Chromosomal Aberrations in Head and Neck Squamous Cell Carcinomas in Norwegian and Sudanese Populations by Array Comparative Genomic Hybridization
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Variation in FCN1 Affects Biosynthesis of Ficolin-1 and Is Associated With
Genes and Immunity (2012) 13, 515–522 & 2012 Macmillan Publishers Limited All rights reserved 1466-4879/12 www.nature.com/gene ORIGINAL ARTICLE Variation in FCN1 affects biosynthesis of ficolin-1 and is associated with outcome of systemic inflammation L Munthe-Fog1, T Hummelshoj1, C Honore´ 1, ME Moller1, MO Skjoedt1, I Palsgaard1, N Borregaard2, HO Madsen1 and P Garred1 Ficolin-1 is a recognition molecule of the lectin complement pathway. The ficolin-1 gene FCN1 is polymorphic, but the functional and clinical consequences are unknown.The concentration of ficolin-1 in plasma and FCN1 polymorphisms in positions À 1981 (rs2989727), À 791 (rs28909068), À 542 (rs10120023), À 271 (rs28909976), À 144 (rs10117466) and þ 7918 (rs1071583) were determined in 100 healthy individuals. FCN1 expression by isolated monocytes and granulocytes and ficolin-1 levels in monocyte culture supernatants were assessed in 21 FCN1-genotyped individuals. FCN1 polymorphisms were determined in a cohort of 251 patients with systemic inflammation. High ficolin-1 plasma levels were significantly associated with the minor alleles in position À 542 and À 144. These alleles were also significantly associated with high FCN1 mRNA expression. The level of ficolin-1 in culture supernatants was significantly higher in individuals homozygous for the minor alleles at positions À 542 and À 144. Homozygosity for these alleles was significantly associated with fatal outcome in patients with systemic inflammation. None of the other investigated polymorphisms were associated with FCN1 and ficolin-1 expression, concentration or disease outcome. Functional polymorphic sites in the promoter region of FCN1 regulate both the expression and synthesis of ficolin-1 and are associated with outcome in severe inflammation. -

SPP1 and AGER As Potential Prognostic Biomarkers for Lung Adenocarcinoma
7028 ONCOLOGY LETTERS 15: 7028-7036, 2018 SPP1 and AGER as potential prognostic biomarkers for lung adenocarcinoma WEIGUO ZHANG, JUNLI FAN, QIANG CHEN, CAIPENG LEI, BIN QIAO and QIN LIU Henan Key Laboratory of Cancer Epigenetics, Department of Oncology Surgery, Cancer Institute and College of Clinical Medicine, The First Affiliated Hospital of Henan University of Science and Technology, Luoyang, Henan 471003, P.R. China Received August 11, 2017; Accepted January 5, 2018 DOI: 10.3892/ol.2018.8235 Abstract. Overdue treatment and prognostic evaluation lead Introduction to low survival rates in patients with lung adenocarcinoma (LUAD). To date, effective biomarkers for prognosis are still Lung adenocarcinoma (LUAD) is one of the three major required. The aim of the present study was to screen differen- histopathological subtypes along with squamous cell carci- tially expressed genes (DEGs) as biomarkers for prognostic noma (SqCLC) and large cell carcinoma (1). Currently, LUAD evaluation of LUAD. DEGs in tumor and normal samples has become the most common lung cancer with increasing were identified and analyzed for Kyoto Encyclopedia of Genes morbidity. This uptrend of incidence may be due to increasing and Genomes/Gene Ontology functional enrichments. The smoking rate and air pollution (2). Furthermore, LUAD is common genes that are up and downregulated were selected diagnosed at late stages (stage III and IV), when the cancer for prognostic analysis using RNAseq data in The Cancer has spread to nearby tissues and metastasis has occurred (3). Genome Atlas. Differential expression analysis was performed As well as overdue diagnosis that leads to delayed treatment, with 164 samples in GSE10072 and GSE7670 datasets. -

Genome-Wide Approach to Identify Risk Factors for Therapy-Related Myeloid Leukemia
Leukemia (2006) 20, 239–246 & 2006 Nature Publishing Group All rights reserved 0887-6924/06 $30.00 www.nature.com/leu ORIGINAL ARTICLE Genome-wide approach to identify risk factors for therapy-related myeloid leukemia A Bogni1, C Cheng2, W Liu2, W Yang1, J Pfeffer1, S Mukatira3, D French1, JR Downing4, C-H Pui4,5,6 and MV Relling1,6 1Department of Pharmaceutical Sciences, The University of Tennessee, Memphis, TN, USA; 2Department of Biostatistics, The University of Tennessee, Memphis, TN, USA; 3Hartwell Center, The University of Tennessee, Memphis, TN, USA; 4Department of Pathology, The University of Tennessee, Memphis, TN, USA; 5Department of Hematology/Oncology St Jude Children’s Research Hospital, The University of Tennessee, Memphis, TN, USA; and 6Colleges of Medicine and Pharmacy, The University of Tennessee, Memphis, TN, USA Using a target gene approach, only a few host genetic risk therapy increases, the importance of identifying host factors for factors for treatment-related myeloid leukemia (t-ML) have been secondary neoplasms increases. defined. Gene expression microarrays allow for a more 4 genome-wide approach to assess possible genetic risk factors Because DNA microarrays interrogate multiple ( 10 000) for t-ML. We assessed gene expression profiles (n ¼ 12 625 genes in one experiment, they allow for a ‘genome-wide’ probe sets) in diagnostic acute lymphoblastic leukemic cells assessment of genes that may predispose to leukemogenesis. from 228 children treated on protocols that included leukemo- DNA microarray analysis of gene expression has been used to genic agents such as etoposide, 13 of whom developed t-ML. identify distinct expression profiles that are characteristic of Expression of 68 probes, corresponding to 63 genes, was different leukemia subtypes.13,14 Studies using this method have significantly related to risk of t-ML. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -
Download Validation Data
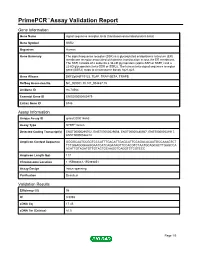
PrimePCR™Assay Validation Report Gene Information Gene Name signal sequence receptor, beta (translocon-associated protein beta) Gene Symbol SSR2 Organism Human Gene Summary The signal sequence receptor (SSR) is a glycosylated endoplasmic reticulum (ER) membrane receptor associated with protein translocation across the ER membrane. The SSR consists of 2 subunits a 34-kD glycoprotein (alpha-SSR or SSR1) and a 22-kD glycoprotein (beta-SSR or SSR2). The human beta-signal sequence receptor gene (SSR2) maps to chromosome bands 1q21-q23. Gene Aliases DKFZp686F19123, TLAP, TRAP-BETA, TRAPB RefSeq Accession No. NC_000001.10, NT_004487.19 UniGene ID Hs.74564 Ensembl Gene ID ENSG00000163479 Entrez Gene ID 6746 Assay Information Unique Assay ID qHsaCID0014663 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000295702, ENST00000529008, ENST00000480567, ENST00000531917, ENST00000526212 Amplicon Context Sequence GGGGCAATCCGGTCCCATTTGACATTGAGCATTCCAGACACAATGCCAAAGTCT TCTGGAGGGAAGGAATCATCAGATAGTTCCACGTCTAATGCAGCACTTGAGCCA ACATTGTAGATGTTGTACTGCAAGGTCAGGTCTCGTCCC Amplicon Length (bp) 117 Chromosome Location 1:155988061-155989851 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9998 cDNA Cq 17.45 cDNA Tm (Celsius) 81.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SSR2, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference -

(12) Patent Application Publication (10) Pub. No.: US 2015/0072349 A1 Diamandis Et Al
US 201500 72349A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2015/0072349 A1 Diamandis et al. (43) Pub. Date: Mar. 12, 2015 (54) CANCER BOMARKERS AND METHODS OF (52) U.S. Cl. USE CPC. G0IN33/57484 (2013.01); G0IN 2333/705 (2013.01) (71) Applicant: University Health Network, Toronto USPC ......................................... 435/6.12: 435/7.94 (CA) (57) ABSTRACT A method of evaluating a probability a Subject has a cancer, (72) Inventors: Eleftherios P. Diamandis, Toronto diagnosing a cancer and/or monitoring cancer progression (CA); Ioannis Prassas, Toronto (CA); comprising: a. measuring an amount of a biomarker selected Shalini Makawita, Toronto (CA); from the group consisting of CUZD1 and/or LAMC2 and/or Caitlin Chrystoja, Toronto (CA); Hari the group CUZD1, LAMC2, AQP8, CELA2B, CELA3B, M. Kosanam, Maple (CA) CTRB1, CTRB2, GCG, IAPP, INS, KLK1, PNLIPRP1, PNLIPRP2, PPY, PRSS3, REG3G, SLC30A8, KLK3, NPY, (21) Appl. No.: 14/385,449 PSCA, RLN1, SLC45A3, DSP GP73, DSG2, CEACAM7, CLCA1, GPA33, LEFTY1, ZG16, IRX5, LAMP3, MFAP4, (22) PCT Fled: Mar. 15, 2013 SCGB1A1, SFTPC, TMEM100, NPY, PSCA RLN1 and/or SLC45A3 in a test sample from a subject with cancer; (86) PCT NO.: PCT/CA2O13/OOO248 wherein the cancer is pancreas cancer if CUZD1, LAMC2, S371 (c)(1), AQP8, CELA2B, CELA3B, CTRB1, CTRB2, GCG, LAPP (2) Date: Sep. 23, 2014 INS, KLK1, PNLIPRP1, PNLIPRP2, PPY, PRSS3, REG3G, SLC30A8, DSP GP73 and/or DSG2 is selected; the cancer is colon cancer if CEACAM7, CLCA1, GPA33, LEFTY 1 and/ Related U.S. Application Data or ZG16 is selected, the cancer is lung cancer if IRX5, (60) Provisional application No. -

IGH Rearrangement in Myeloid Neoplasms Sion and Activation
CASE REPORTS genes such as MYC and BCL2, leading to their overexpres- IGH rearrangement in myeloid neoplasms sion and activation. Here, we found two IGH rearrange- ments in myeloid tumors, including an IGH-MECOM in a Though immunoglobulin genes are typically expressed myelodysplastic syndrome (MDS) and an IGH-CCNG1 in in B lymphocytes, recent studies found ectopic an AML. Our studies provide the first evidence that a once immunoglobulin expression in non-B-cell tumor cells believed B-cell tumor-specific oncogenic mechanism is including acute myeloid leukemia (AML). The also present in myeloid tumors. immunoglobulin genes, including immunoglobulin heavy Case #1: A 60-year-old female with a history of breast chain genes (IGH), light kappa (k) chain genes (IGK) and cancer presented with fatigue, bilateral flank pain, and 9 light lambda (λ) chain genes (IGL), are frequently hematuria. Complete blood count showed 24.82x10 /L of rearranged in B-cell tumors. These rearrangements result white blood cells, 11.2 g/dL of hemoglobin, 33.9% of in a juxtaposition of IG enhancers to the vicinity of onco- hematocrit, and 68x109/L of platelets. The bone marrow A B C D E Figure 1. Acute myeloid leukemia with an IGH-CCNG1 rearrangement. (A) Chromosome analysis of the bone marrow aspirate showed a complex karyotype, including an unbalanced translocation involving chromosomes 5, 14 and 19. Arrows indicate clonal aberrations. (B) Fluorescence in situ hybridization (FISH) on abnormal metaphase showed the 3'IGH (red) remaining on the derivative chromosome 14 and the 5'IGH lost, consistent with an unbalanced IGH rearrange- ment. -

Telomere-To-Telomere Assembly of a Complete Human X Chromosome W
https://doi.org/10.1038/s41586-020-2547-7 Accelerated Article Preview Telomere-to-telomere assembly of a W complete human X chromosome E VI Received: 30 July 2019 Karen H. Miga, Sergey Koren, Arang Rhie, Mitchell R. Vollger, Ariel Gershman, Andrey Bzikadze, Shelise Brooks, Edmund Howe, David Porubsky, GlennisE A. Logsdon, Accepted: 29 May 2020 Valerie A. Schneider, Tamara Potapova, Jonathan Wood, William Chow, Joel Armstrong, Accelerated Article Preview Published Jeanne Fredrickson, Evgenia Pak, Kristof Tigyi, Milinn Kremitzki,R Christopher Markovic, online 14 July 2020 Valerie Maduro, Amalia Dutra, Gerard G. Bouffard, Alexander M. Chang, Nancy F. Hansen, Amy B. Wilfert, Françoise Thibaud-Nissen, Anthony D. Schmitt,P Jon-Matthew Belton, Cite this article as: Miga, K. H. et al. Siddarth Selvaraj, Megan Y. Dennis, Daniela C. Soto, Ruta Sahasrabudhe, Gulhan Kaya, Telomere-to-telomere assembly of a com- Josh Quick, Nicholas J. Loman, Nadine Holmes, Matthew Loose, Urvashi Surti, plete human X chromosome. Nature Rosa ana Risques, Tina A. Graves Lindsay, RobertE Fulton, Ira Hall, Benedict Paten, https://doi.org/10.1038/s41586-020-2547-7 Kerstin Howe, Winston Timp, Alice Young, James C. Mullikin, Pavel A. Pevzner, (2020). Jennifer L. Gerton, Beth A. Sullivan, EvanL E. Eichler & Adam M. Phillippy C This is a PDF fle of a peer-reviewedI paper that has been accepted for publication. Although unedited, the Tcontent has been subjected to preliminary formatting. Nature is providing this early version of the typeset paper as a service to our authors and readers. The text andR fgures will undergo copyediting and a proof review before the paper is published in its fnal form. -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

Associated Antigen 9 Exerts in Vivo Tumor- Promoting Effects Via Its Coiled-Coil Region
INTERNATIONAL JOURNAL OF ONCOLOGY 39: 41-49, 2011 Intracellular estrogen receptor-binding fragment- associated antigen 9 exerts in vivo tumor- promoting effects via its coiled-coil region YOSHIHIRO MAEYAMA1,2, MAKOTO OTSU3, SHUJI KUBO4, TOMOKI YAMANO5, YASUAKI IIMURA1,2, MASAFUMI ONODERA6, SATOSHI KONDO2, YUKIO SAKIYAMA1 and TADASHI ARIGA1,7 1Group of Human Gene Therapy, 2Department of Surgical Oncology, Hokkaido University Graduate School of Medicine, N15W7 Sapporo, Hokkaido 060-8638; 3Division of Stem Cell Therapy, Center for Stem Cell Biology and Regenerative Medicine, The Institute of Medical Science, The University of Tokyo, 4-6-1 Shirokanedai, Minato-ku, Tokyo 108-8639; 4Department of Genetics and 5Division of Lower Gastrointestinal Surgery, Department of Surgery, Hyogo College of Medicine, 1-1 Mukogawa-cho, Nishinomiya, Hyogo 663-8501; 6Department of Human Genetics, National Center for Child Health and Development, 2-10-1 Okura, Setagaya-ku, Tokyo 157-8535; 7Department of Pediatrics, Hokkaido University Graduate School of Medicine, N15W7 Sapporo, Hokkaido 060-8638, Japan Received February 22, 2011; Accepted April 6, 2011 DOI: 10.3892/ijo.2011.1026 Abstract. Estrogen receptor-binding fragment-associated showed a predominantly cytoplasmic localization in the antigen 9 (EBAG9) is a tumor-promoting factor of largely tumor cells. Collectively, these results suggest that EBAG9 unknown function. To assess a causative role of EBAG9 in overexpression can be causative in enhancing the malignant advanced malignancies, we generated the EG7-OVA and properties of tumor cells, and that tumor promotion likely MethA murine tumor cell lines that stably express full-length requires EBAG9 intracellular association with as yet unidentified or truncated EBAG9 protein, using retroviral-mediated gene binding partners via the coiled-coil region. -

Estrogen Receptor–Binding Fragment–Associated Antigen 9 Is a Tumor-Promoting and Prognostic Factor for Renal Cell Carcinoma
Research Article Estrogen Receptor–Binding Fragment–Associated Antigen 9 Is a Tumor-Promoting and Prognostic Factor for Renal Cell Carcinoma Tetsuo Ogushi,1 Satoru Takahashi,1 Takumi Takeuchi,1 Tomohiko Urano,2 Kuniko Horie-Inoue,3 Jinpei Kumagai,1 Tadaichi Kitamura,1 Yasuyoshi Ouchi,2 Masami Muramatsu,3 and Satoshi Inoue2,3 Departments of 1Urology and 2Geriatric Medicine, Faculty of Medicine, The University of Tokyo, Hongo, Bunkyo-ku, Tokyo, Japan and 3Research Center for Genomic Medicine, Saitama Medical School, Yamane, Hidaka-shi, Saitama, Japan Abstract target organs as well as several other tissues such as brain, liver, The estrogen receptor–binding fragment–associated antigen and kidney (2). The protein expression of EBAG9 is estrogen inducible, as it has been shown in ovariectomized mice treated 9(EBAG9) has been identified as a primary estrogen- h responsive gene in human breast cancer MCF7 cells. A high with 17 -estradiol administration (2). The physiologic function of expression of EBAG9 has been observed in invasive breast EBAG9 has not been well defined, yet the molecule may be cancer and advanced prostate cancer, suggesting a tumor- implicated in cancer pathophysiology, with several lines of evidence promoting role of the protein in malignancies. Here we show of the protein expression in malignancies, including breast (3), that intratumoral (i.t.) administration of small interfering ovarian (4), prostate (5), and hepatocellular carcinomas (6). In RNA against EBAG9 exerted overt regression of tumors prostate cancer (5), EBAG9 expression significantly correlated with following s.c. implantation of murine renal cell carcinoma advanced pathologic stages and high Gleason score (P = 0.0305 and P < 0.0001, respectively), suggesting the abundance of EBAG9 may (RCC) Renca cells. -

Prognostic Significance of TIMP-1 in Non-Small Cell Lung Cancer
ANTICANCER RESEARCH 31: 4031-4038 (2011) Prognostic Significance of TIMP-1 in Non-small Cell Lung Cancer MARTIN PESTA1, VLASTIMIL KULDA2, RADEK KUCERA1, MILOS PESEK3, JINDRA VRZALOVA1, VACLAV LISKA4, LADISLAV PECEN1, VLADISLAV TRESKA4, JARMIL SAFRANEK4, MARKETA PRAZAKOVA1, ONDREJ VYCITAL4, JAN BRUHA4, LUBOS HOLUBEC5 and ONDREJ TOPOLCAN1 Departments of 1Internal Medicine II and 2Biochemistry, Faculty of Medicine in Pilsen, Charles University in Prague, Czech Republic; 3TRN Clinic, University Hospital and Medical Faculty in Pilsen, Czech Republic; Departments of 4Surgery and 5Oncology, University Teaching Hospital, Pilsen, Czech Republic Abstract. Tissue inhibitor of metalloproteinases 1 (TIMP1) the mortality 48.7/100 000 per year (1). Non-small cell lung regulates not only extracellular matrix catabolism but the cancer (NSCLC) accounts for 80% of all cases (2). major effect in tumor tissue is promotion of cell growth and Although there are tumor markers routinely used for solid anti-apoptotic activity. The aim of our study was to evaluate tumors, some of them having become gold standards (3), plasma TIMP1 levels and tissue TIMP1 mRNA expression as there is still an effort being made to search for new ones. prognostic markers in NSCLC patients. Patients and Tissue inhibitor of metalloproteinases 1 (TIMP1) has been Methods: We studied a group of 108 patients with NSCLC studied intensively over the last ten years and in colorectal who had undergone lung surgery. Estimation of TIMP1 cancer it seems to be promising as a prognostic marker in mRNA was performed by quantitative polymerase chain clinical use (4). A Danish-Australian endoscopy study group reaction (qPCR) and estimation of plasma TIMP1 protein on colorectal cancer detection is trying to use TIMP1 as a using enzyme-linked immunosorbent assay (ELISA).