In Human Metabolism
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mitoxplorer, a Visual Data Mining Platform To
mitoXplorer, a visual data mining platform to systematically analyze and visualize mitochondrial expression dynamics and mutations Annie Yim, Prasanna Koti, Adrien Bonnard, Fabio Marchiano, Milena Dürrbaum, Cecilia Garcia-Perez, José Villaveces, Salma Gamal, Giovanni Cardone, Fabiana Perocchi, et al. To cite this version: Annie Yim, Prasanna Koti, Adrien Bonnard, Fabio Marchiano, Milena Dürrbaum, et al.. mitoXplorer, a visual data mining platform to systematically analyze and visualize mitochondrial expression dy- namics and mutations. Nucleic Acids Research, Oxford University Press, 2020, 10.1093/nar/gkz1128. hal-02394433 HAL Id: hal-02394433 https://hal-amu.archives-ouvertes.fr/hal-02394433 Submitted on 4 Dec 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License Nucleic Acids Research, 2019 1 doi: 10.1093/nar/gkz1128 Downloaded from https://academic.oup.com/nar/advance-article-abstract/doi/10.1093/nar/gkz1128/5651332 by Bibliothèque de l'université la Méditerranée user on 04 December 2019 mitoXplorer, a visual data mining platform to systematically analyze and visualize mitochondrial expression dynamics and mutations Annie Yim1,†, Prasanna Koti1,†, Adrien Bonnard2, Fabio Marchiano3, Milena Durrbaum¨ 1, Cecilia Garcia-Perez4, Jose Villaveces1, Salma Gamal1, Giovanni Cardone1, Fabiana Perocchi4, Zuzana Storchova1,5 and Bianca H. -

Enhanced CRISPR-Based DNA Demethylation by Casilio-ME-Mediated RNA-Guided Coupling of Methylcytosine Oxidation and DNA Repair Pathways
ARTICLE https://doi.org/10.1038/s41467-019-12339-7 OPEN Enhanced CRISPR-based DNA demethylation by Casilio-ME-mediated RNA-guided coupling of methylcytosine oxidation and DNA repair pathways Aziz Taghbalout1, Menghan Du1, Nathaniel Jillette1, Wojciech Rosikiewicz1, Abhijit Rath2, Christopher D. Heinen2, Sheng Li1 & Albert W. Cheng 1,3,4* Casilio-ME 1234567890():,; Here we develop a methylation editing toolbox, , that enables not only RNA-guided methylcytosine editing by targeting TET1 to genomic sites, but also by co-delivering TET1 and protein factors that couple methylcytosine oxidation to DNA repair activities, and/or promote TET1 to achieve enhanced activation of methylation-silenced genes. Delivery of TET1 activity by Casilio-ME1 robustly alters the CpG methylation landscape of promoter regions and activates methylation-silenced genes. We augment Casilio-ME1 to simultaneously deliver the TET1-catalytic domain and GADD45A (Casilio-ME2) or NEIL2 (Casilio-ME3) to streamline removal of oxidized cytosine intermediates to enhance activation of targeted genes. Using two-in-one effectors or modular effectors, Casilio-ME2 and Casilio-ME3 remarkably boost gene activation and methylcytosine demethylation of targeted loci. We expand the toolbox to enable a stable and expression-inducible system for broader application of the Casilio-ME platforms. This work establishes a platform for editing DNA methylation to enable research investigations interrogating DNA methylomes. 1 The Jackson Laboratory for Genomic Medicine, 10 Discovery Drive, Farmington, CT 06032, USA. 2 Center for Molecular Oncology, University of Connecticut Health, 263 Farmington Avenue, Farmington, CT 06030, USA. 3 Department of Genetics and Genome Sciences, University of Connecticut Health, 400 Farmington Avenue, Farmington, CT 06030, USA. -

Protein Data Bank Contents Guide: Atomic Coordinate Entry Format
Protein Data Bank Contents Guide: Atomic Coordinate Entry Format Description Version 3.20 Document Published by the wwPDB This format complies with the PDB Exchange Dictionary (PDBx) http://mmcif.pdb.org/dictionaries/mmcif_pdbx.dic/Index/index.html. ©2008 wwPDB PDB File Format v. 3.2 Page i Table of Contents 1. Introduction................................................................................................................................... 1 Basic Notions of the Format Description ............................................................................................3 Record Format....................................................................................................................................5 Types of Records................................................................................................................................6 PDB Format Change Policy................................................................................................................9 Order of Records ..............................................................................................................................10 Sections of an Entry..........................................................................................................................12 Field Formats and Data Types .........................................................................................................14 2. Title Section............................................................................................................................... -

Accurate Detection of Fusion Genes and Transcription-Induced Chimeras from RNA-Seq Data
bioRxiv preprint doi: https://doi.org/10.1101/070888; this version posted August 22, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. ChimPipe: Accurate detection of fusion genes and transcription-induced chimeras from RNA-seq data Bernardo Rodr´ıguez-Mart´ın1,2,3, Emilio Palumbo1,2, Santiago Marco-Sola4, Thasso Griebel4, Paolo Ribeca4,5, Graciela Alonso6, Alberto Rastrojo6, Begona˜ Aguado6, Roderic Guigo´ 1,2,7 and Sarah Djebali*1,2,8 1Centre for Genomic Regulation (CRG), The Barcelona Institute of Science and Technology, Dr. Aiguader 88, 08003 Barcelona, Spain 2Universitat Pompeu Fabra (UPF), Barcelona, Spain 3Joint IRB-BSC Program in Computational Biology, Barcelona Supercomputing center (BSC), Jordi Girona 31, 08034 Barcelona, Spain 4Centro Nacional de Analisis´ Genomico,´ Baldiri Reixac, 4, Barcelona Science Park - Tower I, 08028 Barcelona, Spain 5Integrative Biology, The Pirbright Institute, Ash Road, Pirbright, Woking, GU24 0NF, United Kingdom 6Centro de Biolog´ıa Molecular Severo Ochoa (CSIC - UAM), Nicolas´ Cabrera 1, Cantoblanco, 28049 Madrid, Spain 7Institut Hospital del Mar d’Investigacions Mediques (IMIM), 08003 Barcelona, Spain 8GenPhySE, Universite´ de Toulouse, INRA, INPT, ENVT, Castanet 1 bioRxiv preprint doi: https://doi.org/10.1101/070888; this version posted August 22, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. -

(Slx-2119), a ROCK2-Specific Inhibitor, in 3T3-L1 Cells
www.nature.com/scientificreports OPEN Anti-adipogenic efects of KD025 (SLx-2119), a ROCK2-specifc inhibitor, in 3T3-L1 cells Received: 6 January 2017 Duy Trong Vien Diep1, Kyungki Hong1, Triyeng Khun1, Mei Zheng 1, Asad ul-Haq1, Accepted: 24 January 2018 Hee-Sook Jun1,3,4, Young-Bum Kim2,3 & Kwang-Hoon Chun1 Published: xx xx xxxx Adipose tissue is a specialized organ that synthesizes and stores fat. During adipogenesis, Rho and Rho- associated kinase (ROCK) 2 are inactivated, which enhances the expression of pro-adipogenic genes and induces the loss of actin stress fbers. Furthermore, pan ROCK inhibitors enhance adipogenesis in 3T3-L1 cells. Here, we show that KD025 (formerly known as SLx-2119), a ROCK2-specifc inhibitor, suppresses adipogenesis in 3T3-L1 cells partially through a ROCK2-independent mechanism. KD025 downregulated the expression of key adipogenic transcription factors PPARγ and C/EBPα during adipogenesis in addition to lipogenic factors FABP4 and Glut4. Interestingly, adipogenesis was blocked by KD025 during days 1~3 of diferentiation; after diferentiation terminated, lipid accumulation was unafected. Clonal expansion occurred normally in KD025-treated cells. These results suggest that KD025 could function during the intermediate stage after clonal expansion. Data from depletion of ROCKs showed that KD025 suppressed cell diferentiation partially independent of ROCK’s activity. Furthermore, no further loss of actin stress fbers emerged in KD025-treated cells during and after diferentiation compared to control cells. These results indicate that in contrast to the pro-adipogenic efect of pan-inhibitors, KD025 suppresses adipogenesis in 3T3-L1 cells by regulating key pro- adipogenic factors. -

Rabbit Anti-GEM/FITC Conjugated Antibody-SL13331R-FITC
SunLong Biotech Co.,LTD Tel: 0086-571- 56623320 Fax:0086-571- 56623318 E-mail:[email protected] www.sunlongbiotech.com Rabbit Anti-GEM/FITC Conjugated antibody SL13331R-FITC Product Name: Anti-GEM/FITC Chinese Name: FITC标记的GTP结合丝裂原诱导T细胞蛋白抗体 GTP binding mitogen induced T cell protein; GTP binding protein expressed in mitogen stimulated T cells; GTP binding protein GEM; GTP binding protein overexpressed in Alias: skeletal muscle; Kinase inducible Ras like protein; KIR; MGC26294; RAS like protein KIR; GEM_HUMAN. Organism Species: Rabbit Clonality: Polyclonal React Species: Human,Mouse,Rat,Dog,Pig,Cow,Horse, ICC=1:50-200IF=1:50-200 Applications: not yet tested in other applications. optimal dilutions/concentrations should be determined by the end user. Molecular weight: 34kDa Cellular localization: The cell membrane Form: Lyophilized or Liquid Concentration: 1mg/ml immunogen: KLH conjugated synthetic peptide derived from human GEM Lsotype: IgGwww.sunlongbiotech.com Purification: affinity purified by Protein A Storage Buffer: 0.01M TBS(pH7.4) with 1% BSA, 0.03% Proclin300 and 50% Glycerol. Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. The lyophilized antibody is stable at room temperature for at least one month and for greater than a year Storage: when kept at -20°C. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 °C. background: Gem belongs to the Rad/Gem/Kir (RGK) subfamily of Ras-related GTPases, which lack typical C-terminal amino acid motifs for isoprenylation. Rad and Gem bind calmodulin Product Detail: in a Ca2+-dependent manner via this C-terminal extension, involving residues 278–297 in human Rad. -

Mouse Pathology Spontaneous Phenotypes and Diseases (NON
2017 Brayton MICE NON infectious 32p OPC Disclosures Mouse Pathology No financial disclosures that I know of… Spontaneous Phenotypes All opinions expressed and implied in and Diseases (NON infectious) this presentation are solely those of Dr. Brayton. The content of the presentation does Cory Brayton, DVM, DipACLAM, DipACVP not represent or reflect the views of Associate Professor, Molecular and Comparative Pathobiology Director, Phenotyping Core Johns Hopkins University or Johns Johns Hopkins University, School of Medicine Hopkins Health system. Baltimore, MD 21205 [email protected] http://www.hopkinsmedicine.org/mcp/PHENOCORE/index.html 3 General References Resources online 1. ACLAM Blue books https://www.elsevier.com/books/book‐series/american‐college‐of‐ laboratory‐animal‐medicine 1. Brayton: Spontaneous diseases in commonly used mouse strains and stocks (2013) http://phenome.jax.org/projects/Brayton1 – The Mouse in Biomedical Research. Fox, Barthold, Davisson et al. Vol I‐IV 2006 – Lab manual at http://www.hopkinsmedicine.org/mcp/PHENOCORE/courseCURRENT.html – Laboratory Animal Medicine Fox et al 2015 2. DORA Diseases of Research Animals http://dora.missouri.edu/ 2. Barthold S, Griffey S, Percy D. 2016. Pathology of Laboratory Rodents and Rabbits, Wiley‐ Blackwell. 3. Frith and Ward. 1988. A Color Atlas of Neoplastic and Non Neoplastic Lesions in Aging Mice. http://www.informatics.jax.org/frithbook/ also print on Demand from 3. Brayton C. 2006. Spontaneous Diseases in Commonly Used Mouse Strains The Mouse in http://store.cldavis.org/ Biomedical Research. Fox, Barthold, Davisson et al. New York, Elsevier (Academic Press). II. Diseases: 623‐717. outline at http://phenome.jax.org/projects/Brayton1 4. -
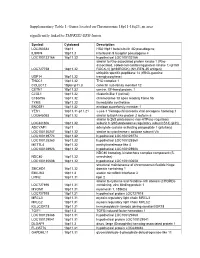
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

393LN V 393P 344SQ V 393P Probe Set Entrez Gene
393LN v 393P 344SQ v 393P Entrez fold fold probe set Gene Gene Symbol Gene cluster Gene Title p-value change p-value change chemokine (C-C motif) ligand 21b /// chemokine (C-C motif) ligand 21a /// chemokine (C-C motif) ligand 21c 1419426_s_at 18829 /// Ccl21b /// Ccl2 1 - up 393 LN only (leucine) 0.0047 9.199837 0.45212 6.847887 nuclear factor of activated T-cells, cytoplasmic, calcineurin- 1447085_s_at 18018 Nfatc1 1 - up 393 LN only dependent 1 0.009048 12.065 0.13718 4.81 RIKEN cDNA 1453647_at 78668 9530059J11Rik1 - up 393 LN only 9530059J11 gene 0.002208 5.482897 0.27642 3.45171 transient receptor potential cation channel, subfamily 1457164_at 277328 Trpa1 1 - up 393 LN only A, member 1 0.000111 9.180344 0.01771 3.048114 regulating synaptic membrane 1422809_at 116838 Rims2 1 - up 393 LN only exocytosis 2 0.001891 8.560424 0.13159 2.980501 glial cell line derived neurotrophic factor family receptor alpha 1433716_x_at 14586 Gfra2 1 - up 393 LN only 2 0.006868 30.88736 0.01066 2.811211 1446936_at --- --- 1 - up 393 LN only --- 0.007695 6.373955 0.11733 2.480287 zinc finger protein 1438742_at 320683 Zfp629 1 - up 393 LN only 629 0.002644 5.231855 0.38124 2.377016 phospholipase A2, 1426019_at 18786 Plaa 1 - up 393 LN only activating protein 0.008657 6.2364 0.12336 2.262117 1445314_at 14009 Etv1 1 - up 393 LN only ets variant gene 1 0.007224 3.643646 0.36434 2.01989 ciliary rootlet coiled- 1427338_at 230872 Crocc 1 - up 393 LN only coil, rootletin 0.002482 7.783242 0.49977 1.794171 expressed sequence 1436585_at 99463 BB182297 1 - up 393 -

The Role of H3K36 Methylation and Associated Methyltransferases In
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.04.433843; this version posted March 5, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 The role of H3K36 methylation and associated 2 methyltransferases in chromosome‐specific gene regulation 3 4 Henrik Lindehell1, Alexander Glotov1, Eshagh Dorafshan1, Yuri B. Schwartz1 and Jan Larsson1* 5 1Department of Molecular Biology, Umeå University, SE‐90187 Umeå, Sweden 6 7 *Corresponding author: Jan Larsson, Department of Molecular Biology, Umeå University, SE‐ 8 90187 Umeå, Sweden, Tel: +46 (0)90 7856785; Fax: +46 (0)90 778007; Email: 9 [email protected] 10 11 Data deposition footnote: The RNA‐seq data reported in this paper have been deposited in 12 the Gene Expression Omnibus database (GSE166934). 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.04.433843; this version posted March 5, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 13 Abstract 14 In Drosophila, two chromosomes require special mechanisms to balance their transcriptional 15 output to the rest of the genome. These are the male‐specific lethal complex targeting the 16 male X‐chromosome, and Painting of fourth targeting chromosome 4. The two systems are 17 evolutionarily linked to dosage compensation of the X‐chromosome and the chromosomes 18 involved display specific chromatin structures. Here we explore the role of histone H3 tri‐ 19 methylated at lysine 36 (H3K36me3) and the associated methyltransferases in these two 20 chromosome‐specific systems. -

New Mechanisms of Resistance to MEK Inhibitors in Melanoma Revealed by Intravital Imaging
Author Manuscript Published OnlineFirst on November 27, 2017; DOI: 10.1158/0008-5472.CAN-17-1653 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. New mechanisms of resistance to MEK inhibitors in melanoma revealed by intravital imaging Hailey E. Brighton1,2,+; Steven P. Angus3,+; Tao Bo1; Jose Roques1; Alicia C. Tagliatela1,2; David B. Darr1; Kubra Karagoz4, Noah Sciaky3, Michael L. Gatza4 Norman E. Sharpless1,5,6; Gary L. Johnson1,3; James E. Bear*1,2,3 + equal contribution 1UNC Lineberger Comprehensive Cancer Center, UNC Chapel Hill, Chapel Hill, NC, USA 2Department of Cell Biology and Physiology, UNC Chapel Hill, Chapel Hill, NC, USA 3Department of Pharmacology, UNC Chapel Hill, Chapel Hill, NC, USA 4Department of Radiation Oncology, Rutgers Cancer Institute of New Jersey, New Brunswick, NJ, USA 5Department of Medicine, UNC Chapel Hill, Chapel Hill, NC, USA 6Department of Genetics, UNC Chapel Hill, Chapel Hill, NC, USA *Corresponding Author: James E. Bear UNC Lineberger Comprehensive Cancer Center CB 7295 Chapel Hill, NC 27599 Phone: 919-966-5471 [email protected] Running Title: Intravital Imaging of Melanoma Growth and Response to MAPKi Abbreviations List: GEM: genetically engineered mouse, CL: collagen, ECM: extracellular matrix, SHG: second harmonic generation, 4-HT: 4-Hydroxytamoxifen, PBT: Tyr::CreER; BRAFCA;PTENlox/lox GEM model, wpa: weeks post-application, dpa: days post-application, MT: masson’s trichrome, TRAMETINIB&E: Hematoxylin and Eosin staining, RTK: receptor tyrosine kinase, CI: confidence interval, MIB/MS: multiplexed-inhibitor bead affinity chromatography coupled with mass spectrometry, EMT: epithelial to mesenchymal transition, M-E-T: mesenchymal to epithelial transition, WB: western blotting, IHC: immunohistochemistry Financial Support: This work was supported by HHMI funds to J.E. -

16P11.2 Transcription Factor MAZ Is a Dosage-Sensitive Regulator Of
16p11.2 transcription factor MAZ is a dosage-sensitive PNAS PLUS regulator of genitourinary development Meade Hallera,b,c,1, Jason Aub,c, Marisol O’Neilla,b,c, and Dolores J. Lamba,b,c aDepartment of Molecular and Cellular Biology, Baylor College of Medicine, Houston, TX 77030; bCenter for Reproductive Medicine, Baylor College of Medicine, Houston, TX 77030; and cDepartment of Urology, Baylor College of Medicine, Houston, TX 77030 Edited by Frank Costantini, Columbia University, New York, NY, and accepted by Editorial Board Member Kathryn V. Anderson January 12, 2018 (received for review September 12, 2017) Genitourinary (GU) birth defects are among the most common yet novo or inherited, and their allele frequency is influenced by both least studied congenital malformations. Congenital anomalies of the the rate of their spontaneous occurrence as well as their compati- kidney and urinary tract (CAKUTs) have high morbidity and mortality bility with life and with fertility (6). While many CNVs are consid- rates and account for ∼30% of structural birth defects. Copy number ered benign (7), specific CNVs have well-documented associations variation (CNV) mapping revealed that 16p11.2 is a hotspot for GU with a wide variety of birth defects (8). The specific congenital development. The only gene covered collectivelybyallofthemapped malformationsassociatedwitheachpathogenicCNVdependonthe GU-patient CNVs was MYC-associated zinc finger transcription factor specific genes disrupted within the genomic loci and whether the (MAZ), and MAZ CNV frequency is enriched in nonsyndromic GU- CNV is a duplication or a deletion. A CNV is considered syndromic abnormal patients. Knockdown of MAZ in HEK293 cells results in if multiple organs fail to develop normally in affected patients.