Induced Polyploidy: a Tool for Forage Species Improvement
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Ray Grass Perenne Pdf
Ray grass perenne pdf Continue Published in Temperate Climate and Cold Description: Unlike the annual rye grass, this variety is characterized by lasting over two years, being able to reach up to three or four years. It has a shallow fibrous radical system. It adapts to a temperate climate, does not tolerate high temperatures. Its growth is straight, with bright green leaves. It supports trampling, frost and is well competitive with other species. Soil: Suitable for all soils (except sandy soils), although it prefers fertile and moist soils and ph is close to neutrality. Nutrients and water: Requires irrigation, especially in summer and high fertilization. Shadow: Not suitable. Landing density: 3 to 15 kg / 100 m2. It is preferable to sow in early autumn, but can also be done in summer. Please note that the depth is no more than 2 cm. Cutting: 3 to 5 cm. Frequency: increased. Use: Widely used in parks and sports grounds, in mixes. It belongs to the family of grass, which is widely used all over the world and is considered one of the most valuable species of meadows. It is one of the most commonly used species both alone and in the mix. It has excellent tolerance to use, rapid germination (7 days in spring and 10 days in winter) and excellent installation speed. On the other hand, it is a species that is not drought-tolerant and requires high maintenance. Crossing different varieties has produced a wide range of varieties that differ in their characteristics, such as: tolerance to use, winter strength, color, resistance to disease and tolerance to heat and drought. -

Initial Study Appendix B
Uvas Road at Little Uvas Creek Bridge Replacement Project Biological Assessment Biological Assessment Uvas Road over Little Uvas Creek Bridge Replacement Project (37C-0095/37C-0601 [new]) Near Morgan Hill, Santa Clara County, California 04-SCL-0-CR Federal Project Number BRLO 5937(124) Caltrans District 04 November 2015 Biological Assessment Uvas Road over Little Uvas Creek Bridge Replacement Project (37C-0095/37C-0601 [new]) Near Morgan Hill, Santa Clara County, California 04-SCL-0-CR Federal Project Number BRLO 5937(124) Caltrans District 04 November 2015 STATE OF CALIFORNIA Department of Transportation and Santa Clara County Roads and Airports Department Prepared By: ___________________________________ Date: ____________ Patrick Boursier, Principal (408) 458-3204 H. T. Harvey & Associates Los Gatos, California Approved By: ___________________________________ Date: ____________ Solomon Tegegne, Associate Civil Engineer Santa Clara County Roads and Airports Department Highway and Bridge Design 408-573-2495 Concurred By: ___________________________________ Date: ____________ Tom Holstein Environmental Branch Chief Office of Local Assistance Caltrans, District 4 Oakland, California 510-286-5250 For individuals with sensory disabilities, this document is available in Braille, large print, on audiocassette, or computer disk. To obtain a copy in one of these alternate formats, please call or write to the Santa Clara County Roads and Airports Department: Solomon Tegegne Santa Clara County Roads and Airports Department 101 Skyport Drive San Jose, CA 95110 408-573-2495 Summary of Findings, Conclusions and Determinations Summary of Findings, Conclusions and Determinations The Uvas Road at Little Uvas Creek Bridge Replacement Project (proposed project) is proposed by the County of Santa Clara Roads and Airports Department in cooperation with the Office of Local Assistance of the California Department of Transportation (Caltrans), and this Biological Assessment (BA) has been prepared following Caltrans’ procedures. -
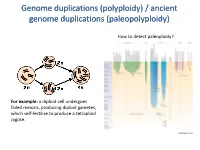
Polyploidy) / Ancient Genome Duplications (Paleopolyploidy
Genome duplications (polyploidy) / ancient genome duplications (paleopolyploidy) How to detect paleoploidy? For example: a diploid cell undergoes failed meiosis, producing diploid gametes, which self-fertilize to produce a tetraploid zygote. Timing of duplication by trees (phylogenetic timing) Phylogenetic timing of duplicates b Paramecium genome duplications Comparison of two scaffolds originating from a common ancestor at the recent WGD Saccharomyces cerevisiae Just before genome duplication Just after genome duplication More time after genome duplication Unaligned view (removing gaps just like in cerev has occurred) Saccharomyces cerevisiae Problem reciprocal gene loss (extreme case); how to solve? Problem reciprocal gene loss (extreme case); how to solve? Just before genome duplication Outgroup! Just after genome duplication Outgroup Just after genome duplication Outgroup More time after genome duplication Outgroup Problem (extreme case); how to solve? Outgroup Outgroup Outgroup Outgroup Outgroup Using other genomes Wong et al. 2002 PNAS Centromeres Vertebrate genome duplication Nature. 2011 Apr 10. [Epub ahead of print] Ancestral polyploidy in seed plants and angiosperms. Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, Tomsho LP, Hu Y, Liang H, Soltis PS, Soltis DE, Clifton SW, Schlarbaum SE, Schuster SC, Ma H, Leebens-Mack J, Depamphilis CW. Flowering plants Flowering MOSS Vertebrates Teleosts S. serevisiae and close relatives Paramecium Reconstructed map of genome duplications allows unprecedented mapping -

The Diversity of Plant Sex Chromosomes Highlighted Through Advances in Genome Sequencing
G C A T T A C G G C A T genes Review The Diversity of Plant Sex Chromosomes Highlighted through Advances in Genome Sequencing Sarah Carey 1,2 , Qingyi Yu 3,* and Alex Harkess 1,2,* 1 Department of Crop, Soil, and Environmental Sciences, Auburn University, Auburn, AL 36849, USA; [email protected] 2 HudsonAlpha Institute for Biotechnology, Huntsville, AL 35806, USA 3 Texas A&M AgriLife Research, Texas A&M University System, Dallas, TX 75252, USA * Correspondence: [email protected] (Q.Y.); [email protected] (A.H.) Abstract: For centuries, scientists have been intrigued by the origin of dioecy in plants, characterizing sex-specific development, uncovering cytological differences between the sexes, and developing theoretical models. Through the invention and continued improvements in genomic technologies, we have truly begun to unlock the genetic basis of dioecy in many species. Here we broadly review the advances in research on dioecy and sex chromosomes. We start by first discussing the early works that built the foundation for current studies and the advances in genome sequencing that have facilitated more-recent findings. We next discuss the analyses of sex chromosomes and sex-determination genes uncovered by genome sequencing. We synthesize these results to find some patterns are emerging, such as the role of duplications, the involvement of hormones in sex-determination, and support for the two-locus model for the origin of dioecy. Though across systems, there are also many novel insights into how sex chromosomes evolve, including different sex-determining genes and routes to suppressed recombination. We propose the future of research in plant sex chromosomes should involve interdisciplinary approaches, combining cutting-edge technologies with the classics Citation: Carey, S.; Yu, Q.; to unravel the patterns that can be found across the hundreds of independent origins. -

Mitosis Vs. Meiosis
Mitosis vs. Meiosis In order for organisms to continue growing and/or replace cells that are dead or beyond repair, cells must replicate, or make identical copies of themselves. In order to do this and maintain the proper number of chromosomes, the cells of eukaryotes must undergo mitosis to divide up their DNA. The dividing of the DNA ensures that both the “old” cell (parent cell) and the “new” cells (daughter cells) have the same genetic makeup and both will be diploid, or containing the same number of chromosomes as the parent cell. For reproduction of an organism to occur, the original parent cell will undergo Meiosis to create 4 new daughter cells with a slightly different genetic makeup in order to ensure genetic diversity when fertilization occurs. The four daughter cells will be haploid, or containing half the number of chromosomes as the parent cell. The difference between the two processes is that mitosis occurs in non-reproductive cells, or somatic cells, and meiosis occurs in the cells that participate in sexual reproduction, or germ cells. The Somatic Cell Cycle (Mitosis) The somatic cell cycle consists of 3 phases: interphase, m phase, and cytokinesis. 1. Interphase: Interphase is considered the non-dividing phase of the cell cycle. It is not a part of the actual process of mitosis, but it readies the cell for mitosis. It is made up of 3 sub-phases: • G1 Phase: In G1, the cell is growing. In most organisms, the majority of the cell’s life span is spent in G1. • S Phase: In each human somatic cell, there are 23 pairs of chromosomes; one chromosome comes from the mother and one comes from the father. -

Review Cell Fusion and Some Subcellular Properties Of
REVIEW CELL FUSION AND SOME SUBCELLULAR PROPERTIES OF HETEROKARYONS AND HYBRIDS SAIMON GORDON From the Genetics Laboratory, Department of Biochemistry, The University of Oxford, England, and The Rockefeller University, New York 10021 I. INTRODUCTION The technique of somatic cell fusion has made it first cell hybrids obtained by this method. The iso- possible to study cell biology in an unusual and lation of hybrid cells from such mixed cultures direct way. When cells are mixed in the presence of was greatly facilitated by the Szybalski et al. (6) Sendai virus, their membranes coalesce, the cyto- and Littlefield (7, 8) adaptation of a selective me- plasm becomes intermingled, and multinucleated dium containing hypoxanthine, aminopterin, and homo- and heterokaryons are formed by fusion of thymidine (HAT) to mammalian systems. similar or different cells, respectively (1, 2). By Further progress followed the use of Sendai fusing cells which contrast in some important virus by Harris and Watkins to increase the biologic property, it becomes possible to ask ques- frequency of heterokaryon formation (9). These tions about dominance of control processes, nu- authors exploited the observation by Okada that cleocytoplasmic interactions, and complementa- UV irradiation could be used to inactivate Sendai tion in somatic cell heterokaryons. The multinucle- virus without loss of fusion efficiency (10). It was ate cell may divide and give rise to mononuclear therefore possible to eliminate the problem of virus cells containing chromosomes from both parental replication in fused cells. Since many cells carry cells and become established as a hybrid cell line receptors for Sendai virus, including those of able to propagate indefinitely in vitro. -

Gene Therapy Product Quality Aspects in the Production of Vectors and Genetically Modified Somatic Cells
_____________________________________________________________ 3AB6a ■ GENE THERAPY PRODUCT QUALITY ASPECTS IN THE PRODUCTION OF VECTORS AND GENETICALLY MODIFIED SOMATIC CELLS Guideline Title Gene Therapy Product Quality Aspects in the Production of Vectors and Genetically Modified Somatic Cells Legislative basis Directive 75/318/EEC as amended Date of first adoption December 1994 Date of entry into July 1995 force Status Last revised December 1994 Previous titles/other Gene Therapy Products - Quality, Safety and Efficacy references Aspects in the Production of Vectors and Genetically Modified Somatic Cells/ III/5863/93 Additional Notes This note for guidance is intended to facilitate the collection and submission of data to support applications for marketing authorisation within the EC for gene therapy products derived by biotechnology/high technology and intended for medicinal use in man. CONTENTS 1. INTRODUCTION 2. POINTS TO CONSIDER IN MANUFACTURE 3. DEVELOPMENT GENETICS 4. PRODUCTION 5. PURIFICATION 6. PRODUCT CHARACTERISATION 7. CONSISTENCY AND ROUTINE BATCH CONTROL OF FINAL PROCESSED PRODUCT 8. SAFETY REGULATIONS 275 _____________________________________________________________ 3AB6a ■ GENE THERAPY PRODUCT QUALITY ASPECTS IN THE PRODUCTION OF VECTORS AND GENETICALLY MODIFIED SOMATIC CELLS 1. INTRODUCTION Somatic gene therapy encompasses medical interventions which involve the deliberate modification of the genetic material of somatic cells. Scientific progress over the past decade has led to the development of novel methods for the transfer of new genetic material into patients’ cells. The aims of these methods include the efficient transfer and functional expression or manifestation of the transferred genetic material in a target somatic cell population for therapeutic, prophylactic or diagnostic purposes. Although in the majority of cases the intention is the addition and expression of a gene to yield a protein product, the transfer of nucleic acids with the aim of modifying the function or expression of an endogenous gene, e.g. -

Human Germline Genome Editing: Fact Sheet
Human germline genome editing: fact sheet Purpose • To contribute to evidence-informed discussions about human germline genome editing. KEY TAKEAWAYS • Gene editing offers the potential to improve human health in ways not previously possible. • Making changes to human genes that can be passed on to future generations is prohibited in Australia. • Unresolved questions remain on the possible long-term impacts, unintended consequences, and ethical issues associated with introducing heritable changes by editing of the genome of human gametes (sperm and eggs) and embryos. • AusBiotech believes the focus of human gene editing should remain on non-inheritable changes until such time as the scientific evidence, regulatory frameworks and health care models have progressed sufficiently to warrant consideration of any heritable genetic edits. Gene editing Gene editing is the insertion, deletion, or modification of DNA to modify an organism’s specific genetic characteristics. New and evolving gene editing techniques and tools (e.g. CRISPR) allow editing of genes with a level of precision that increases its applications across the health, agricultural, and industrial sectors. These breakthrough techniques potentially offer a range of different options for treating devastating human diseases and delivering environmentally sustainable food production systems that can feed the world’s growing population, which is expected to exceed nine billion by 2050. The current primary application of human gene editing is on non-reproductive cells (‘somatic’ cells) -

Checklist of the Vascular Plants of Redwood National Park
Humboldt State University Digital Commons @ Humboldt State University Botanical Studies Open Educational Resources and Data 9-17-2018 Checklist of the Vascular Plants of Redwood National Park James P. Smith Jr Humboldt State University, [email protected] Follow this and additional works at: https://digitalcommons.humboldt.edu/botany_jps Part of the Botany Commons Recommended Citation Smith, James P. Jr, "Checklist of the Vascular Plants of Redwood National Park" (2018). Botanical Studies. 85. https://digitalcommons.humboldt.edu/botany_jps/85 This Flora of Northwest California-Checklists of Local Sites is brought to you for free and open access by the Open Educational Resources and Data at Digital Commons @ Humboldt State University. It has been accepted for inclusion in Botanical Studies by an authorized administrator of Digital Commons @ Humboldt State University. For more information, please contact [email protected]. A CHECKLIST OF THE VASCULAR PLANTS OF THE REDWOOD NATIONAL & STATE PARKS James P. Smith, Jr. Professor Emeritus of Botany Department of Biological Sciences Humboldt State Univerity Arcata, California 14 September 2018 The Redwood National and State Parks are located in Del Norte and Humboldt counties in coastal northwestern California. The national park was F E R N S established in 1968. In 1994, a cooperative agreement with the California Department of Parks and Recreation added Del Norte Coast, Prairie Creek, Athyriaceae – Lady Fern Family and Jedediah Smith Redwoods state parks to form a single administrative Athyrium filix-femina var. cyclosporum • northwestern lady fern unit. Together they comprise about 133,000 acres (540 km2), including 37 miles of coast line. Almost half of the remaining old growth redwood forests Blechnaceae – Deer Fern Family are protected in these four parks. -

Cell Division- Ch 5
Cell Division- Mitosis and Meiosis When do cells divide? Cell size . One of most important factors affecting size of the cell is size of cell membrane . Cell must remain relatively small to survive (why?) – Cell membrane has to be big enough to take in nutrients and eliminate wastes – As cells get bigger, the volume increases faster than the surface area – Small cells have a larger surface area to volume ratio than larger cells to help with nutrient intake and waste elimination . When a cell reaches its max size, the nucleus starts cell division: called MITOSIS or MEIOSIS Mitosis . General Information – Occurs in somatic (body) cells ONLY!! – Nickname: called “normal” cell division – Produces somatic cells only . Background Info – Starts with somatic cell in DIPLOID (2n) state . Cell contains homologous chromosomes- chromosomes that control the same traits but not necessarily in the same way . 1 set from mom and 1 set from dad – Ends in diploid (2n) state as SOMATIC cells – Goes through one set of divisions – Start with 1 cell and end with 2 cells Mitosis (cont.) . Accounts for three essential life processes – Growth . Result of cell producing new cells . Develop specialized shapes/functions in a process called differentiation . Rate of cell division controlled by GH (Growth Hormone) which is produced in the pituitary gland . Ex. Nerve cell, intestinal cell, etc. – Repair . Cell regenerates at the site of injury . Ex. Skin (replaced every 28 days), blood vessels, bone Mitosis (cont.) – Reproduction . Asexual – Offspring produced by only one parent – Produce offspring that are genetically identical – MITOSIS – Ex. Bacteria, fungi, certain plants and animals . -

© 2020 Hui Chen ALL RIGHTS RESERVED
© 2020 Hui Chen ALL RIGHTS RESERVED TRAFFIC TOLERANCE OF FINE FESCUES: TECHNIQUES FOR SCREENING GERMPLASM by HUI CHEN A dissertation submitted to the School of Graduate Studies Rutgers, The State University of New Jersey in partial fulfillment of the requirements for the degree of Doctor of Philosophy Graduate Program in Plant Biology written under the direction of James A. Murphy and approved by ________________________ ________________________ ________________________ ________________________ ________________________ New Brunswick, New Jersey JANUARY 2020 ABSTRACT OF THE DISSERTATION Traffic tolerance of fine fescues: Techniques for screening germplasm BY HUI CHEN Dissertation Director: Dr, James A. Murphy The term fine fescue refers to several Festuca spp. that have a very fine leaf texture compared to most other turfgrass species. These species are adapted to low-input management systems and have been used in mixtures with other cool-season grasses. However, fine fescues are not utilized to the same extent as other species partially due to their poor traffic tolerance and recuperative ability. Improvement in traffic tolerance of fine fescues would enable use of these grasses beyond turf systems that experience little to no traffic. The purpose of this dissertation was to develop and evaluate germplasm screening techniques that improve selecting efficiency for traffic tolerant fine fescues. The specific objectives of this research were: i) to evaluate the effect of traffic form (abrasive wear vs. cleated traffic) and season (spring vs. summer vs. autumn) on the assessment of fine fescue traffic tolerance (Chapters 1 and 2); ii) to evaluate the effect of nitrogen fertilization and harvest time on cell wall composition of fine fescues (Chapters 3 and 4); iii) to investigate the correlation between cell wall composition and wear tolerance of fine ii fescues; and iv) to develop near-infrared reflectance spectroscopy (NIRS) models to determine the cell wall composition of fine fescues. -

Bio 103 Lecture
Biology 103 Lecture Mitosis and Meiosis Study Guide (Cellular Basis of Reproduction - Chapter 8) Like begets like, more or less · does sexual reproduction and genetics allow for a maple tree to produce a sea star? · do offspring inherit genetic material from both parents in asexual reproduction? · do offspring inherit genetic material from both parents in sexual reproduction? · what is the name of structures that contain most of an organism's DNA? Cells arise only from preexisting cells · what two main roles does cell division play in perpetuating the life cycle of animals and other multicellular organisms? Prokaryotes reproduce by binary fission · what is the name of the type of cell division used by prokaryotes to reproduce themselves? · do prokaryotes have DNA? · do prokaryotes replicate their chromosome prior to division? · is binary fission considered asexual or sexual reproduction and why? The large, complex chromosomes of eukaryotes duplicate with each cell division · in what structure of the eukaryotic cell do most of the genes occur? · what are genes and where are they located? · what is chromatin and how is it different from chromosomes? · understand chromosome duplication and distribution The cell cycle multiplies cells · define cell cycle · what are the major components of the cell cycle? · in what phase of the cell cycle does the cell spend the most time? · what occurs during cytokinesis? Cell division is a continuum of dynamic changes · know that the main things that occur during the four stages of mitosis are formation