Isolation and Identification of Oleaginous Yeasts
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Phylogenetic Circumscription of Saccharomyces, Kluyveromyces
FEMS Yeast Research 4 (2003) 233^245 www.fems-microbiology.org Phylogenetic circumscription of Saccharomyces, Kluyveromyces and other members of the Saccharomycetaceae, and the proposal of the new genera Lachancea, Nakaseomyces, Naumovia, Vanderwaltozyma and Zygotorulaspora Cletus P. Kurtzman à Microbial Genomics and Bioprocessing Research Unit, National Center for Agricultural Utilization Research, Agricultural Research Service, U.S. Department of Agriculture, 1815 N. University Street, Peoria, IL 61604, USA Received 22 April 2003; received in revised form 23 June 2003; accepted 25 June 2003 First published online Abstract Genera currently assigned to the Saccharomycetaceae have been defined from phenotype, but this classification does not fully correspond with species groupings determined from phylogenetic analysis of gene sequences. The multigene sequence analysis of Kurtzman and Robnett [FEMS Yeast Res. 3 (2003) 417^432] resolved the family Saccharomycetaceae into 11 well-supported clades. In the present study, the taxonomy of the Saccharomyctaceae is evaluated from the perspective of the multigene sequence analysis, which has resulted in reassignment of some species among currently accepted genera, and the proposal of the following five new genera: Lachancea, Nakaseomyces, Naumovia, Vanderwaltozyma and Zygotorulaspora. ß 2003 Federation of European Microbiological Societies. Published by Elsevier B.V. All rights reserved. Keywords: Saccharomyces; Kluyveromyces; New ascosporic yeast genera; Molecular systematics; Multigene phylogeny 1. Introduction support the maintenance of three distinct genera. Yarrow [8^10] revived the concept of three genera and separated The name Saccharomyces was proposed for bread and Torulaspora and Zygosaccharomyces from Saccharomyces, beer yeasts by Meyen in 1838 [1], but it was Reess in 1870 although species assignments were often di⁄cult. -

(Vles) in the Yeast Debaryomyces Hansenii
toxins Article New Cytoplasmic Virus-Like Elements (VLEs) in the Yeast Debaryomyces hansenii Xymena Połomska 1,* ,Cécile Neuvéglise 2, Joanna Zyzak 3, Barbara Zarowska˙ 1, Serge Casaregola 4 and Zbigniew Lazar 1 1 Department of Biotechnology and Food Microbiology, Faculty of Biotechnology and Food Science, Wrocław University of Environmental and Life Sciences (WUELS), 50-375 Wroclaw, Poland; [email protected] (B.Z.);˙ [email protected] (Z.L.) 2 SPO, INRAE, Montpellier SupAgro, Université de Montpellier, 34060 Montpellier, France; [email protected] 3 Department of Microbiology, Laboratory of Microbiome Immunobiology, Ludwik Hirszfeld Institute of Immunology and Experimental Therapy, Polish Academy of Sciences, 53-114 Wroclaw, Poland; [email protected] 4 INRAE, AgroParisTech, Micalis Institute, CIRM-Levures, Université Paris-Saclay, 78350 Jouy-en-Josas, France; [email protected] * Correspondence: [email protected]; Tel.: +48-71-3207-791 Abstract: Yeasts can have additional genetic information in the form of cytoplasmic linear dsDNA molecules called virus-like elements (VLEs). Some of them encode killer toxins. The aim of this work was to investigate the prevalence of such elements in D. hansenii killer yeast deposited in culture collections as well as in strains freshly isolated from blue cheeses. Possible benefits to the host from harboring such VLEs were analyzed. VLEs occurred frequently among fresh D. hansenii isolates (15/60 strains), as opposed to strains obtained from culture collections (0/75 strains). Eight new different systems were identified: four composed of two elements and four of three elements. Full sequences of three new VLE systems obtained by NGS revealed extremely high conservation Citation: Połomska, X.; Neuvéglise, among the largest molecules in these systems except for one ORF, probably encoding a protein C.; Zyzak, J.; Zarowska,˙ B.; resembling immunity determinant to killer toxins of VLE origin in other yeast species. -

Thesis Contents
Genome diversity in Torulaspora microellipsoides and its contribution to the evolution of the Saccharomyces genus 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 0.1 This thesis is presented for the PhD degree of the University of Valencia Thesis Director: Dr. Eladio Barrio Esparducer Thesis Supervisor: Dr. Mercedes Costell Roselló Adriana Mena Romero Valencia, June 2018 El Dr. Eladio Barrio Esparducer, Profesor Titular del Departamento de Genética de la Universitat de València, adscrito como investigador al Departamento de Biotecnología del Instituto de Agroquímica y Tecnología de los Alimentos, CSIC. CERTIFICA Que el presente trabajo titulado “Genome diversity in Torulaspora microellipsoides and its contribution to the evolution of the Saccharomyces genus”, que presenta Dª Adriana Mena Romero para optar al grado de doctor en Biotecnología por la Universitat de València, ha sido realizado bajo su dirección en el Departamento de Genética de la Universidad de Valencia y en el Departamento de Biotecnología del Instituto de Agroquímica y Tecnología de los Alimentos, CSIC. Y para que conste para los trámites de lectura y defensa de la tesis doctoral, en cumplimiento de la legislación vigente, firma el presente certificado en Valencia a 15 de Junio de 2018 Fdo. Eladio Barrio Esparducer Agradecimientos (Acknowledgements) Agradecimientos Todo este trabajo no tendría sentido sin la gente que lleva años apoyándome para sacarlo adelante. -

Yeasts and Yeast-Like Fungi As an Element of Purity Assessment of Surface Waters
Polish J. of Environ. Stud. Vol. 20, No. 2 (2011), 267-274 Original Research Yeasts and Yeast-Like Fungi as an Element of Purity Assessment of Surface Waters Anna Biedunkiewicz*, Ewelina Baranowska Chair of Mycology, University of Warmia and Mazury, Oczapowskiego 1A, 10-957 Olsztyn, Poland Received: 2 August 2010 Accepted: 9 November 2010 Abstract The aim of our study was to determine species biodiversity of yeasts and yeast-like fungi in the stratum of surface water of Lake Tyrsko, as well as to evaluate the condition of waters of that lake based on the iso- lated species of fungi. The study was carried out from June 2007 to June 2008. The isolation and determination of the number of fungal colonies were conducted with the use of the membrane filter method. Fungal diagnostics were con- ducted based on the morphological and biochemical characteristics of the fungi. These characteristics were additionally used for taxonomic identification. In the course of the study we isolated 56 species of yeasts and yeast-like fungi belonging to 26 genera, with the predominating genus being Kluyveromyces (13 species), followed by less frequent genera: Candida (8 species), and Debaryomyces (7 species). The highest number of isolates were obtained in the spring (51), fewer in the summer (36), and the lowest in autumn (15). Species indicatory of the self-purifying process of the water examined were frequently isolated (Debaryomyces hansenii, Pichia membranifaciens). Results obtained in the study confirm suggestions of other authors that yeasts and yeast-like fungi may be useful as indicatory organisms showing the extent and character of water contamination and as bioindicators of sanitary and hygienic evaluation of water. -

Phylogenetic Circumscription of Saccharomyces, Kluyveromyces
FEMS Yeast Research 4 (2003) 233^245 www.fems-microbiology.org Phylogenetic circumscription of Saccharomyces, Kluyveromyces and other members of the Saccharomycetaceae, and the proposal of the new genera Lachancea, Nakaseomyces, Naumovia, Vanderwaltozyma and Zygotorulaspora Downloaded from https://academic.oup.com/femsyr/article-abstract/4/3/233/562841 by guest on 29 May 2020 Cletus P. Kurtzman à Microbial Genomics and Bioprocessing Research Unit, National Center for Agricultural Utilization Research, Agricultural Research Service, U.S. Department of Agriculture, 1815 N. University Street, Peoria, IL 61604, USA Received 22 April 2003; received in revised form 23 June 2003; accepted 25 June 2003 First published online Abstract Genera currently assigned to the Saccharomycetaceae have been defined from phenotype, but this classification does not fully correspond with species groupings determined from phylogenetic analysis of gene sequences. The multigene sequence analysis of Kurtzman and Robnett [FEMS Yeast Res. 3 (2003) 417^432] resolved the family Saccharomycetaceae into 11 well-supported clades. In the present study, the taxonomy of the Saccharomyctaceae is evaluated from the perspective of the multigene sequence analysis, which has resulted in reassignment of some species among currently accepted genera, and the proposal of the following five new genera: Lachancea, Nakaseomyces, Naumovia, Vanderwaltozyma and Zygotorulaspora. ß 2003 Federation of European Microbiological Societies. Published by Elsevier B.V. All rights reserved. Keywords: Saccharomyces; Kluyveromyces; New ascosporic yeast genera; Molecular systematics; Multigene phylogeny 1. Introduction support the maintenance of three distinct genera. Yarrow [8^10] revived the concept of three genera and separated The name Saccharomyces was proposed for bread and Torulaspora and Zygosaccharomyces from Saccharomyces, beer yeasts by Meyen in 1838 [1], but it was Reess in 1870 although species assignments were often di⁄cult. -

Comparative Genomics of Protoploid Saccharomycetaceae
Downloaded from genome.cshlp.org on October 5, 2021 - Published by Cold Spring Harbor Laboratory Press Evolution of protoploid yeast genomes ___________________________________________________________________________ Comparative genomics of protoploid Saccharomycetaceae. The Génolevures Consortium (1) Running title: Evolution of protoploid yeast genomes Key words: protein families, synteny, tandems, annotation, SONS, ancestor genome Corresponding author: Jean Luc Souciet Université de Strasbourg, CNRS, UMR 7156 Institut de Botanique, 28 rue Goethe, F-67000 Strasbourg, France Tel: 33 3 90 24 18 17 FAX: 33 3 90 24 20 28 e-mail: [email protected] (1) List of participants and affiliations appear at the end of the paper 1 Downloaded from genome.cshlp.org on October 5, 2021 - Published by Cold Spring Harbor Laboratory Press Evolution of protoploid yeast genomes ___________________________________________________________________________ Abstract Our knowledge on yeast genomes remains largely dominated by the extensive studies on Saccharomyces cerevisiae and the consequences of its ancestral duplication, leaving the evolution of the entire class of hemiascomycetes only partly explored. We concentrate here on five species of Saccharomycetaceae, a large subdivision of hemiascomycetes, that we call “protoploid” because they diverged from the S. cerevisiae lineage prior to its genome duplication. We determined the complete genome sequences of three of these species, Kluyveromyces (Lachancea) thermotolerans and Saccharomyces (Lachancea) kluyveri (two members of the newly described Lachancea clade) and Zygosaccharomyces rouxii. We included in our comparisons the previously available sequences of Klyveromyces lactis and Ashbya (Eremothecium) gossypii. Despite their broad evolutionary range and significant individual variations in each lineage, the five protoploid Saccharomycetaceae share a core repertoire of ca. -
Supplementary File 1
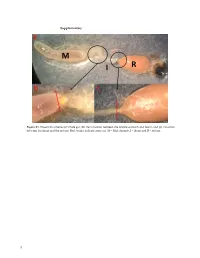
Supplementary Figure S1. Dissection scheme (a) whole gut, (b) the transition between the middle stomach and ileum, and (c) transition between the ileum and the rectum. Red streaks indicate areas cut. M = Mid stomach, I = ileum and R = rectum. 1 Figure S2. Relative abundance of the 20 most abundant fungal OTUs across months per gut part. Taxonomic assignments are provided in Table S1. 2 Figure S3. Relative abundance of the bacterial OUTs (with abundance > 1%) across months per gut part. The color code is given at the genus level. 3 Table S1. Taxonomic assignments of fungal OTUs by Blast. OTU# Taxonomy by blast nt collection 11/11‐2020 Phylum Accesion# OTU_1 Aureobasidium pullulans Ascomycota MW085051 OTU_2 Meyerozyma guilliermondii Ascomycota MT988167 OTU_3 Starmerella apicola Ascomycota KY101940 OTU_4 Debaryomyces hansenii Ascomycota MW051606 OTU_5 Engyodontium sp. Ascomycota MN905797 Basidiomy‐ OTU_6 Mrakia sp. MT505696 cota Basidiomy‐ OTU_7 Tausonia pullulans MN900123 cota OTU_8 Unknown OTU_9 Unknown OTU_10 Penicillium corylophilum Ascomycota MT906500 OTU_11 Penicillium sp. Ascomycota MT993349 OTU_12 Cladosporium sp. Ascomycota MW077705 Phaffia rhodozyma/Xanthophyllomyces dendro‐ Basidiomy‐ OTU_13 KY104501/DQ904243 rhous cota Basidiomy‐ OTU_14 Filobasidium wieringae MN899199 cota OTU_15 Hanseniaspora uvarum Ascomycota MN556596 Basidiomy‐ OTU_16 Mrakia gelida MN460370 cota OTU_17 Uncultured fungus clone MK717974 OTU_18 Cladosporium allicinum Ascomycota MT974153 OTU_20 Taphrina carpini Ascomycota MK782181 Basidiomy‐ OTU_25 Papiliotrema -

High-Level Classification of the Fungi and a Tool for Evolutionary Ecological Analyses
Fungal Diversity (2018) 90:135–159 https://doi.org/10.1007/s13225-018-0401-0 (0123456789().,-volV)(0123456789().,-volV) High-level classification of the Fungi and a tool for evolutionary ecological analyses 1,2,3 4 1,2 3,5 Leho Tedersoo • Santiago Sa´nchez-Ramı´rez • Urmas Ko˜ ljalg • Mohammad Bahram • 6 6,7 8 5 1 Markus Do¨ ring • Dmitry Schigel • Tom May • Martin Ryberg • Kessy Abarenkov Received: 22 February 2018 / Accepted: 1 May 2018 / Published online: 16 May 2018 Ó The Author(s) 2018 Abstract High-throughput sequencing studies generate vast amounts of taxonomic data. Evolutionary ecological hypotheses of the recovered taxa and Species Hypotheses are difficult to test due to problems with alignments and the lack of a phylogenetic backbone. We propose an updated phylum- and class-level fungal classification accounting for monophyly and divergence time so that the main taxonomic ranks are more informative. Based on phylogenies and divergence time estimates, we adopt phylum rank to Aphelidiomycota, Basidiobolomycota, Calcarisporiellomycota, Glomeromycota, Entomoph- thoromycota, Entorrhizomycota, Kickxellomycota, Monoblepharomycota, Mortierellomycota and Olpidiomycota. We accept nine subkingdoms to accommodate these 18 phyla. We consider the kingdom Nucleariae (phyla Nuclearida and Fonticulida) as a sister group to the Fungi. We also introduce a perl script and a newick-formatted classification backbone for assigning Species Hypotheses into a hierarchical taxonomic framework, using this or any other classification system. We provide an example -

The 100 Years of the Fungus Collection Mucl 1894-1994
THE 100 YEARS OF THE FUNGUS COLLECTION MUCL 1894-1994 Fungal Taxonomy and Tropical Mycology: Quo vadis ? Taxonomy and Nomenclature of the Fungi Grégoire L. Hennebert Catholic University of Louvain, Belgium Notice of the editor This document is now published as an archive It is available on www.Mycotaxon.com It is also produced on CD and in few paperback copies G. L. Hennebert ed. Published by Mycotaxon, Ltd. Ithaca, New York, USA December 2010 ISBN 978-0-930845-18-6 (www pdf version) ISBN 978-0-930845-17-9 (paperback version) DOI 10.5248/2010MUCL.pdf 1894-1994 MUCL Centenary CONTENTS Lists of participants 8 Forword John Webser 13 PLENARY SESSION The 100 Year Fungus Culture Collection MUCL, June 29th, 1994 G.L. Hennebert, UCL Mycothèque de l'Université Catholique de Louvain (MUCL) 17 D. Hawksworth, IMI, U.K. Fungal genetic resource collections and biodiversity. 27 D. van der Mei, CBS, MINE, Netherlands The fungus culture collections in Europe. 34 J. De Brabandere, BCCM, Belgium The Belgian Coordinated Collections of Microorganisms. 40 Fungal Taxonomy and tropical Mycology G.L. Hennebert, UCL Introduction. Fungal taxonomy and tropical mycology: Quo vadis ? 41 C.P. Kurtzman, NRRL, USA Molecular taxonomy in the yeast fungi: present and future. 42 M. Blackwell, Louisiana State University, USA Phylogeny of filamentous fungi deduced from analysis of molecular characters: present and future. 52 J. Rammeloo, National Botanical Garden, Belgium Importance of morphological and anatomical characters in fungal taxonomy. 57 M.F. Roquebert, Natural History Museum, France Possible progress of modern morphological analysis in fungal taxonomy. 63 A.J. -

Diversity and Distribution of Hidden Cultivable Fungi Associated with Marine Animals of Antarctica Fungal Biology
Fungal Biology 123 (2019) 507e516 Contents lists available at ScienceDirect Fungal Biology journal homepage: www.elsevier.com/locate/funbio Diversity and distribution of hidden cultivable fungi associated with marine animals of Antarctica Valeria Martins Godinho a, Maria Theresa Rafaela de Paula a, Debora Amorim Saraiva Silva a, Karla Paresque b, Aline Paternostro Martins c, * Pio Colepicolo c, Carlos Augusto Rosa a, Luiz Henrique Rosa a, a Departamento de Microbiologia, Instituto de Ci^encias Biologicas, Universidade Federal de Minas Gerais, Belo Horizonte, MG, Brazil b Laboratorio de Bentologia, Universidade Federal de Alagoas, Maceio, AL, Brazil c Departamento de Bioquímica, Instituto de Química, Universidade de Sao~ Paulo, Sao~ Paulo, Brazil article info abstract Article history: In the present study, we surveyed the distribution and diversity of fungal assemblages associated with 10 Received 22 November 2018 species of marine animals from Antarctica. The collections yielded 83 taxa from 27 distinct genera, which Received in revised form were identified using molecular biology methods. The most abundant taxa were Cladosporium sp. 1, 17 April 2019 Debaryomyces hansenii, Glaciozyma martinii, Metschnikowia australis, Pseudogymnoascus destructans, Accepted 1 May 2019 Thelebolus cf. globosus, Pseudogymnoascus pannorum, Tolypocladium tundrense, Metschnikowia australis, Available online 14 May 2019 and different Penicillium species. The diversity, richness, and dominance of fungal assemblages ranged Corresponding Editor: Nik Money among the host; however, in general, the fungal community, which was composed of endemic and cold- adapted cosmopolitan taxa distributed across the different sites of Antarctic Peninsula, displayed high Keywords: diversity, richness, and dominance indices. Our results contribute to knowledge about fungal diversity in Antarctic ocean the marine environment across the Antarctic Peninsula and their phylogenetic relationships with species Extremophiles that occur in other cold, temperate, and tropical regions of the World. -

Biotechnological Importance of Torulaspora Delbrueckii: from the Obscurity to the Spotlight
Journal of Fungi Review Biotechnological Importance of Torulaspora delbrueckii: From the Obscurity to the Spotlight Ticiana Fernandes 1,2 , Flávia Silva-Sousa 1,2 ,Fábio Pereira 1,2, Teresa Rito 1,2 , Pedro Soares 1,2, Ricardo Franco-Duarte 1,2,* and Maria João Sousa 1,2 1 CBMA (Centre of Molecular and Environmental Biology), Department of Biology, University of Minho, 4710-057 Braga, Portugal; [email protected] (T.F.); sousa.s.fl[email protected] (F.S.-S.); [email protected] (F.P.); [email protected] (T.R.); [email protected] (P.S.); [email protected] (M.J.S.) 2 Institute of Science and Innovation for Bio-Sustainability (IB-S), University of Minho, 4710-057 Braga, Portugal * Correspondence: ricardofi[email protected] or [email protected]; Tel.: +351-253-604-310; Fax: +351-253-678-980 Abstract: Torulaspora delbrueckii has attracted interest in recent years, especially due to its biotech- nological potential, arising from its flavor- and aroma-enhancing properties when used in wine, beer or bread dough fermentation, as well as from its remarkable resistance to osmotic and freez- ing stresses. In the present review, genomic, biochemical, and phenotypic features of T. delbrueckii are described, comparing them with other species, particularly with the biotechnologically well- established yeast, Saccharomyces cerevisiae. We conclude about the aspects that make this yeast a promising biotechnological model to be exploited in a wide range of industries, particularly in wine and bakery. A phylogenetic analysis was also performed, using the core proteome of T. delbrueckii, to compare the number of homologous proteins relative to the most closely related species, under- Citation: Fernandes, T.; Silva-Sousa, standing the phylogenetic placement of this species with robust support. -

Descriptions of Medical Fungi
DESCRIPTIONS OF MEDICAL FUNGI THIRD EDITION (revised November 2016) SARAH KIDD1,3, CATRIONA HALLIDAY2, HELEN ALEXIOU1 and DAVID ELLIS1,3 1NaTIONal MycOlOgy REfERENcE cENTRE Sa PaTHOlOgy, aDElaIDE, SOUTH aUSTRalIa 2clINIcal MycOlOgy REfERENcE labORatory cENTRE fOR INfEcTIOUS DISEaSES aND MIcRObIOlOgy labORatory SERvIcES, PaTHOlOgy WEST, IcPMR, WESTMEaD HOSPITal, WESTMEaD, NEW SOUTH WalES 3 DEPaRTMENT Of MOlEcUlaR & cEllUlaR bIOlOgy ScHOOl Of bIOlOgIcal ScIENcES UNIvERSITy Of aDElaIDE, aDElaIDE aUSTRalIa 2016 We thank Pfizera ustralia for an unrestricted educational grant to the australian and New Zealand Mycology Interest group to cover the cost of the printing. Published by the authors contact: Dr. Sarah E. Kidd Head, National Mycology Reference centre Microbiology & Infectious Diseases Sa Pathology frome Rd, adelaide, Sa 5000 Email: [email protected] Phone: (08) 8222 3571 fax: (08) 8222 3543 www.mycology.adelaide.edu.au © copyright 2016 The National Library of Australia Cataloguing-in-Publication entry: creator: Kidd, Sarah, author. Title: Descriptions of medical fungi / Sarah Kidd, catriona Halliday, Helen alexiou, David Ellis. Edition: Third edition. ISbN: 9780646951294 (paperback). Notes: Includes bibliographical references and index. Subjects: fungi--Indexes. Mycology--Indexes. Other creators/contributors: Halliday, catriona l., author. Alexiou, Helen, author. Ellis, David (David H.), author. Dewey Number: 579.5 Printed in adelaide by Newstyle Printing 41 Manchester Street Mile End, South australia 5031 front cover: Cryptococcus neoformans, and montages including Syncephalastrum, Scedosporium, Aspergillus, Rhizopus, Microsporum, Purpureocillium, Paecilomyces and Trichophyton. back cover: the colours of Trichophyton spp. Descriptions of Medical Fungi iii PREFACE The first edition of this book entitled Descriptions of Medical QaP fungi was published in 1992 by David Ellis, Steve Davis, Helen alexiou, Tania Pfeiffer and Zabeta Manatakis.