EPH/Ephrin Profile and EPHB2 Expression Predicts Patient Survival in Breast Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Targeting Synthetic Lethality Between the SRC Kinase and the EPHB6 Receptor May Benefit Cancer Treatment
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 7, No. 31 Research Paper Targeting synthetic lethality between the SRC kinase and the EPHB6 receptor may benefit cancer treatment James M. Paul1,*, Behzad Toosi2,*, Frederick S. Vizeacoumar2,*, Kalpana Kalyanasundaram Bhanumathy2, Yue Li3,4,5, Courtney Gerger2, Amr El Zawily2,6, Tanya Freywald7, Deborah H. Anderson7, Darrell Mousseau8, Rani Kanthan2, Zhaolei Zhang3,4, Franco J. Vizeacoumar2,7, Andrew Freywald2 1Department of Biochemistry, University of Saskatchewan, Saskatoon, SK, S7N 5E5, Canada 2Department of Pathology and Laboratory Medicine, College of Medicine, University of Saskatchewan, Royal University Hospital, Saskatoon, SK, S7N 0W8, Canada 3Department of Computer Science, University of Toronto, Toronto, ON, M5S 3G4, Canada 4The Donnelly Centre, University of Toronto, Toronto, ON, M5S 3E1, Canada 5Present address: Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, MA, 02139, USA 6Faculty of Science, Damanhour University, Damanhour, 22516, Egypt 7Cancer Research, Saskatchewan Cancer Agency, Saskatoon, SK, S7N 5E5, Canada 8Cell Signaling Laboratory, Neuroscience Cluster, University of Saskatchewan, Saskatoon, SK, S7N 5E5, Canada *These authors contributed equally to this work Correspondence to: Franco J. Vizeacoumar, email: [email protected] Andrew Freywald, email: [email protected] Keywords: breast cancer, genetic interaction, synthetic lethality, EPHB6, SRC kinase Received: April 22, 2016 Accepted: June 17, 2016 Published: July 13, 2016 ABSTRACT Application of tumor genome sequencing has identified numerous loss-of-function alterations in cancer cells. While these alterations are difficult to target using direct interventions, they may be attacked with the help of the synthetic lethality (SL) approach. In this approach, inhibition of one gene causes lethality only when another gene is also completely or partially inactivated. -
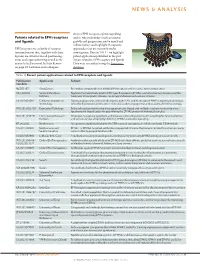
Patents Related to EPH Receptors and Ligands
NEWS & ANALYSIS discuss EPH receptor–ephrin signalling Patents related to EPH receptors and its role in disorders such as tumour and ligands growth and progression, nerve injury and inflammation, and highlight therapeutic EPH receptors are a family of receptor approaches that are currently under tyrosine kinases that, together with their investigation. Here in TABLE 1 we highlight ligands, are involved in cell positioning, patent applications published in the past tissue and organ patterning as well as the 3 years related to EPH receptors and ligands. control of cell survival. In their Review Data were researched using the Espacenet on page 39, Lackman and colleagues database. Table 1 | Recent patent applications related to EPH receptors and ligands Nature Reviews | Drug Discovery Publication Applicants Subject numbers NZ 581397 AstraZeneca Pyrimidine compounds that inhibit EPH receptors and are useful for treating cancer HK 1108702 Sanford-Burnham Peptides that selectively bind to EPH type-B receptors (EPHBs); useful for tumour imaging and the Institute treatment of neoplastic disease, neurological disease and vascular disease US 2013091591 California Institute of During angiogenesis, arterial cells express ephrin B2, and its receptor EPHB4 is expressed on venous Technology cells; this distinction can be used in methods to alter angiogenesis and to assess the effect of drugs WO 2013052710 Expression Pathology Selected reaction monitoring mass spectrometry-based and multiple reaction monitoring mass spectrometry-based assays for quantifying -

The Kinase Defective EPHB6 Receptor Tyrosine Kinase Activates MAP Kinase Signaling in Lung Adenocarcinoma
175-179.qxd 29/5/2009 01:21 ÌÌ ™ÂÏ›‰·175 INTERNATIONAL JOURNAL OF ONCOLOGY 35: 175-179, 2009 175 The kinase defective EPHB6 receptor tyrosine kinase activates MAP kinase signaling in lung adenocarcinoma JUN YU1,2, ETMAR BULK1, PING JI1, ANTJE HASCHER1, STEFFEN KOSCHMIEDER1, WOLFGANG E. BERDEL1 and CARSTEN MÜLLER-TIDOW1 1Department of Medicine, Hematology and Oncology, University of Münster, Münster, Germany; 2Department of Preclinical Experiment Center, Fourth Military Medical University, Xi'an, P.R. China Received January 28, 2009; Accepted March 13, 2009 DOI: 10.3892/ijo_00000326 Abstract. Decreased expression levels of EPHB6, a member Elk-1 (8,9). ERK1/2 are negatively regulated by a family of of the receptor tyrosine kinases (RTKs), are associated with dual-specificity (Thr/Tyr) MAPK phosphatases, known as an increased risk of metastasis development in early stage DUSPs or MKPs, and pharmacologically by MEK inhibitors non-small cell lung cancer (NSCLC). However, the signaling such as U0126 and PD98059 (10). properties of the kinase-defective EPHB6 receptor are not EPH receptors form the largest known subfamily of well-understood. Here, we show that expression of EPHB6 receptor tyrosine kinases, and to date, the EPH subfamily in A549 lung adenocarinoma cells led to phosphorylation of contains 16 members in vertebrates (11,12). The EPH receptors the MAP kinase ERK. Conversely, siRNA based knockdown interact with a family of ligands located on the surfaces of of EPHB6 reversed ERK phosphorylation. Intriguingly, adjacent cells, named Ephrins including Ephrin-As and EPHB6-induced phosphorylation of ERK was uncoupled Ephrin-Bs subgroups. The EPH receptors are also grouped by activation of the Elk-1 transcriptional factor. -

A Review of VEGF/VEGFR-Targeted Therapeutics for Recurrent Glioblastoma
414 Original Article A Review of VEGF/VEGFR-Targeted Therapeutics for Recurrent Glioblastoma David A. Reardon, MDa,b; Scott Turner, MDc; Katherine B. Peters, MD, PhDc; Annick Desjardins, MDc; Sridharan Gururangan, MDa,b; John H. Sampson, MD, PhDa; Roger E. McLendon, MDd; James E. Herndon II, PhDe; Lee W. Jones, PhDf; John P. Kirkpatrick, MD, PhDf; Allan H. Friedman, MDa; James J. Vredenburgh, MDc; Darell D. Bigner, MD, PhDd; and Henry S. Friedman, MDa,b; Durham, North Carolina Key Words ability effect of VEGF inhibitors, the Radiologic Assessment in Neu- Glioblastoma, angiogenesis, vascular endothelial growth factor, ro-Oncology (RANO) criteria were recently implemented to bet- malignant glioma ter assess response among patients with glioblastoma. Although bevacizumab improves survival and quality of life, eventual tumor progression is the norm. Better understanding of resistance mech- Abstract anisms to VEGF inhibitors and identification of effective therapy Glioblastoma, the most common primary malignant brain tumor after bevacizumab progression are currently a critical need for pa- among adults, is a highly angiogenic and deadly tumor. Angiogen- tients with glioblastoma. (JNCCN 2011;9:414–427) esis in glioblastoma, driven by hypoxia-dependent and indepen- dent mechanisms, is primarily mediated by vascular endothelial growth factor (VEGF), and generates blood vessels with distinctive features. The outcome for patients with recurrent glioblastoma is poor because of ineffective therapies. However, recent encourag- ing rates of radiographic response and progression-free survival, Malignant gliomas, including the most common sub- and adequate safety, led the FDA to grant accelerated approval of type of glioblastoma, are rapidly growing destructive tu- bevacizumab, a humanized monoclonal antibody against VEGF, for mors that extensively invade locally but rarely metasta- the treatment of recurrent glioblastoma in May 2009. -

Profiling Data
Compound Name DiscoveRx Gene Symbol Entrez Gene Percent Compound Symbol Control Concentration (nM) JNK-IN-8 AAK1 AAK1 69 1000 JNK-IN-8 ABL1(E255K)-phosphorylated ABL1 100 1000 JNK-IN-8 ABL1(F317I)-nonphosphorylated ABL1 87 1000 JNK-IN-8 ABL1(F317I)-phosphorylated ABL1 100 1000 JNK-IN-8 ABL1(F317L)-nonphosphorylated ABL1 65 1000 JNK-IN-8 ABL1(F317L)-phosphorylated ABL1 61 1000 JNK-IN-8 ABL1(H396P)-nonphosphorylated ABL1 42 1000 JNK-IN-8 ABL1(H396P)-phosphorylated ABL1 60 1000 JNK-IN-8 ABL1(M351T)-phosphorylated ABL1 81 1000 JNK-IN-8 ABL1(Q252H)-nonphosphorylated ABL1 100 1000 JNK-IN-8 ABL1(Q252H)-phosphorylated ABL1 56 1000 JNK-IN-8 ABL1(T315I)-nonphosphorylated ABL1 100 1000 JNK-IN-8 ABL1(T315I)-phosphorylated ABL1 92 1000 JNK-IN-8 ABL1(Y253F)-phosphorylated ABL1 71 1000 JNK-IN-8 ABL1-nonphosphorylated ABL1 97 1000 JNK-IN-8 ABL1-phosphorylated ABL1 100 1000 JNK-IN-8 ABL2 ABL2 97 1000 JNK-IN-8 ACVR1 ACVR1 100 1000 JNK-IN-8 ACVR1B ACVR1B 88 1000 JNK-IN-8 ACVR2A ACVR2A 100 1000 JNK-IN-8 ACVR2B ACVR2B 100 1000 JNK-IN-8 ACVRL1 ACVRL1 96 1000 JNK-IN-8 ADCK3 CABC1 100 1000 JNK-IN-8 ADCK4 ADCK4 93 1000 JNK-IN-8 AKT1 AKT1 100 1000 JNK-IN-8 AKT2 AKT2 100 1000 JNK-IN-8 AKT3 AKT3 100 1000 JNK-IN-8 ALK ALK 85 1000 JNK-IN-8 AMPK-alpha1 PRKAA1 100 1000 JNK-IN-8 AMPK-alpha2 PRKAA2 84 1000 JNK-IN-8 ANKK1 ANKK1 75 1000 JNK-IN-8 ARK5 NUAK1 100 1000 JNK-IN-8 ASK1 MAP3K5 100 1000 JNK-IN-8 ASK2 MAP3K6 93 1000 JNK-IN-8 AURKA AURKA 100 1000 JNK-IN-8 AURKA AURKA 84 1000 JNK-IN-8 AURKB AURKB 83 1000 JNK-IN-8 AURKB AURKB 96 1000 JNK-IN-8 AURKC AURKC 95 1000 JNK-IN-8 -

Anti-Ephb6 (E5029)
Anti-EphB6 produced in goat, affinity isolated antibody Catalog Number E5029 Product Description do not induce receptor autophosphorylation and Anti-Mouse EphB6 is produced in goat using as activation.1 The ephrin ligands and Eph receptors immunogen a purified recombinant mouse EphB6, display reciprocal expression in vivo.2 Developing and extracellular domain, expressed in mouse NSO cells. adult neural tissue express nearly all of the Eph Affinity isolated antibody is obtained from goat anti- receptors and ephrin ligands.2 Ephs and ephrins play a EphB6 antiserum by immuno-specific purification which significant role in angiogenesis.2 removes essentially all goat serum proteins, including immunoglobulins, which do not specifically bind to the Reagent peptide. Supplied as a lyophilized powder from a 0.2 mm filtered solution of phosphate buffered saline (PBS), pH 7.4, Anti-Mouse EphB6 recognizes recombinant mouse with 5% trehalose. EphB6 by immunoblotting and direct ELISA. The antibody shows less than 1% cross-reactivity with Preparation Instructions recombinant mouse EphB2, recombinant mouse To one vial of lyophilized powder, add 0.5 ml of sterile EphB3, and recombinant mouse EphB4. phosphate buffered saline, pH 7.4, to produce a 0.2 mg/ml stock solution of antibody. EphB6, also known as Mep, is a member of the Eph receptor family, which binds members of the Ephrin Storage/Stability ligand family. Two classes of receptors exist, Prior to reconstitution, store at -20 °C. Reconstituted designated A and B, that have an extracellular domain product may be stored at 2-8 °C for up to one month. made up of a globular domain, a cysteine-rich domain, For prolonged storage, freeze in working aliquots at and two fibronectin type III domains, followed by the -20 °C for up to 6 months. -

Recombinant Human Epha8 Fc Chimera Catalog Number: 6828-A8
Recombinant Human EphA8 Fc Chimera Catalog Number: 6828-A8 DESCRIPTION Source Mouse myeloma cell line, NS0derived Human EphA8 Human IgG (Glu31Thr542) IEGRMD 1 (Pro100Lys330) Accession # NP_065387 Nterminus Cterminus Nterminal Sequence Glu31 Analysis Structure / Form Disulfidelinked homodimer Predicted Molecular 83.2 kDa (monomer) Mass SPECIFICATIONS SDSPAGE 94 kDa, reducing conditions Activity Measured by its binding ability in a functional ELISA. When Recombinant Human EphA8 Fc Chimera is coated at 2 μg/mL (100 μL/well), the concentration of Biotinylayed Recombinant Human EphrinA5 Fc Chimera (Catalog # BT374) that produces 50% of the optimal binding response is 212 ng/mL. Endotoxin Level <0.01 EU per 1 μg of the protein by the LAL method. Purity >90%, by SDSPAGE under reducing conditions and visualized by silver stain. Formulation Supplied as a 0.2 μm filtered solution in MES, NaCl, PEG, CHAPS and Imidazole. See Certificate of Analysis for details. PREPARATION AND STORAGE Shipping The product is shipped with dry ice or equivalent. Upon receipt, store it immediately at the temperature recommended below. Stability & Storage Use a manual defrost freezer and avoid repeated freezethaw cycles. l 12 months from date of receipt, 70 °C as supplied. l 1 month, 2 to 8 °C under sterile conditions after opening. l 3 months, 20 to 70 °C under sterile conditions after opening. BACKGROUND EphA8, also known as Hek3 and Eek, is a 120 kDa glycosylated member of the Eph family of transmembrane receptor tyrosine kinases (1, 2). The A and B classes of Eph proteins are distinguished by Ephrin ligand binding preference but have a common structural organization. -

A Tyrosine Kinase Expression Signature Predicts the Post-Operative Clinical Outcome in Triple Negative Breast Cancers
cancers Article A Tyrosine Kinase Expression Signature Predicts the Post-Operative Clinical Outcome in Triple Negative Breast Cancers Alexandre de Nonneville 1, Pascal Finetti 2 , José Adelaide 2, Éric Lambaudie 3, Patrice Viens 1, Anthony Gonçalves 1 , Daniel Birnbaum 2, Emilie Mamessier 2 and François Bertucci 1,2,* 1 Department of Medical Oncology, Institut Paoli-Calmettes, Aix-Marseille Univ, CRCM, CNRS, INSERM, 13000 Marseille, France 2 Laboratory of Predictive Oncology, Centre de Recherche en Cancérologie de Marseille, Institut Paoli-Calmettes, Inserm UMR1068, CNRS UMR725, Aix-Marseille Université, 13000 Marseille, France 3 Department of Surgical Oncology, Institut Paoli-Calmettes, Aix-Marseille Univ, CNRS, INSERM, CRCM, 13000 Marseille, France * Correspondence: [email protected]; Tel.: +33-4-91-22-35-37; Fax: +33-4-91-22-36-70 Received: 15 July 2019; Accepted: 9 August 2019; Published: 13 August 2019 Abstract: Triple negative breast cancer (TNBC) represent 15% of breast cancers. Histoclinical features and marketed prognostic gene expression signatures (GES) failed to identify good- and poor-prognosis patients. Tyrosine kinases (TK) represent potential prognostic and/or therapeutic targets for TNBC. We sought to define a prognostic TK GES in a large series of TNBC. mRNA expression and histoclinical data of 6379 early BCs were collected from 16 datasets. We searched for a TK-based GES associated with disease-free survival (DFS) and tested its robustness in an independent validation set. A total of 1226 samples were TNBC. In the learning set of samples (N = 825), we identified a 13-TK GES associated with DFS. This GES was associated with cell proliferation and immune response. -

Functional Analysis of Somatic Mutations Affecting Receptor Tyrosine Kinase Family in Metastatic Colorectal Cancer
Author Manuscript Published OnlineFirst on March 29, 2019; DOI: 10.1158/1535-7163.MCT-18-0582 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Functional analysis of somatic mutations affecting receptor tyrosine kinase family in metastatic colorectal cancer Leslie Duplaquet1, Martin Figeac2, Frédéric Leprêtre2, Charline Frandemiche3,4, Céline Villenet2, Shéhérazade Sebda2, Nasrin Sarafan-Vasseur5, Mélanie Bénozène1, Audrey Vinchent1, Gautier Goormachtigh1, Laurence Wicquart6, Nathalie Rousseau3, Ludivine Beaussire5, Stéphanie Truant7, Pierre Michel8, Jean-Christophe Sabourin9, Françoise Galateau-Sallé10, Marie-Christine Copin1,6, Gérard Zalcman11, Yvan De Launoit1, Véronique Fafeur1 and David Tulasne1 1 Univ. Lille, CNRS, Institut Pasteur de Lille, UMR 8161 - M3T – Mechanisms of Tumorigenesis and Target Therapies, F-59000 Lille, France. 2 Univ. Lille, Plateau de génomique fonctionnelle et structurale, CHU Lille, F-59000 Lille, France 3 TCBN - Tumorothèque Caen Basse-Normandie, F-14000 Caen, France. 4 Réseau Régional de Cancérologie – OncoBasseNormandie – F14000 Caen – France. 5 Normandie Univ, UNIROUEN, Inserm U1245, IRON group, Rouen University Hospital, Normandy Centre for Genomic and Personalized Medicine, F-76000 Rouen, France. 6 Tumorothèque du C2RC de Lille, F-59037 Lille, France. 7 Department of Digestive Surgery and Transplantation, CHU Lille, Univ Lille, 2 Avenue Oscar Lambret, 59037, Lille Cedex, France. 8 Department of hepato-gastroenterology, Rouen University Hospital, Normandie Univ, UNIROUEN, Inserm U1245, IRON group, F-76000 Rouen, France. 9 Department of Pathology, Normandy University, INSERM 1245, Rouen University Hospital, F 76 000 Rouen, France. 10 Department of Pathology, MESOPATH-MESOBANK, Centre León Bérard, Lyon, France. 11 Thoracic Oncology Department, CIC1425/CLIP2 Paris-Nord, Hôpital Bichat-Claude Bernard, Paris, France. -

Inhibition of Src Family Kinases and Receptor Tyrosine Kinases by Dasatinib: Possible Combinations in Solid Tumors
Published OnlineFirst June 13, 2011; DOI: 10.1158/1078-0432.CCR-10-2616 Clinical Cancer Molecular Pathways Research Inhibition of Src Family Kinases and Receptor Tyrosine Kinases by Dasatinib: Possible Combinations in Solid Tumors Juan Carlos Montero1, Samuel Seoane1, Alberto Ocaña2,3, and Atanasio Pandiella1 Abstract Dasatinib is a small molecule tyrosine kinase inhibitor that targets a wide variety of tyrosine kinases implicated in the pathophysiology of several neoplasias. Among the most sensitive dasatinib targets are ABL, the SRC family kinases (SRC, LCK, HCK, FYN, YES, FGR, BLK, LYN, and FRK), and the receptor tyrosine kinases c-KIT, platelet-derived growth factor receptor (PDGFR) a and b, discoidin domain receptor 1 (DDR1), c-FMS, and ephrin receptors. Dasatinib inhibits cell duplication, migration, and invasion, and it triggers apoptosis of tumoral cells. As a consequence, dasatinib reduces tumoral mass and decreases the metastatic dissemination of tumoral cells. Dasatinib also acts on the tumoral microenvironment, which is particularly important in the bone, where dasatinib inhibits osteoclastic activity and favors osteogenesis, exerting a bone-protecting effect. Several preclinical studies have shown that dasatinib potentiates the antitumoral action of various drugs used in the oncology clinic, paving the way for the initiation of clinical trials of dasatinib in combination with standard-of-care treatments for the therapy of various neoplasias. Trials using combinations of dasatinib with ErbB/HER receptor antagonists are being explored in breast, head and neck, and colorectal cancers. In hormone receptor–positive breast cancer, trials using combina- tions of dasatinib with antihormonal therapies are ongoing. Dasatinib combinations with chemother- apeutic agents are also under development in prostate cancer (dasatinib plus docetaxel), melanoma (dasatinib plus dacarbazine), and colorectal cancer (dasatinib plus oxaliplatin plus capecitabine). -

Ephrin-A Binding and Epha Receptor Expression Delineate the Matrix Compartment of the Striatum
The Journal of Neuroscience, June 15, 1999, 19(12):4962–4971 Ephrin-A Binding and EphA Receptor Expression Delineate the Matrix Compartment of the Striatum L. Scott Janis, Robert M. Cassidy, and Lawrence F. Kromer Department of Cell Biology and Interdisciplinary Program in Neuroscience, Georgetown University Medical Center, Washington, DC 20007 The striatum integrates limbic and neocortical inputs to regulate matrix neurons. In situ hybridization for EphA RTKs reveals that sensorimotor and psychomotor behaviors. This function is de- the two different ligand binding patterns strictly match pendent on the segregation of striatal projection neurons into the mRNA expression patterns of EphA4 and EphA7. anatomical and functional components, such as the striosome Ligand–receptor binding assays indicate that ephrin-A1 and and matrix compartments. In the present study the association ephrin-A4 selectively bind EphA4 but not EphA7 in the lysates of ephrin-A cell surface ligands and EphA receptor tyrosine of striatal tissue. Conversely, ephrin-A2, ephrin-A3, and kinases (RTKs) with the organization of these compartments ephrin-A5 bind EphA7 but not EphA4. These observations im- was determined in postnatal rats. Ephrin-A1 and ephrin-A4 plicate selective interactions between ephrin-A molecules and selectively bind to EphA receptors on neurons restricted to the EphA RTKs as potential mechanisms for regulating the com- matrix compartment. Binding is absent from the striosomes, partmental organization of the striatum. which were identified by m-opioid -

Supplementary Table 1. in Vitro Side Effect Profiling Study for LDN/OSU-0212320. Neurotransmitter Related Steroids
Supplementary Table 1. In vitro side effect profiling study for LDN/OSU-0212320. Percent Inhibition Receptor 10 µM Neurotransmitter Related Adenosine, Non-selective 7.29% Adrenergic, Alpha 1, Non-selective 24.98% Adrenergic, Alpha 2, Non-selective 27.18% Adrenergic, Beta, Non-selective -20.94% Dopamine Transporter 8.69% Dopamine, D1 (h) 8.48% Dopamine, D2s (h) 4.06% GABA A, Agonist Site -16.15% GABA A, BDZ, alpha 1 site 12.73% GABA-B 13.60% Glutamate, AMPA Site (Ionotropic) 12.06% Glutamate, Kainate Site (Ionotropic) -1.03% Glutamate, NMDA Agonist Site (Ionotropic) 0.12% Glutamate, NMDA, Glycine (Stry-insens Site) 9.84% (Ionotropic) Glycine, Strychnine-sensitive 0.99% Histamine, H1 -5.54% Histamine, H2 16.54% Histamine, H3 4.80% Melatonin, Non-selective -5.54% Muscarinic, M1 (hr) -1.88% Muscarinic, M2 (h) 0.82% Muscarinic, Non-selective, Central 29.04% Muscarinic, Non-selective, Peripheral 0.29% Nicotinic, Neuronal (-BnTx insensitive) 7.85% Norepinephrine Transporter 2.87% Opioid, Non-selective -0.09% Opioid, Orphanin, ORL1 (h) 11.55% Serotonin Transporter -3.02% Serotonin, Non-selective 26.33% Sigma, Non-Selective 10.19% Steroids Estrogen 11.16% 1 Percent Inhibition Receptor 10 µM Testosterone (cytosolic) (h) 12.50% Ion Channels Calcium Channel, Type L (Dihydropyridine Site) 43.18% Calcium Channel, Type N 4.15% Potassium Channel, ATP-Sensitive -4.05% Potassium Channel, Ca2+ Act., VI 17.80% Potassium Channel, I(Kr) (hERG) (h) -6.44% Sodium, Site 2 -0.39% Second Messengers Nitric Oxide, NOS (Neuronal-Binding) -17.09% Prostaglandins Leukotriene,