sell theses le monde, a des fins commerciales ou autres, worldwide, for commercial or non sur support microforme, papier, electronique commercial purposes, in microform, et/ou autres formats. paper, electronic and/or any other formats. The author retains copyright L'auteur conserve la propriete du droit d'auteur ownership and moral rights in et des droits moraux qui protege cette these. this thesis. Neither the thesis Ni la these ni des extraits substantiels de nor substantial extracts from it celle-ci ne doivent etre imprimes ou autrement may be printed or otherwise reproduits sans son autorisation. -
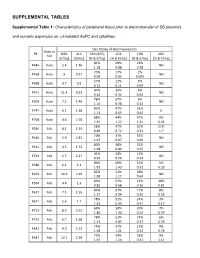
Supplemental Tables
SUPPLEMENTAL TABLES Supplemental Table 1: Characteristics of peripheral blood prior to electrotransfer of SB plasmids and numeric expansion on γ-irradiated AaPC and cytokines Day 0 (Day of electroporation) Auto or P# WBC ALC CD3 (ATC) CD4 CD8 ABC Allo [K/mL] [K/mL] [% & K/mL] [% & K/mL] [% & K/mL] [% & K/mL] 81% 60% 19% P446 Auto 5.4 1.46 ND 1.18 0.88 0.28 23% 17% 2% P458 Auto 9 0.27 ND 0.06 0.05 0.005 17% 12% 5% P468 Auto 4.7 0.9 ND 0.15 0.11 0.04 47% 35% 5% P471 Auto 11.4 0.93 ND 0.44 0.32 0.04 78% 67% 9% P509 Auto 7.2 1.46 ND 1.14 0.98 0.13 82% 47% 32% P747 Auto 6.1 1.38 0 1.13 0.65 0.44 88% 44% 47% 0% P708 Auto 4.6 2.78 2.45 1.22 1.31 0.05 58% 47% 20% 21% P396 Allo 8.1 1.54 0.89 0.72 0.31 1.7 70% 31% 32% P410 Allo 5.9 2.81 ND 1.97 0.87 0.90 80% 48% 32% P411 Allo 4.5 1.72 ND 1.38 0.83 0.55 41% 24% 15% P513 Allo 6.7 2.27 ND 0.93 0.54 0.34 86% 68% 15% 5% P580 Allo 6.1 2.1 1.81 1.43 0.32 0.29 82% 51% 28% P459 Allo 10.6 2.29 ND 1.88 1.17 0.64 64% 52% 12% 18% P564 Allo 4.4 1.3 0.83 0.68 0.16 0.81 82% 61% 17% 8% P617 Allo 7.1 1.55 1.27 0.94 0.26 0.58 78% 53% 24% 3% P671 Allo 5.4 1.7 1.33 0.90 0.41 0.17 69% 50% 20% 7% P723 Allo 8.6 2.61 1.80 1.30 0.52 0.59 78% 52% 22% 6% P732 Allo 6.7 1.68 1.31 0.87 0.37 0.39 74% 57% 15% 9% P641 Allo 9.0 2.13 1.58 1.21 0.32 0.78 73% 54% 18% 9% P647 Allo 12.1 2.29 1.67 1.24 0.41 1.11 75% 59% 11% P716 Allo 3.0 1.83 0 1.37 1.08 0.20 87% 73% 12% 1% P718 Allo 6.2 1.99 1.73 1.45 0.24 0.07 83% 69% 13% 6% P771 Allo 13.1 3.23 2.68 2.23 0.42 0.79 73% 42% 27% 6% P783 Allo 8.1 1.54 1.12 0.65 0.42 0.52 WBC = white blood -

Integrated Bioinformatic Analysis of RNA Binding Proteins in Hepatocellular Carcinoma
www.aging-us.com AGING 2021, Vol. 13, No. 2 Research Paper Integrated bioinformatic analysis of RNA binding proteins in hepatocellular carcinoma Ling Wang1,*, Zhen Zhang2,3,*, Yuan Li1, Yanyan Wan1, Baocai Xing1,& 1Key Laboratory of Carcinogenesis and Translational Research, Ministry of Education, Hepatopancreatobiliary Surgery Department I, Peking University Cancer Hospital and Institute, Beijing 100142, China 2Department of Gastroenterological Surgery, Peking University People’s Hospital, Beijing 100044, China 3Laboratory of Surgical Oncology, Beijing Key Laboratory of Colorectal Cancer Diagnosis and Treatment Research, Peking University People’s Hospital, Beijing 100044, China *Equal contribution Correspondence to: Baocai Xing; email: [email protected] Keywords: RNA binding protein, hepatocellular carcinoma, biomarker, transcriptomics, proteomics Received: June 8, 2020 Accepted: November 3, 2020 Published: December 19, 2020 Copyright: © 2020 Wang et al. This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT RNA binding proteins (RBPs) are aberrantly expressed in a tissue-specific manner across many tumors. These proteins, which play a vital role in post-transcriptional gene regulation, are involved in RNA splicing, maturation, transport, stability, degradation, and translation. We set out to establish an accurate risk score model based on RBPs to estimate prognosis in hepatocellular carcinoma (HCC). RNA-sequencing data, proteomic data and corresponding clinical information were acquired from the Cancer Genome Atlas database and the Clinical Proteomic Tumor Analysis Consortium database respectively. We identified 406 differentially expressed RBPs between HCC tumor and normal tissues at the transcriptional and protein level. -

P53 — 30 Yreviewsears On
FOCUS ON P53 — 30 YREVIEWSEARS ON p53 — a Jack of all trades but master of none Melissa R. Junttila and Gerard I. Evan Abstract | Cancers are rare because their evolution is actively restrained by a range of tumour suppressors. Of these p53 seems unusually crucial as either it or its attendant upstream or downstream pathways are inactivated in virtually all cancers. p53 is an evolutionarily ancient coordinator of metazoan stress responses. Its role in tumour suppression is likely to be a relatively recent adaptation, which is only necessary when large, long-lived organisms acquired the sufficient size and somatic regenerative capacity to necessitate specific mechanisms to reign in rogue proliferating cells. However, such evolutionary reappropriation of this venerable transcription factor entails compromises that restrict its efficacy as a tumour suppressor. Cnidarians–bilaterians Cancer is a genetic pathology that arises in the adult tissues questions are important for several reasons. Knowing Cinidarians comprise an animal of long-lived organisms, such as vertebrates, whose tis- how p53 discriminates between normal and tumour cells phylum of ~9,000 radially sues retain a substantial regenerative capacity throughout might point to attributes of cancer cells that qualitatively symmetrical, mostly marine life and, consequently, whose somatic cells accumulate distinguish them from normal somatic cells and could organisms. Most other animals are bilaterally symmetrical mutations. Occasionally, such mutations corrupt the be used as tumour-specific targets. Knowing what sig- and are classed as bilateria. regulatory mechanisms that suppress untoward somatic nals drive selection for loss of p53 function in cancers The cnidarians and bilaterians cell proliferation, survival and migration, resulting in the would help us understand what constrains and dictates last shared a common ancestor progressive outgrowth of a somatic clone. -

Predicting Clinical Response to Treatment with a Soluble Tnf-Antagonist Or Tnf, Or a Tnf Receptor Agonist
(19) TZZ _ __T (11) EP 2 192 197 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 02.06.2010 Bulletin 2010/22 C12Q 1/68 (2006.01) (21) Application number: 08170119.5 (22) Date of filing: 27.11.2008 (84) Designated Contracting States: (72) Inventor: The designation of the inventor has not AT BE BG CH CY CZ DE DK EE ES FI FR GB GR yet been filed HR HU IE IS IT LI LT LU LV MC MT NL NO PL PT RO SE SI SK TR (74) Representative: Habets, Winand Designated Extension States: Life Science Patents AL BA MK RS PO Box 5096 6130 PB Sittard (NL) (71) Applicant: Vereniging voor Christelijk Hoger Onderwijs, Wetenschappelijk Onderzoek en Patiëntenzorg 1081 HV Amsterdam (NL) (54) Predicting clinical response to treatment with a soluble tnf-antagonist or tnf, or a tnf receptor agonist (57) The invention relates to methods for predicting a clinical response to a therapy with a soluble TNF antagonist, TNF or a TNF receptor agonist and a kit for use in said methods. EP 2 192 197 A1 Printed by Jouve, 75001 PARIS (FR) EP 2 192 197 A1 Description [0001] The invention relates to methods for predicting a clinical response to a treatment with a soluble TNF antagonist, with TNF or a TNF receptor agonist using expression levels of genes of the Type I INF pathway and a kit for use in said 5 methods. In another aspect, the invention relates to a method for evaluating a pharmacological effect of a treatment with a soluble TNF antagonist, TNF or a TNF receptor agonist. -

Rrna Types in Zebrafish Development
UvA-DARE (Digital Academic Repository) Distinct maternal and somatic rRNA types in zebrafish development Locati, M. Publication date 2017 Document Version Final published version License Other Link to publication Citation for published version (APA): Locati, M. (2017). Distinct maternal and somatic rRNA types in zebrafish development. General rights It is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), other than for strictly personal, individual use, unless the work is under an open content license (like Creative Commons). Disclaimer/Complaints regulations If you believe that digital publication of certain material infringes any of your rights or (privacy) interests, please let the Library know, stating your reasons. In case of a legitimate complaint, the Library will make the material inaccessible and/or remove it from the website. Please Ask the Library: https://uba.uva.nl/en/contact, or a letter to: Library of the University of Amsterdam, Secretariat, Singel 425, 1012 WP Amsterdam, The Netherlands. You will be contacted as soon as possible. UvA-DARE is a service provided by the library of the University of Amsterdam (https://dare.uva.nl) Download date:04 Oct 2021 DISTINCT MATERNAL AND SOMATIC rRNA TYPES IN ZEBRAFISH DEVELOPMENT MAURO LOCATI DISTINCT MATERNAL AND SOMATIC rRNA TYPES IN ZEBRAFISH DEVELOPMENT MAURO LOCATI Distinct Maternal and Somatic rRNA Types in Zebrafish Development ACADEMISCH PROEFSCHRIFT ter verkrijging van de graad van doctor aan de Universiteit van Amsterdam op gezag van de Rector Magnificus prof. dr. ir. K.I.J. Maex ten overstaan van een door het College voor Promoties ingestelde commissie, in het openbaar te verdedigen in de Aula der Universiteit op woensdag 6 december 2017, te 13:00 uur door Mauro Locati geboren te Palermo, Italië Promotiecommissie: Promotor: Prof. -

Intersections Spring Symposium
INTERSECTIONS SPRING SYMPOSIUM APRIL TH, 8 Last Name First Name Project Title Page Analyzing the Effect of Urbanization on Abboud Bissan Phylogenetic signal in the Species Columba 1 Abouzahra Sharifah Peripheral IV Catheter Sterilization Device 2 Development of a Liquid Crystal Based Electron Adkins Raymond Shower Detector 2 Purification of Secreted Protein, Acidic and Rich in Cysteine (SPARC) Implicated for its Effects on Al Bahrani Zaid Intraocular Pressure 3 A Window into Metabolite Changes of Live Aljawad Yaqeen Bacteria Following Drug Infusion 3 Anderson Emily In-situ Resource Utilization on Mars 4 The Relationship Between Subjective and Objective Language Assessments For Second- Arredondo Kaitlynn Language Learners 4 Atkinson Scott MotionSense Wave: Toucheless Faucet 5 Ayyar Aneeka Hand Tremor Reduction Device 6 Modifying Low-Emissivity Glass for Use with Bailey Jessica Surface Enhanced Raman Spectroscopy 5 Automated Segmentation and Radiomic Characterization of Visceral Fat on Bowel MRIs for Barbur Iulia Crohn's Disease 6 Periodic Liquid Crystal Order: Theory and Barnaby Gavin-Rae Simulations 7 The Role of Dentistry in the U.S. Opioid Epidemic: Basore Makenna A Literature Review 7 Berends Hannah A Leader’s Role in a Nation’s Stability 8 A Study on Daily Cycles in Activity Levels of Bhatia Shireen Coyotes (Canis latrans) 8 Co-creation of an Intervention with African American Older Adults to Manage Stress Blackshire Gabrielle Associated with Self-management of 9 Developing to Developed: Entrepreneurship in Blatt Noah Latin America -

Integration of 198 Chip-Seq Datasets Reveals Human Cis-Regulatory Regions
Integration of 198 ChIP-seq datasets reveals human cis-regulatory regions Hamid Bolouri & Larry Ruzzo Thanks to Stephen Tapscott, Steven Henikoff & Zizhen Yao Slides will be available from: http://labs.fhcrc.org/bolouri/ Email [email protected] for manuscript (Bolouri & Ruzzo, submitted) Kleinjan & van Heyningen, Am. J. Hum. Genet., 2005, (76)8–32 Epstein, Briefings in Func. Genom. & Protemoics, 2009, 8(4)310-16 Regulation of SPi1 (Sfpi1, PU.1 protein) expression – part 1 miR155*, miR569# ~750nt promoter ~250nt promoter The antisense RNA • causes translational stalling • has its own promoter • requires distal SPI1 enhancer • is transcribed with/without SPI1. # Hikami et al, Arthritis & Rheumatism, 2011, 63(3):755–763 * Vigorito et al, 2007, Immunity 27, 847–859 Ebralidze et al, Genes & Development, 2008, 22: 2085-2092. Regulation of SPi1 expression – part 2 (mouse coordinates) Bidirectional ncRNA transcription proportional to PU.1 expression PU.1/ELF1/FLI1/GLI1 GATA1 GATA1 Sox4/TCF/LEF PU.1 RUNX1 SP1 RUNX1 RUNX1 SP1 ELF1 NF-kB SATB1 IKAROS PU.1 cJun/CEBP OCT1 cJun/CEBP 500b 500b 500b 500b 500b 750b 500b -18Kb -14Kb -12Kb -10Kb -9Kb Chou et al, Blood, 2009, 114: 983-994 Hoogenkamp et al, Molecular & Cellular Biology, 2007, 27(21):7425-7438 Zarnegar & Rothenberg, 2010, Mol. & cell Biol. 4922-4939 An NF-kB binding-site variant in the SPI1 URE reduces PU.1 expression & is GGGCCTCCCC correlated with AML GGGTCTTCCC Bonadies et al, Oncogene, 2009, 29(7):1062-72. SATB1 binding site A distal single nucleotide polymorphism alters long- range regulation of the PU.1 gene in acute myeloid leukemia Steidl et al, J Clin Invest. -

BMC Genomics Biomed Central
BMC Genomics BioMed Central Methodology article Open Access "NeuroStem Chip": a novel highly specialized tool to study neural differentiation pathways in human stem cells Sergey V Anisimov*, Nicolaj S Christophersen, Ana S Correia, Jia-Yi Li and Patrik Brundin Address: Neuronal Survival Unit, Wallenberg Neuroscience Center, Lund University, 221 84 Lund, Sweden Email: Sergey V Anisimov* - [email protected]; Nicolaj S Christophersen - [email protected]; Ana S Correia - [email protected]; Jia-Yi Li - [email protected]; Patrik Brundin - [email protected] * Corresponding author Published: 8 February 2007 Received: 21 September 2006 Accepted: 8 February 2007 BMC Genomics 2007, 8:46 doi:10.1186/1471-2164-8-46 This article is available from: http://www.biomedcentral.com/1471-2164/8/46 © 2007 Anisimov et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Human stem cells are viewed as a possible source of neurons for a cell-based therapy of neurodegenerative disorders, such as Parkinson's disease. Several protocols that generate different types of neurons from human stem cells (hSCs) have been developed. Nevertheless, the cellular mechanisms that underlie the development of neurons in vitro as they are subjected to the specific differentiation protocols are often poorly understood. Results: We have designed a focused DNA (oligonucleotide-based) large-scale microarray platform (named "NeuroStem Chip") and used it to study gene expression patterns in hSCs as they differentiate into neurons. -

Prostate News
Prostate Cancer and Prostatic Diseases (2005) 8, 2–6 & 2005 Nature Publishing Group All rights reserved 1365-7852/05 $30.00 www.nature.com/pcan Research Highlights Prostate News Prostate Cancer and Prostatic Diseases (2005) 8, 2–6. doi:10.1038/sj.pcan.4500793 Pokemon’s central role in cancer progression of prostate cancer. Epigenetic changes are reversible, which make them attractive targets for cancer development treatment with modulators that demethylate DNA and Researchers at New York’s Memorial Sloan-Kettering inhibit histone deacetylases, leading to reactivation of Cancer Center report novel findings in Nature regarding silenced genes. DNA methylation and histone modifica- a new cellular oncogene implicated in the development tions are important epigenetic mechanisms of gene of human cancers, including prostate neoplasia. Poke- regulation, and both play essential roles in tumor mon, a pop-culture inspired acronym for ‘POK erythroid initiation and progression. In prostate cancer, aberrant myeloid ontogenic factor’, is a POK protein family epigenetic events such as DNA hypo- and hypermethy- member encoded by the ZbTb7 gene and characterized lation and altered histone acetylation have been by its critical transcription function in cellular differ- observed to affect a large number of genes. However, entiation. Takahiro Maeda, Pier Paolo Pandolfi and no plenitude is without peril: although the number of colleagues show that Pokemon acts as a genuine proto- genes that undergo aberrant epigenetic inactivation oncogene when overexpressed by inhibiting the expres- associated with prostate cancer seems to be growing, sion of tumor suppressors, such as ARF. very few of these genes have yielded promising results To establish this novel finding, the authors conducted as potential tumor biomarkers for early diagnosis and a complete battery of genetic, clinical, and pathological risk assessment. -

RNA Polymerase I Transcription and Pre-Rrna Processing Are Linked by Specific SSU Processome Components
Downloaded from genesdev.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press RNA polymerase I transcription and pre-rRNA processing are linked by specific SSU processome components Jennifer E.G. Gallagher,2 David A. Dunbar,3 Sander Granneman,1 Brianna M. Mitchell,3 Yvonne Osheim,4 Ann L. Beyer,4 and Susan J. Baserga1,2,3,5 1Department of Molecular Biophysics and Biochemistry, 2Department of Genetics, and 3Department of Therapeutic Radiology, Yale University School of Medicine, New Haven, Connecticut 06520-8024, USA; 4Department of Microbiology, University of Virginia, Charlottesville, Virginia 22904, USA Sequential events in macromolecular biosynthesis are often elegantly coordinated. The small ribosomal subunit (SSU) processome is a large ribonucleoprotein (RNP) required for processing of precursors to the small subunit RNA, the 18S, of the ribosome. We have found that a subcomplex of SSU processome proteins, the t-Utps, is also required for optimal rRNA transcription in vivo in the yeast Saccharomyces cerevisiae.The t-Utps are ribosomal chromatin (r-chromatin)-associated, and they exist in a complex in the absence of the U3 snoRNA. Transcription is required neither for the formation of the subcomplex nor for its r-chromatin association. The t-Utps are associated with the pre-18S rRNAs independent of the presence of the U3 snoRNA. This association may thus represent an early step in the formation of the SSU processome. Our results indicate that rRNA transcription and pre-rRNA processing are coordinated via specific components of the SSU processome. [Keywords: Ribosome; RNA; transcription; RNA processing; SSU processome] Received May 27, 2004; revised version accepted August 19, 2004. -

Establishment of Mouse Primed Stem Cells by Combination of Activin and LIF Signaling
fcell-09-713503 July 31, 2021 Time: 12:43 # 1 ORIGINAL RESEARCH published: 05 August 2021 doi: 10.3389/fcell.2021.713503 Establishment of Mouse Primed Stem Cells by Combination of Activin and LIF Signaling Mengyi Wei1,2†, Yanglin Chen1,2,3†, Chaoyue Zhao1,2, Li Zheng1,2, Baojiang Wu1,2, Chen Chen1,2, Xihe Li1,2,4 and Siqin Bao1,2* 1 State Key Laboratory of Reproductive Regulation and Breeding of Grassland Livestock, Inner Mongolia University, Hohhot, China, 2 Institute of Animal Genetic Research of Mongolia Plateau, College of Life Sciences, Inner Mongolia University, Hohhot, China, 3 School of Basic Medical Sciences, Southern Medical University, Guangzhou, China, 4 Inner Mongolia Saikexing Institute of Breeding and Reproductive Biotechnology in Domestic Animal, Hohhot, China In mice, embryonic stem cells (ESCs) and epiblast stem cells (EpiSCs) are established from pre- and post-implantation embryos and represent the naive and primed state, respectively. Herein we used mouse leukemia inhibitory factor (LIF), which supports Edited by: Wei Jiang, ESCs self-renewal and Activin A (Act A), which is the main factor in maintaining EpiSCs Wuhan University, China in post-implantation epiblast cultures, to derive a primed stem cell line named ALSCs. Reviewed by: Like EpiSCs, ALSCs express key pluripotent genes Oct4, Sox2, and Nanog; one X Tetsuya S. Tanaka, Elixirgen Scientific, Inc., United States chromosome was inactivated; and the cells failed to contribute to chimera formation Fabiana Passaro, in vivo. Notably, compared to EpiSCs, ALSCs efficiently reversed to ESCs (rESCs) University of Naples Federico II, Italy on activation of Wnt signaling. Moreover, we also discovered that culturing EpiSCs in *Correspondence: AL medium for several passages favored Wnt signaling-driven naive pluripotency.