Casilio-ME: Enhanced CRISPR-Based DNA Demethylation by RNA-Guided
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Fine–Mapping Identifies NAD–ME1 As a Candidate Underlying a Major
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.07.285429; this version posted September 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Fine–mapping identifies NAD–ME1 as a candidate underlying 2 a major locus controlling temporal variation in primary and 3 specialized metabolism in Arabidopsis 4 5 Marta Francisco1, Daniel J. Kliebenstein2,3, Víctor M. Rodríguez1, Pilar Soengas1, 6 Rosaura Abilleira1, María E. Cartea1 7 1Misión Biológica de Galicia, (MBG-CSIC), P.O. Box 28, 36080, Pontevedra, Spain 8 2Department of Plant Sciences, University of California at Davis, Davis, CA 95616, 9 USA. 10 3DynaMo Center of Excellence, University of Copenhagen, Thorvaldsensvej 40, DK- 11 1871 Frederiksberg C, Denmark. 12 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.09.07.285429; this version posted September 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 13 Summary 14 Plant metabolism is modulated by a complex interplay between internal signals and 15 external cues. A major goal of all quantitative metabolomic studies is to clone the 16 underlying genes to understand the mechanistic basis of this variation. Using fine-scale 17 genetic mapping, in this work we report the identification and initial characterization of 18 NAD-DEPENDENT MALIC ENZYME 1 (NAD-ME1) as the candidate gene underlying 19 the pleiotropic network Met.II.15 QTL controlling variation in plant metabolism and 20 circadian clock outputs in the Bay × Sha Arabidopsis population. -

In Human Metabolism
Supporting Information (SI Appendix) Framework and resource for more than 11,000 gene-transcript- protein-reaction associations (GeTPRA) in human metabolism SI Appendix Materials and Methods Standardization of Metabolite IDs with MNXM IDs Defined in the MNXref Namespace. Information on metabolic contents of the Recon 2Q was standardized using MNXM IDs defined in the MNXref namespace available at MetaNetX (1-3). This standardization was to facilitate the model refinement process described below. Each metabolite ID in the Recon 2Q was converted to MNXM ID accordingly. For metabolite IDs that were not converted to MNXM IDs, they were manually converted to MNXM IDs by comparing their compound structures and synonyms. In the final resulting SBML files, 97 metabolites were assigned with arbitrary IDs (i.e., “MNXMK_” followed by four digits) because they were not covered by the MNXref namespace (i.e., metabolite IDs not converted to MNXM IDs). Refinement or Removal of Biochemically Inconsistent Reactions. Recon 2 was built upon metabolic genes and reactions collected from EHMN (4, 5), the first genome-scale human liver metabolic model HepatoNet1 (6), an acylcarnitine and fatty-acid oxidation model Ac-FAO (7), and a small intestinal enterocyte model hs_eIEC611 (8). Flux variability analysis (9) of the Recon 2Q identified blocked reactions coming from these four sources of metabolic reaction data. The EHMN caused the greatest number of blocked reactions in the Recon 2Q (1,070 reactions corresponding to 69.3% of all the identified blocked reactions). To refine the EHMN reactions, following reactions were initially disregarded: 1) reactions having metabolite IDs not convertible to MNXM IDs; and 2) reactions without genes. -

Profiling Cellular Processes in Adipose Tissue During Weight Loss Using Time Series Gene Expression
G C A T T A C G G C A T genes Article Profiling Cellular Processes in Adipose Tissue during Weight Loss Using Time Series Gene Expression Samar H. K. Tareen 1,* , Michiel E. Adriaens 1,* , Ilja C. W. Arts 1,2, Theo M. de Kok 1,3, Roel G. Vink 4, Nadia J. T. Roumans 4, Marleen A. van Baak 4, Edwin C. M. Mariman 4, Chris T. Evelo 1,5,* and Martina Kutmon 1,5,* 1 Maastricht Centre for Systems Biology (MaCSBio), Maastricht University, 6211ER Maastricht, The Netherlands; [email protected] (I.C.W.A.); [email protected] (T.M.d.K.) 2 Department of Epidemiology, CARIM School for Cardiovascular Diseases, Maastricht University, 6211ER Maastricht, The Netherlands 3 Department of Toxicogenomics, GROW School of Oncology and Developmental Biology, Maastricht University, 6211ER Maastricht, The Netherlands 4 Department of Human Biology, NUTRIM Research School, Maastricht University, 6211ER Maastricht, The Netherlands; [email protected] (R.G.V.); [email protected] (N.J.T.R.); [email protected] (M.A.v.B.); [email protected] (E.C.M.M.) 5 Department of Bioinformatics—BiGCaT, NUTRIM Research School, Maastricht University, 6211ER Maastricht, The Netherlands * Correspondence: [email protected] (S.H.K.T.); [email protected] (M.E.A.); [email protected] (C.T.E.); [email protected] (M.K.) Received: 28 September 2018; Accepted: 22 October 2018; Published: 29 October 2018 Abstract: Obesity is a global epidemic identified as a major risk factor for multiple chronic diseases and, consequently, diet-induced weight loss is used to counter obesity. -

Mitoxplorer, a Visual Data Mining Platform To
mitoXplorer, a visual data mining platform to systematically analyze and visualize mitochondrial expression dynamics and mutations Annie Yim, Prasanna Koti, Adrien Bonnard, Fabio Marchiano, Milena Dürrbaum, Cecilia Garcia-Perez, José Villaveces, Salma Gamal, Giovanni Cardone, Fabiana Perocchi, et al. To cite this version: Annie Yim, Prasanna Koti, Adrien Bonnard, Fabio Marchiano, Milena Dürrbaum, et al.. mitoXplorer, a visual data mining platform to systematically analyze and visualize mitochondrial expression dy- namics and mutations. Nucleic Acids Research, Oxford University Press, 2020, 10.1093/nar/gkz1128. hal-02394433 HAL Id: hal-02394433 https://hal-amu.archives-ouvertes.fr/hal-02394433 Submitted on 4 Dec 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License Nucleic Acids Research, 2019 1 doi: 10.1093/nar/gkz1128 Downloaded from https://academic.oup.com/nar/advance-article-abstract/doi/10.1093/nar/gkz1128/5651332 by Bibliothèque de l'université la Méditerranée user on 04 December 2019 mitoXplorer, a visual data mining platform to systematically analyze and visualize mitochondrial expression dynamics and mutations Annie Yim1,†, Prasanna Koti1,†, Adrien Bonnard2, Fabio Marchiano3, Milena Durrbaum¨ 1, Cecilia Garcia-Perez4, Jose Villaveces1, Salma Gamal1, Giovanni Cardone1, Fabiana Perocchi4, Zuzana Storchova1,5 and Bianca H. -

Enhanced CRISPR-Based DNA Demethylation by Casilio-ME-Mediated RNA-Guided Coupling of Methylcytosine Oxidation and DNA Repair Pathways
ARTICLE https://doi.org/10.1038/s41467-019-12339-7 OPEN Enhanced CRISPR-based DNA demethylation by Casilio-ME-mediated RNA-guided coupling of methylcytosine oxidation and DNA repair pathways Aziz Taghbalout1, Menghan Du1, Nathaniel Jillette1, Wojciech Rosikiewicz1, Abhijit Rath2, Christopher D. Heinen2, Sheng Li1 & Albert W. Cheng 1,3,4* Casilio-ME 1234567890():,; Here we develop a methylation editing toolbox, , that enables not only RNA-guided methylcytosine editing by targeting TET1 to genomic sites, but also by co-delivering TET1 and protein factors that couple methylcytosine oxidation to DNA repair activities, and/or promote TET1 to achieve enhanced activation of methylation-silenced genes. Delivery of TET1 activity by Casilio-ME1 robustly alters the CpG methylation landscape of promoter regions and activates methylation-silenced genes. We augment Casilio-ME1 to simultaneously deliver the TET1-catalytic domain and GADD45A (Casilio-ME2) or NEIL2 (Casilio-ME3) to streamline removal of oxidized cytosine intermediates to enhance activation of targeted genes. Using two-in-one effectors or modular effectors, Casilio-ME2 and Casilio-ME3 remarkably boost gene activation and methylcytosine demethylation of targeted loci. We expand the toolbox to enable a stable and expression-inducible system for broader application of the Casilio-ME platforms. This work establishes a platform for editing DNA methylation to enable research investigations interrogating DNA methylomes. 1 The Jackson Laboratory for Genomic Medicine, 10 Discovery Drive, Farmington, CT 06032, USA. 2 Center for Molecular Oncology, University of Connecticut Health, 263 Farmington Avenue, Farmington, CT 06030, USA. 3 Department of Genetics and Genome Sciences, University of Connecticut Health, 400 Farmington Avenue, Farmington, CT 06030, USA. -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

Mouse Me1 Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Me1 Knockout Project (CRISPR/Cas9) Objective: To create a Me1 knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Me1 gene (NCBI Reference Sequence: NM_001198933 ; Ensembl: ENSMUSG00000032418 ) is located on Mouse chromosome 9. 14 exons are identified, with the ATG start codon in exon 1 and the TAA stop codon in exon 14 (Transcript: ENSMUST00000185374). Exon 2~3 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a spontaneous allele exhibit decreased body weight on a medium fat diet, altered cytoplasmic malic enzyme activity, and a male-specific reduction in food intake on a high fat diet. Exon 2 starts from about 1.15% of the coding region. Exon 2~3 covers 17.15% of the coding region. The size of effective KO region: ~2089 bp. The KO region does not have any other known gene. Page 1 of 9 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 2 3 14 Legends Exon of mouse Me1 Knockout region Page 2 of 9 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of Exon 2 is aligned with itself to determine if there are tandem repeats. No significant tandem repeat is found in the dot plot matrix. So this region is suitable for PCR screening or sequencing analysis. -

Dual Specificity Phosphatases from Molecular Mechanisms to Biological Function
International Journal of Molecular Sciences Dual Specificity Phosphatases From Molecular Mechanisms to Biological Function Edited by Rafael Pulido and Roland Lang Printed Edition of the Special Issue Published in International Journal of Molecular Sciences www.mdpi.com/journal/ijms Dual Specificity Phosphatases Dual Specificity Phosphatases From Molecular Mechanisms to Biological Function Special Issue Editors Rafael Pulido Roland Lang MDPI • Basel • Beijing • Wuhan • Barcelona • Belgrade Special Issue Editors Rafael Pulido Roland Lang Biocruces Health Research Institute University Hospital Erlangen Spain Germany Editorial Office MDPI St. Alban-Anlage 66 4052 Basel, Switzerland This is a reprint of articles from the Special Issue published online in the open access journal International Journal of Molecular Sciences (ISSN 1422-0067) from 2018 to 2019 (available at: https: //www.mdpi.com/journal/ijms/special issues/DUSPs). For citation purposes, cite each article independently as indicated on the article page online and as indicated below: LastName, A.A.; LastName, B.B.; LastName, C.C. Article Title. Journal Name Year, Article Number, Page Range. ISBN 978-3-03921-688-8 (Pbk) ISBN 978-3-03921-689-5 (PDF) c 2019 by the authors. Articles in this book are Open Access and distributed under the Creative Commons Attribution (CC BY) license, which allows users to download, copy and build upon published articles, as long as the author and publisher are properly credited, which ensures maximum dissemination and a wider impact of our publications. The book as a whole is distributed by MDPI under the terms and conditions of the Creative Commons license CC BY-NC-ND. Contents About the Special Issue Editors .................................... -
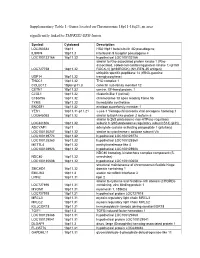
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

393LN V 393P 344SQ V 393P Probe Set Entrez Gene
393LN v 393P 344SQ v 393P Entrez fold fold probe set Gene Gene Symbol Gene cluster Gene Title p-value change p-value change chemokine (C-C motif) ligand 21b /// chemokine (C-C motif) ligand 21a /// chemokine (C-C motif) ligand 21c 1419426_s_at 18829 /// Ccl21b /// Ccl2 1 - up 393 LN only (leucine) 0.0047 9.199837 0.45212 6.847887 nuclear factor of activated T-cells, cytoplasmic, calcineurin- 1447085_s_at 18018 Nfatc1 1 - up 393 LN only dependent 1 0.009048 12.065 0.13718 4.81 RIKEN cDNA 1453647_at 78668 9530059J11Rik1 - up 393 LN only 9530059J11 gene 0.002208 5.482897 0.27642 3.45171 transient receptor potential cation channel, subfamily 1457164_at 277328 Trpa1 1 - up 393 LN only A, member 1 0.000111 9.180344 0.01771 3.048114 regulating synaptic membrane 1422809_at 116838 Rims2 1 - up 393 LN only exocytosis 2 0.001891 8.560424 0.13159 2.980501 glial cell line derived neurotrophic factor family receptor alpha 1433716_x_at 14586 Gfra2 1 - up 393 LN only 2 0.006868 30.88736 0.01066 2.811211 1446936_at --- --- 1 - up 393 LN only --- 0.007695 6.373955 0.11733 2.480287 zinc finger protein 1438742_at 320683 Zfp629 1 - up 393 LN only 629 0.002644 5.231855 0.38124 2.377016 phospholipase A2, 1426019_at 18786 Plaa 1 - up 393 LN only activating protein 0.008657 6.2364 0.12336 2.262117 1445314_at 14009 Etv1 1 - up 393 LN only ets variant gene 1 0.007224 3.643646 0.36434 2.01989 ciliary rootlet coiled- 1427338_at 230872 Crocc 1 - up 393 LN only coil, rootletin 0.002482 7.783242 0.49977 1.794171 expressed sequence 1436585_at 99463 BB182297 1 - up 393 -

The Role of H3K36 Methylation and Associated Methyltransferases In
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.04.433843; this version posted March 5, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 The role of H3K36 methylation and associated 2 methyltransferases in chromosome‐specific gene regulation 3 4 Henrik Lindehell1, Alexander Glotov1, Eshagh Dorafshan1, Yuri B. Schwartz1 and Jan Larsson1* 5 1Department of Molecular Biology, Umeå University, SE‐90187 Umeå, Sweden 6 7 *Corresponding author: Jan Larsson, Department of Molecular Biology, Umeå University, SE‐ 8 90187 Umeå, Sweden, Tel: +46 (0)90 7856785; Fax: +46 (0)90 778007; Email: 9 [email protected] 10 11 Data deposition footnote: The RNA‐seq data reported in this paper have been deposited in 12 the Gene Expression Omnibus database (GSE166934). 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.04.433843; this version posted March 5, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 13 Abstract 14 In Drosophila, two chromosomes require special mechanisms to balance their transcriptional 15 output to the rest of the genome. These are the male‐specific lethal complex targeting the 16 male X‐chromosome, and Painting of fourth targeting chromosome 4. The two systems are 17 evolutionarily linked to dosage compensation of the X‐chromosome and the chromosomes 18 involved display specific chromatin structures. Here we explore the role of histone H3 tri‐ 19 methylated at lysine 36 (H3K36me3) and the associated methyltransferases in these two 20 chromosome‐specific systems. -

Dissecting the Genetic Etiology of Lupus at ETS1 Locus
Dissecting the Genetic Etiology of Lupus at ETS1 Locus A dissertation submitted to the Graduate School of the University of Cincinnati in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Immunobiology of the College of Medicine 2017 by Xiaoming Lu B.S. Sun Yat-sen University, P.R. China June 2011 Dissertation Committee: John B. Harley, MD, PhD Harinder Singh, PhD Leah C. Kottyan, PhD Matthew T. Weirauch, PhD Kasper Hoebe, PhD Lili Ding, PhD i Abstract Systemic lupus erythematosus (SLE) is a complex autoimmune disease with strong evidence for genetics factor involvement. Genome-wide association studies have identified 84 risk loci associated with SLE. However, the specific genotype-dependent (allelic) molecular mechanisms connecting these lupus-genetic risk loci to immunological dysregulation are mostly still unidentified. ~ 90% of these loci contain variants that are non-coding, and are thus likely to act by impacting subtle, comparatively hard to predict mechanisms controlling gene expression. Here, we developed a strategic approach to prioritize non-coding variants, and screen them for their function. This approach involves computational prioritization using functional genomic databases followed by experimental analysis of differential binding of transcription factors (TFs) to risk and non-risk alleles. For both electrophoretic mobility shift assay (EMSA) and DNA affinity precipitation assay (DAPA) analysis of genetic variants, a synthetic DNA oligonucleotide (oligo) is used to identify factors in the nuclear lysate of disease or phenotype-relevant cells. This strategic approach was then used for investigating SLE association at ETS1 locus. Genetic variants at chromosomal region 11q23.3, near the gene ETS1, have been associated with systemic lupus erythematosus (SLE), or lupus, in independent cohorts of Asian ancestry.