Network Propagation-Based Prioritization of Long Tail Genes In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

POLR2L Antibody Cat
POLR2L Antibody Cat. No.: XW-7445 POLR2L Antibody Specifications HOST SPECIES: Chicken SPECIES REACTIVITY: Human, Mouse, Rat IMMUNOGEN: 1-67 TESTED APPLICATIONS: WB POLR2L antibody can be used for the detection of POLR2L by Western blot, may also work APPLICATIONS: for IHC and ICC. PREDICTED MOLECULAR 7.6 kDa (calculated) WEIGHT: Properties PURIFICATION: Antigen affinity-purified CLONALITY: Polyclonal CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: Phosphate-Buffered Saline. No preservatives added. CONCENTRATION: 1 mg/mL October 1, 2021 1 https://www.prosci-inc.com/polr2l-antibody-7445.html POLR2L antibody can be stored at 4˚C for short term (weeks). Long term storage should STORAGE CONDITIONS: be at -20˚C. As with all antibodies care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. Additional Info OFFICIAL SYMBOL: POLR2L DNA-directed RNA polymerases I, II, and III subunit RPABC5, DNA-directed RNA ALTERNATE NAMES: polymerase III subunit L, RNA polymerases I, and III subunit ABC5, RBP10, RPB10, RPABC5, RPB7.6, hRPB7.6, RPB10beta, POLR2L ACCESSION NO.: NP_066951.1 PROTEIN GI NO.: 10863925 GENE ID: 5441 USER NOTE: Optimal dilutions for each application to be determined by the researcher. Background and References DNA directed RNA polymerase II polypeptide L; polymerase (RNA) II (DNA directed) polypeptide L (7.6kD); RNA polymerase II subunit. This protein is a subunit of RNA polymerase II, the polymerase responsible for synthesizing messenger RNA in eukaryotes. BACKGROUND: It contains four conserved cysteines characteristic of an atypical zinc-binding domain. Like its counterpart in yeast, this subunit may be shared by the other two DNA-directed RNA polymerases. -

A Network Propagation Approach to Prioritize Long Tail Genes in Cancer
bioRxiv preprint doi: https://doi.org/10.1101/2021.02.05.429983; this version posted February 8, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. A Network Propagation Approach to Prioritize Long Tail Genes in Cancer Hussein Mohsen1,*, Vignesh Gunasekharan2, Tao Qing2, Sahand Negahban3, Zoltan Szallasi4, Lajos Pusztai2,*, Mark B. Gerstein1,5,6,3,* 1 Computational Biology & Bioinformatics Program, Yale University, New Haven, CT 06511, USA 2 Breast Medical Oncology, Yale School of Medicine, New Haven, CT 06511, USA 3 Department of Statistics & Data Science, Yale University, New Haven, CT 06511, USA 4 Children’s Hospital Informatics Program, Harvard-MIT Division of Health Sciences and Technology, Harvard Medical School, Boston, MA 02115, USA 5 Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06511, USA 6 Department of Computer Science, Yale University, New Haven, CT 06511, USA * Corresponding author Abstract Introduction. The diversity of genomic alterations in cancer pose challenges to fully understanding the etiologies of the disease. Recent interest in infrequent mutations, in genes that reside in the “long tail” of the mutational distribution, uncovered new genes with significant implication in cancer development. The study of these genes often requires integrative approaches with multiple types of biological data. Network propagation methods have demonstrated high efficacy in uncovering genomic patterns underlying cancer using biological interaction networks. Yet, the majority of these analyses have focused their assessment on detecting known cancer genes or identifying altered subnetworks. -

SLFN11 Promotes CDT1 Degradation by CUL4 in Response to Replicative DNA Damage, While Its Absence Leads to Synthetic Lethality with ATR/CHK1 Inhibitors
SLFN11 promotes CDT1 degradation by CUL4 in response to replicative DNA damage, while its absence leads to synthetic lethality with ATR/CHK1 inhibitors Ukhyun Joa,1, Yasuhisa Muraia, Sirisha Chakkab, Lu Chenb, Ken Chengb, Junko Muraic, Liton Kumar Sahaa, Lisa M. Miller Jenkinsd, and Yves Pommiera,1 aDevelopmental Therapeutics Branch, Laboratory of Molecular Pharmacology, Center for Cancer Research, National Cancer Institute, Bethesda, MD 20814; bNational Center for Advancing Translational Sciences, Functional Genomics Laboratory, NIH, Rockville, MD 20850; cInstitute for Advanced Biosciences, Keio University, 997-0052 Yamagata, Japan; and dLaboratory of Cell Biology, Center for Cancer Research, National Cancer Institute, NIH, Bethesda, MD 20892 Edited by Richard D. Kolodner, Ludwig Institute for Cancer Research, La Jolla, CA, and approved December 8, 2020 (received for review July 29, 2020) Schlafen-11 (SLFN11) inactivation in ∼50% of cancer cells confers condensation related to deposition of H3K27me3 in the gene broad chemoresistance. To identify therapeutic targets and under- body of SLFN11 by EZH2, a histone methyltransferase (11). lying molecular mechanisms for overcoming chemoresistance, we Targeting epigenetic regulators is therefore an attractive com- performed an unbiased genome-wide RNAi screen in SLFN11-WT bination strategy to overcome chemoresistance of SLFN11- and -knockout (KO) cells. We found that inactivation of Ataxia deficient cancers (10, 25, 26). An alternative approach is to at- Telangiectasia- and Rad3-related (ATR), CHK1, BRCA2, and RPA1 tack SLFN11-negative cancer cells by targeting the essential SLFN11 overcome chemoresistance to camptothecin (CPT) in -KO pathways that cells use to overcome replicative damage and cells. Accordingly, we validate that clinical inhibitors of ATR replication stress. -

Molecular Pharmacology of Cancer Therapy in Human Colorectal Cancer by Gene Expression Profiling1,2
[CANCER RESEARCH 63, 6855–6863, October 15, 2003] Molecular Pharmacology of Cancer Therapy in Human Colorectal Cancer by Gene Expression Profiling1,2 Paul A. Clarke,3 Mark L. George, Sandra Easdale, David Cunningham, R. Ian Swift, Mark E. Hill, Diana M. Tait, and Paul Workman Cancer Research UK Centre for Cancer Therapeutics, Institute of Cancer Research, Sutton, Surrey SM2 5NG [P. A. C., M. L. G., S. E., P. W.]; Department of Gastrointestinal Oncology, Royal Marsden Hospital, Sutton, Surrey [D. C., M. E. H., D. M. T.]; and Department of Surgery, Mayday Hospital, Croydon, Surrey [M. L. G., R. I. S.], United Kingdom ABSTRACT ment with a single dose of MMC4 and during a continuous infusion of 5FU. In this study, we report for the first time gene expression Global gene expression profiling has potential for elucidating the com- profiling in cancer patients before, and critically, during the period of plex cellular effects and mechanisms of action of novel targeted anticancer exposure to chemotherapy. We have demonstrated that the approach agents or existing chemotherapeutics for which the precise molecular is feasible, and we have detected a novel molecular response that mechanism of action may be unclear. In this study, decreased expression would not have been predicted from in vitro studies and that would of genes required for RNA and protein synthesis, and for metabolism were have otherwise been missed by conventional approaches. The results detected in rectal cancer biopsies taken from patients during a 5-fluorou- also suggest a possible new therapeutic approach. Overall our obser- racil infusion. Our observations demonstrate that this approach is feasible and can detect responses that may have otherwise been missed by con- vations suggest that gene expression profiling in response to treatment ventional methods. -
Attachment PDF Icon
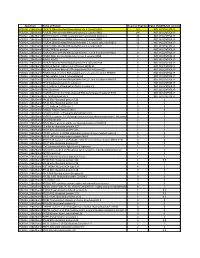
Spectrum Name of Protein Count of Peptides Ratio (POL2RA/IgG control) POLR2A_228kdBand POLR2A DNA-directed RNA polymerase II subunit RPB1 197 NOT IN CONTROL IP POLR2A_228kdBand POLR2B DNA-directed RNA polymerase II subunit RPB2 146 NOT IN CONTROL IP POLR2A_228kdBand RPAP2 Isoform 1 of RNA polymerase II-associated protein 2 24 NOT IN CONTROL IP POLR2A_228kdBand POLR2G DNA-directed RNA polymerase II subunit RPB7 23 NOT IN CONTROL IP POLR2A_228kdBand POLR2H DNA-directed RNA polymerases I, II, and III subunit RPABC3 19 NOT IN CONTROL IP POLR2A_228kdBand POLR2C DNA-directed RNA polymerase II subunit RPB3 17 NOT IN CONTROL IP POLR2A_228kdBand POLR2J RPB11a protein 7 NOT IN CONTROL IP POLR2A_228kdBand POLR2E DNA-directed RNA polymerases I, II, and III subunit RPABC1 8 NOT IN CONTROL IP POLR2A_228kdBand POLR2I DNA-directed RNA polymerase II subunit RPB9 9 NOT IN CONTROL IP POLR2A_228kdBand ALMS1 ALMS1 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2D DNA-directed RNA polymerase II subunit RPB4 6 NOT IN CONTROL IP POLR2A_228kdBand GRINL1A;Gcom1 Isoform 12 of Protein GRINL1A 6 NOT IN CONTROL IP POLR2A_228kdBand RECQL5 Isoform Beta of ATP-dependent DNA helicase Q5 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2L DNA-directed RNA polymerases I, II, and III subunit RPABC5 5 NOT IN CONTROL IP POLR2A_228kdBand KRT6A Keratin, type II cytoskeletal 6A 3 NOT IN CONTROL IP POLR2A_228kdBand POLR2K DNA-directed RNA polymerases I, II, and III subunit RPABC4 2 NOT IN CONTROL IP POLR2A_228kdBand RFC4 Replication factor C subunit 4 1 NOT IN CONTROL IP POLR2A_228kdBand RFC2 -

MAF1 Represses CDKN1A Through a Pol III-Dependent Mechanism Yu-Ling Lee, Yuan-Ching Li, Chia-Hsin Su, Chun-Hui Chiao, I-Hsuan Lin, Ming-Ta Hsu*
RESEARCH ARTICLE elifesciences.org MAF1 represses CDKN1A through a Pol III-dependent mechanism Yu-Ling Lee, Yuan-Ching Li, Chia-Hsin Su, Chun-Hui Chiao, I-Hsuan Lin, Ming-Ta Hsu* Institute of Biochemistry and Molecular Biology, School of Life Science, National Yang-Ming University, Taipei, Taiwan Abstract MAF1 represses Pol III-mediated transcription by interfering with TFIIIB and Pol III. Herein, we found that MAF1 knockdown induced CDKN1A transcription and chromatin looping concurrently with Pol III recruitment. Simultaneous knockdown of MAF1 with Pol III or BRF1 (subunit of TFIIIB) diminished the activation and looping effect, which indicates that recruiting Pol III was required for activation of Pol II-mediated transcription and chromatin looping. Chromatin- immunoprecipitation analysis after MAF1 knockdown indicated enhanced binding of Pol III and BRF1, as well as of CFP1, p300, and PCAF, which are factors that mediate active histone marks, along with the binding of TATA binding protein (TBP) and POLR2E to the CDKN1A promoter. Simultaneous knockdown with Pol III abolished these regulatory events. Similar results were obtained for GDF15. Our results reveal a novel mechanism by which MAF1 and Pol III regulate the activity of a protein- coding gene transcribed by Pol II. DOI: 10.7554/eLife.06283.001 Introduction Transcription by RNA polymerase III (Pol III) is regulated by MAF1, which is a highly conserved protein in eukaryotes (Pluta et al., 2001; Reina et al., 2006). MAF1 represses Pol III transcription through *For correspondence: mth@ym. association with BRF1, a subunit of initiation factor TFIIIB, which prevents attachment of TFIIIB onto edu.tw DNA. -

The Human Genome Project
TO KNOW OURSELVES ❖ THE U.S. DEPARTMENT OF ENERGY AND THE HUMAN GENOME PROJECT JULY 1996 TO KNOW OURSELVES ❖ THE U.S. DEPARTMENT OF ENERGY AND THE HUMAN GENOME PROJECT JULY 1996 Contents FOREWORD . 2 THE GENOME PROJECT—WHY THE DOE? . 4 A bold but logical step INTRODUCING THE HUMAN GENOME . 6 The recipe for life Some definitions . 6 A plan of action . 8 EXPLORING THE GENOMIC LANDSCAPE . 10 Mapping the terrain Two giant steps: Chromosomes 16 and 19 . 12 Getting down to details: Sequencing the genome . 16 Shotguns and transposons . 20 How good is good enough? . 26 Sidebar: Tools of the Trade . 17 Sidebar: The Mighty Mouse . 24 BEYOND BIOLOGY . 27 Instrumentation and informatics Smaller is better—And other developments . 27 Dealing with the data . 30 ETHICAL, LEGAL, AND SOCIAL IMPLICATIONS . 32 An essential dimension of genome research Foreword T THE END OF THE ROAD in Little has been rapid, and it is now generally agreed Cottonwood Canyon, near Salt that this international project will produce Lake City, Alta is a place of the complete sequence of the human genome near-mythic renown among by the year 2005. A skiers. In time it may well And what is more important, the value assume similar status among molecular of the project also appears beyond doubt. geneticists. In December 1984, a conference Genome research is revolutionizing biology there, co-sponsored by the U.S. Department and biotechnology, and providing a vital of Energy, pondered a single question: Does thrust to the increasingly broad scope of the modern DNA research offer a way of detect- biological sciences. -

Functional Dependency Analysis Identifies Potential Druggable
cancers Article Functional Dependency Analysis Identifies Potential Druggable Targets in Acute Myeloid Leukemia 1, 1, 2 3 Yujia Zhou y , Gregory P. Takacs y , Jatinder K. Lamba , Christopher Vulpe and Christopher R. Cogle 1,* 1 Division of Hematology and Oncology, Department of Medicine, College of Medicine, University of Florida, Gainesville, FL 32610-0278, USA; yzhou1996@ufl.edu (Y.Z.); gtakacs@ufl.edu (G.P.T.) 2 Department of Pharmacotherapy and Translational Research, College of Pharmacy, University of Florida, Gainesville, FL 32610-0278, USA; [email protected]fl.edu 3 Department of Physiological Sciences, College of Veterinary Medicine, University of Florida, Gainesville, FL 32610-0278, USA; cvulpe@ufl.edu * Correspondence: [email protected]fl.edu; Tel.: +1-(352)-273-7493; Fax: +1-(352)-273-5006 Authors contributed equally. y Received: 3 November 2020; Accepted: 7 December 2020; Published: 10 December 2020 Simple Summary: New drugs are needed for treating acute myeloid leukemia (AML). We analyzed data from genome-edited leukemia cells to identify druggable targets. These targets were necessary for AML cell survival and had favorable binding sites for drug development. Two lists of genes are provided for target validation, drug discovery, and drug development. The deKO list contains gene-targets with existing compounds in development. The disKO list contains gene-targets without existing compounds yet and represent novel targets for drug discovery. Abstract: Refractory disease is a major challenge in treating patients with acute myeloid leukemia (AML). Whereas the armamentarium has expanded in the past few years for treating AML, long-term survival outcomes have yet to be proven. To further expand the arsenal for treating AML, we searched for druggable gene targets in AML by analyzing screening data from a lentiviral-based genome-wide pooled CRISPR-Cas9 library and gene knockout (KO) dependency scores in 15 AML cell lines (HEL, MV411, OCIAML2, THP1, NOMO1, EOL1, KASUMI1, NB4, OCIAML3, MOLM13, TF1, U937, F36P, AML193, P31FUJ). -

POLR2L Is Over-Expressed in Human Endometrial Cancer
Over-expression of RNA polymerase II subunit L in human endometrial cancer. Shahan Mamoor, MS1 [email protected] East Islip, NY USA Gynecologic cancers including cancers of the endometrium are a clinical problem1-4. We mined published microarray data5,6 to discover genes associated with endometrial cancers by comparing transcriptomes of the normal endometrium and endometrial tumors from humans. We identified RNA polymerase II subunit L, encoded by POLR2L, as among the most differentially expressed genes, transcriptome-wide, in cancers of the endometrium. POLR2L was expressed at significantly higher levels in endometrial tumor tissues as compared to the endometrium. Importantly, primary tumor expression of POLR2L was correlated with recurrence-free survival in patients with endometrial cancer. POLR2L may be a molecule of interest in understanding the etiology or progression of human endometrial cancer. Keywords: endometrial cancer, gynecologic cancers, endometrium, POLR2L, RNA polymerase II subunit L, systems biology of endometrial cancer, targeted therapeutics in endometrial cancer. 1 Endometrial cancer is the most common gynecologic cancer in the developed world1. Over the last three decades, the incidence of endometrial cancer has increased 21%4 and the death rate has increased 100%3. We harnessed the power of independently published microarray datasets5,6 to determine in an unbiased fashion and at the systems-level genes most differentially expressed in endometrial tumors. We report here the differential and increased expression of the RNA polymerase II subunit L (POLR2L) in human endometrial cancer. Methods We utilized datasets GSE636785 and GSE1158106 for this global differential gene expression analysis of human endometrial cancer in conjunction with GEO2R. -

Gene Duplication and Neofunctionalization: POLR3G and POLR3GL
Downloaded from genome.cshlp.org on September 27, 2021 - Published by Cold Spring Harbor Laboratory Press Research Gene duplication and neofunctionalization: POLR3G and POLR3GL Marianne Renaud,1 Viviane Praz,1,2 Erwann Vieu,1,5 Laurence Florens,3 Michael P. Washburn,3,4 Philippe l’Hoˆte,1 and Nouria Hernandez1,6 1Center for Integrative Genomics, Faculty of Biology and Medicine, University of Lausanne, 1015 Lausanne, Switzerland; 2Swiss Institute of Bioinformatics, 1015 Lausanne, Switzerland; 3Stowers Institute for Medical Research, Kansas City, Missouri 64110, USA; 4Department of Pathology and Laboratory Medicine, The University of Kansas Medical Center, Kansas City, Kansas 66160, USA RNA polymerase III (Pol III) occurs in two versions, one containing the POLR3G subunit and the other the closely related POLR3GL subunit. It is not clear whether these two Pol III forms have the same function, in particular whether they recognize the same target genes. We show that the POLR3G and POLR3GL genes arose from a DNA-based gene duplication, probably in a common ancestor of vertebrates. POLR3G- as well as POLR3GL-containing Pol III are present in cultured cell lines and in normal mouse liver, although the relative amounts of the two forms vary, with the POLR3G-containing Pol III relatively more abundant in dividing cells. Genome-wide chromatin immunoprecipitations followed by high-throughput sequencing (ChIP-seq) reveal that both forms of Pol III occupy the same target genes, in very constant proportions within one cell line, suggesting that the two forms of Pol III have a similar function with regard to specificity for target genes. In contrast, the POLR3G promoter—not the POLR3GL promoter—binds the transcription factor MYC, as do all other pro- moters of genes encoding Pol III subunits. -

Renoprotective Effect of Combined Inhibition of Angiotensin-Converting Enzyme and Histone Deacetylase
BASIC RESEARCH www.jasn.org Renoprotective Effect of Combined Inhibition of Angiotensin-Converting Enzyme and Histone Deacetylase † ‡ Yifei Zhong,* Edward Y. Chen, § Ruijie Liu,*¶ Peter Y. Chuang,* Sandeep K. Mallipattu,* ‡ ‡ † | ‡ Christopher M. Tan, § Neil R. Clark, § Yueyi Deng, Paul E. Klotman, Avi Ma’ayan, § and ‡ John Cijiang He* ¶ *Department of Medicine, Mount Sinai School of Medicine, New York, New York; †Department of Nephrology, Longhua Hospital, Shanghai University of Traditional Chinese Medicine, Shanghai, China; ‡Department of Pharmacology and Systems Therapeutics and §Systems Biology Center New York, Mount Sinai School of Medicine, New York, New York; |Baylor College of Medicine, Houston, Texas; and ¶Renal Section, James J. Peters Veterans Affairs Medical Center, New York, New York ABSTRACT The Connectivity Map database contains microarray signatures of gene expression derived from approximately 6000 experiments that examined the effects of approximately 1300 single drugs on several human cancer cell lines. We used these data to prioritize pairs of drugs expected to reverse the changes in gene expression observed in the kidneys of a mouse model of HIV-associated nephropathy (Tg26 mice). We predicted that the combination of an angiotensin-converting enzyme (ACE) inhibitor and a histone deacetylase inhibitor would maximally reverse the disease-associated expression of genes in the kidneys of these mice. Testing the combination of these inhibitors in Tg26 mice revealed an additive renoprotective effect, as suggested by reduction of proteinuria, improvement of renal function, and attenuation of kidney injury. Furthermore, we observed the predicted treatment-associated changes in the expression of selected genes and pathway components. In summary, these data suggest that the combination of an ACE inhibitor and a histone deacetylase inhibitor could have therapeutic potential for various kidney diseases. -

Anti-POLR2L (GW22520F)
3050 Spruce Street, Saint Louis, MO 63103 USA Tel: (800) 521-8956 (314) 771-5765 Fax: (800) 325-5052 (314) 771-5757 email: [email protected] Product Information Anti-POLR2L antibody produced in chicken, affinity isolated antibody Catalog Number GW22520F Formerly listed as GenWay Catalog Number 15-288-22520F, DNA-directed RNA polymerase II 7.6 kDa polypeptide Antibody. The product is a clear, colorless solution in phosphate – Storage Temperature Store at 20 °C buffered saline, pH 7.2, containing 0.02% sodium azide. Synonyms: DNA directed RNA polymerase II polypeptide L, Species Reactivity: Human, mouse EC 2.7.7.6; RPB10; RPB7.6; RPABC5 Tested Applications: WB Product Description Recommended Dilutions: Recommended starting dilution DNA-dependent RNA polymerase catalyzes the transcrip- for Western blot analysis is 1:500, for tissue or cell staining tion of DNA into RNA using the four ribonucleoside 1:200. triphosphates as substrates. Note: Optimal concentrations and conditions for each NCBI Accession number: NP_066951.1 application should be determined by the user. Swiss Prot Accession number: P62875 Precautions and Disclaimer Gene Information: Human .. POLR2L (5441) This product is for R&D use only, not for drug, household, or Immunogen: Recombinant protein DNA directed RNA other uses. Due to the sodium azide content a material polymerase II polypeptide L safety data sheet (MSDS) for this product has been sent to the attention of the safety officer of your institution. Please Immunogen Sequence: GI # 10863925, sequence 1 - 67 consult the Material Safety Data Sheet for information regarding hazards and safe handling practices. Storage/Stability For continuous use, store at 2–8 °C for up to one week.