Pan-Cancer Screen for Mutations in Non-Coding Elements with Conservation and Cancer Specificity Reveals Correlations with Expression and Survival
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
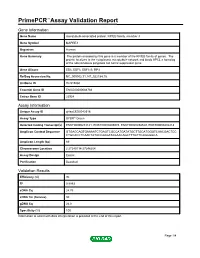
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name microtubule-associated protein, RP/EB family, member 3 Gene Symbol MAPRE3 Organism Human Gene Summary The protein encoded by this gene is a member of the RP/EB family of genes. The protein localizes to the cytoplasmic microtubule network and binds APCL a homolog of the adenomatous polyposis coli tumor suppressor gene. Gene Aliases EB3, EBF3, EBF3-S, RP3 RefSeq Accession No. NC_000002.11, NT_022184.15 UniGene ID Hs.515860 Ensembl Gene ID ENSG00000084764 Entrez Gene ID 22924 Assay Information Unique Assay ID qHsaCED0042518 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000233121, ENST00000405074, ENST00000458529, ENST00000402218 Amplicon Context Sequence GTGACCAGTGAAAATCTGAGTCGCCATGATATGCTTGCATGGGTCAACGACTCC CTGCACCTCAACTATACCAAGATAGAACAGCTTTGTTCAGGGGCA Amplicon Length (bp) 69 Chromosome Location 2:27245114-27246204 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 90 R2 0.9993 cDNA Cq 24.76 cDNA Tm (Celsius) 80 gDNA Cq 26.8 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report MAPRE3, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

EB3 (MAPRE3) (NM 012326) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC204135 EB3 (MAPRE3) (NM_012326) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: EB3 (MAPRE3) (NM_012326) Human Tagged ORF Clone Tag: Myc-DDK Symbol: MAPRE3 Synonyms: EB3; EBF3; EBF3-S; RP3 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RC204135 ORF sequence Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGCCGTCAATGTGTACTCCACATCTGTGACCAGTGAAAATCTGAGTCGCCATGATATGCTTGCATGGG TCAACGACTCCCTGCACCTCAACTATACCAAGATAGAACAGCTTTGTTCAGGGGCAGCCTACTGCCAGTT CATGGACATGCTCTTCCCCGGCTGTGTGCACTTGAGGAAAGTGAAGTTCCAGGCCAAACTAGAGCATGAA TACATCCACAACTTCAAGGTGCTGCAAGCAGCTTTCAAGAAGATGGGTGTTGACAAAATCATTCCTGTAG AGAAATTAGTGAAAGGAAAATTCCAAGATAATTTTGAGTTTATTCAGTGGTTTAAGAAATTCTTTGACGC AAACTATGATGGAAAGGATTACAACCCTCTGCTGGCGCGGCAGGGCCAGGACGTAGCGCCACCTCCTAAC CCAGGTGATCAGATCTTCAACAAATCCAAGAAACTCATTGGCACAGCAGTTCCACAGAGGACGTCCCCCA CAGGCCCAAAAAACATGCAGACCTCTGGCCGGCTGAGCAATGTGGCCCCCCCCTGCATTCTCCGGAAGAA TCCTCCATCAGCCCGAAATGGCGGCCATGAGACTGATGCCCAAATTCTTGAACTCAACCAACAGCTGGTG GACTTGAAGCTGACAGTGGATGGGCTGGAGAAGGAACGTGACTTCTACTTCAGCAAACTTCGTGACATCG AGCTCATCTGCCAGGAGCATGAAAGTGAAAACAGCCCTGTTATCTCAGGCATCATTGGCATCCTCTATGC CACAGAGGAAGGATTCGCACCCCCTGAGGACGATGAGATTGAAGAGCATCAACAAGAAGACCAGGACGAG TAC ACGCGTACGCGGCCGCTCGAGCAGAAACTCATCTCAGAAGAGGATCTGGCAGCAAATGATATCCTGGATT -

Supporting Information
Supporting Information Pouryahya et al. SI Text Table S1 presents genes with the highest absolute value of Ricci curvature. We expect these genes to have significant contribution to the network’s robustness. Notably, the top two genes are TP53 (tumor protein 53) and YWHAG gene. TP53, also known as p53, it is a well known tumor suppressor gene known as the "guardian of the genome“ given the essential role it plays in genetic stability and prevention of cancer formation (1, 2). Mutations in this gene play a role in all stages of malignant transformation including tumor initiation, promotion, aggressiveness, and metastasis (3). Mutations of this gene are present in more than 50% of human cancers, making it the most common genetic event in human cancer (4, 5). Namely, p53 mutations play roles in leukemia, breast cancer, CNS cancers, and lung cancers, among many others (6–9). The YWHAG gene encodes the 14-3-3 protein gamma, a member of the 14-3-3 family proteins which are involved in many biological processes including signal transduction regulation, cell cycle pro- gression, apoptosis, cell adhesion and migration (10, 11). Notably, increased expression of 14-3-3 family proteins, including protein gamma, have been observed in a number of human cancers including lung and colorectal cancers, among others, suggesting a potential role as tumor oncogenes (12, 13). Furthermore, there is evidence that loss Fig. S1. The histogram of scalar Ricci curvature of 8240 genes. Most of the genes have negative scalar Ricci curvature (75%). TP53 and YWHAG have notably low of p53 function may result in upregulation of 14-3-3γ in lung cancer Ricci curvatures. -

In This Table Protein Name, Uniprot Code, Gene Name P-Value
Supplementary Table S1: In this table protein name, uniprot code, gene name p-value and Fold change (FC) for each comparison are shown, for 299 of the 301 significantly regulated proteins found in both comparisons (p-value<0.01, fold change (FC) >+/-0.37) ALS versus control and FTLD-U versus control. Two uncharacterized proteins have been excluded from this list Protein name Uniprot Gene name p value FC FTLD-U p value FC ALS FTLD-U ALS Cytochrome b-c1 complex P14927 UQCRB 1.534E-03 -1.591E+00 6.005E-04 -1.639E+00 subunit 7 NADH dehydrogenase O95182 NDUFA7 4.127E-04 -9.471E-01 3.467E-05 -1.643E+00 [ubiquinone] 1 alpha subcomplex subunit 7 NADH dehydrogenase O43678 NDUFA2 3.230E-04 -9.145E-01 2.113E-04 -1.450E+00 [ubiquinone] 1 alpha subcomplex subunit 2 NADH dehydrogenase O43920 NDUFS5 1.769E-04 -8.829E-01 3.235E-05 -1.007E+00 [ubiquinone] iron-sulfur protein 5 ARF GTPase-activating A0A0C4DGN6 GIT1 1.306E-03 -8.810E-01 1.115E-03 -7.228E-01 protein GIT1 Methylglutaconyl-CoA Q13825 AUH 6.097E-04 -7.666E-01 5.619E-06 -1.178E+00 hydratase, mitochondrial ADP/ATP translocase 1 P12235 SLC25A4 6.068E-03 -6.095E-01 3.595E-04 -1.011E+00 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 Protein kinase C and casein Q9BY11 PACSIN1 3.837E-03 -5.863E-01 3.680E-06 -1.824E+00 kinase substrate in neurons protein 1 Tubulin polymerization- O94811 TPPP 6.466E-03 -5.755E-01 6.943E-06 -1.169E+00 promoting protein MIC C9JRZ6 CHCHD3 2.912E-02 -6.187E-01 2.195E-03 -9.781E-01 Mitochondrial 2- -

Transcriptomic and Epigenomic Characterization of the Developing Bat Wing
ARTICLES OPEN Transcriptomic and epigenomic characterization of the developing bat wing Walter L Eckalbar1,2,9, Stephen A Schlebusch3,9, Mandy K Mason3, Zoe Gill3, Ash V Parker3, Betty M Booker1,2, Sierra Nishizaki1,2, Christiane Muswamba-Nday3, Elizabeth Terhune4,5, Kimberly A Nevonen4, Nadja Makki1,2, Tara Friedrich2,6, Julia E VanderMeer1,2, Katherine S Pollard2,6,7, Lucia Carbone4,8, Jeff D Wall2,7, Nicola Illing3 & Nadav Ahituv1,2 Bats are the only mammals capable of powered flight, but little is known about the genetic determinants that shape their wings. Here we generated a genome for Miniopterus natalensis and performed RNA-seq and ChIP-seq (H3K27ac and H3K27me3) analyses on its developing forelimb and hindlimb autopods at sequential embryonic stages to decipher the molecular events that underlie bat wing development. Over 7,000 genes and several long noncoding RNAs, including Tbx5-as1 and Hottip, were differentially expressed between forelimb and hindlimb, and across different stages. ChIP-seq analysis identified thousands of regions that are differentially modified in forelimb and hindlimb. Comparative genomics found 2,796 bat-accelerated regions within H3K27ac peaks, several of which cluster near limb-associated genes. Pathway analyses highlighted multiple ribosomal proteins and known limb patterning signaling pathways as differentially regulated and implicated increased forelimb mesenchymal condensation in differential growth. In combination, our work outlines multiple genetic components that likely contribute to bat wing formation, providing insights into this morphological innovation. The order Chiroptera, commonly known as bats, is the only group of To characterize the genetic differences that underlie divergence in mammals to have evolved the capability of flight. -

The Genetic Basis of Emotional Behaviour in Mice
European Journal of Human Genetics (2006) 14, 721–728 & 2006 Nature Publishing Group All rights reserved 1018-4813/06 $30.00 www.nature.com/ejhg REVIEW The genetic basis of emotional behaviour in mice Saffron AG Willis-Owen*,1 and Jonathan Flint1 1Wellcome Trust Centre for Human Genetics, Roosevelt Drive, Headington, Oxford, UK The last decade has witnessed a steady expansion in the number of quantitative trait loci (QTL) mapped for complex phenotypes. However, despite this proliferation, the number of successfully cloned QTL has remained surprisingly low, and to a great extent limited to large effect loci. In this review, we follow the progress of one complex trait locus; a low magnitude moderator of murine emotionality identified some 10 years ago in a simple two-strain intercross, and successively resolved using a variety of crosses and fear-related phenotypes. These experiments have revealed a complex underlying genetic architecture, whereby genetic effects fractionate into several separable QTL with some evidence of phenotype specificity. Ultimately, we describe a method of assessing gene candidacy, and show that given sufficient access to genetic diversity and recombination, progression from QTL to gene can be achieved even for low magnitude genetic effects. European Journal of Human Genetics (2006) 14, 721–728. doi:10.1038/sj.ejhg.5201569 Keywords: mouse; emotionality; quantitative trait locus; QTL Introduction tude genetic effects and their interactions (both with other Emotionality is a psychological trait of complex aetiology, genetic loci (ie epistasis) and nongenetic (environmental) which moderates an organism’s response to stress. Beha- factors) ultimately producing a quasi-continuously distrib- vioural evidence of emotionality has been documented uted phenotype. -

Different Contribution of BRINP3 Gene in Chronic
Casado et al. BMC Oral Health (2015) 15:33 DOI 10.1186/s12903-015-0018-6 RESEARCH ARTICLE Open Access Different contribution of BRINP3 gene in chronic periodontitis and peri-implantitis: a cross- sectional study Priscila L Casado1,2, Diego P Aguiar2, Lucas C Costa3, Marcos A Fonseca3, Thays CS Vieira2,4, Claudia CK Alvim-Pereira5, Fabiano Alvim-Pereira6, Kathleen Deeley7, José M Granjeiro1,9, Paula C Trevilatto8 and Alexandre R Vieira7,10* Abstract Background: Peri-implantitis is a chronic inflammation, resulting in loss of supporting bone around implants. Chronic periodontitis is a risk indicator for implant failure. Both diseases have a common etiology regarding inflammatory destructive response. BRINP3 gene is associated with aggressive periodontitis. However, is still unclear if chronic periodontitis and peri-implantitis have the same genetic background. The aim of this work was to investigate the association between BRINP3 genetic variation (rs1342913 and rs1935881) and expression and susceptibility to both diseases. Methods: Periodontal and peri-implant examinations were performed in 215 subjects, divided into: healthy (without chronic periodontitis and peri-implantitis, n = 93); diseased (with chronic periodontitis and peri-implantitis, n = 52); chronic periodontitis only (n = 36), and peri-implantitis only (n = 34). A replication sample of 92 subjects who lost implants and 185 subjects successfully treated with implants were tested. DNA was extracted from buccal cells. Two genetic markers of BRINP3 (rs1342913 and rs1935881) were genotyped using TaqMan chemistry. Chi-square (p < 0.05) compared genotype and allele frequency between groups. A subset of subjects (n = 31) had gingival biopsies ΔΔCT harvested. The BRINP3 mRNA levels were studied by CT method (2 ). -

Castration Delays Epigenetic Aging and Feminizes DNA
RESEARCH ARTICLE Castration delays epigenetic aging and feminizes DNA methylation at androgen- regulated loci Victoria J Sugrue1, Joseph Alan Zoller2, Pritika Narayan3, Ake T Lu4, Oscar J Ortega-Recalde1, Matthew J Grant3, C Simon Bawden5, Skye R Rudiger5, Amin Haghani4, Donna M Bond1, Reuben R Hore6, Michael Garratt1, Karen E Sears7, Nan Wang8, Xiangdong William Yang8,9, Russell G Snell3, Timothy A Hore1†*, Steve Horvath4†* 1Department of Anatomy, University of Otago, Dunedin, New Zealand; 2Department of Biostatistics, Fielding School of Public Health, University of California, Los Angeles, Los Angeles, United States; 3Applied Translational Genetics Group, School of Biological Sciences, Centre for Brain Research, The University of Auckland, Auckland, New Zealand; 4Department of Human Genetics, David Geffen School of Medicine, University of California, Los Angeles, Los Angeles, United States; 5Livestock and Farming Systems, South Australian Research and Development Institute, Roseworthy, Australia; 6Blackstone Hill Station, Becks, RD2, Omakau, New Zealand; 7Department of Ecology and Evolutionary Biology, UCLA, Los Angeles, United States; 8Center for Neurobehavioral Genetics, Semel Institute for Neuroscience and Human Behavior, University of California, Los Angeles (UCLA), Los Angeles, United States; 9Department of Psychiatry and Biobehavioral Sciences, David Geffen School of Medicine at UCLA, Los Angeles, United States *For correspondence: Abstract In mammals, females generally live longer than males. Nevertheless, the mechanisms [email protected] (TAH); underpinning sex-dependent longevity are currently unclear. Epigenetic clocks are powerful [email protected] (SH) biological biomarkers capable of precisely estimating chronological age and identifying novel †These authors contributed factors influencing the aging rate using only DNA methylation data. In this study, we developed the equally to this work first epigenetic clock for domesticated sheep (Ovis aries), which can predict chronological age with a median absolute error of 5.1 months. -

PRODUCT SPECIFICATION Product Datasheet
Product Datasheet QPrEST PRODUCT SPECIFICATION Product Name QPrEST MARE3 Mass Spectrometry Protein Standard Product Number QPrEST38970 Protein Name Microtubule-associated protein RP/EB family member 3 Uniprot ID Q9UPY8 Gene MAPRE3 Product Description Stable isotope-labeled standard for absolute protein quantification of Microtubule- associated protein RP/EB family member 3. Lys (13C and 15N) and Arg (13C and 15N) metabolically labeled recombinant human protein fragment. Application Absolute protein quantification using mass spectrometry Sequence (excluding YDGKDYNPLLARQGQDVAPPPNPGDQIFNKSKKLIGTAVPQRTSPTGPKN fusion tag) MQTSGRLSNVAPPCILRKNPPSARNGGHETDAQILELNQQLVDLKLTVDG LEKERDFYFSKLR Theoretical MW 30368 Da including N-terminal His6ABP fusion tag Fusion Tag A purification and quantification tag (QTag) consisting of a hexahistidine sequence followed by an Albumin Binding Protein (ABP) domain derived from Streptococcal Protein G. Expression Host Escherichia coli LysA ArgA BL21(DE3) Purification IMAC purification Purity >90% as determined by Bioanalyzer Protein 230 Purity Assay Isotopic Incorporation >99% Concentration >5 μM after reconstitution in 100 μl H20 Concentration Concentration determined by LC-MS/MS using a highly pure amino acid analyzed internal Determination reference (QTag), CV ≤10%. Amount >0.5 nmol per vial, two vials supplied. Formulation Lyophilized in 100 mM Tris-HCl 5% Trehalose, pH 8.0 Instructions for Spin vial before opening. Add 100 μL ultrapure H2O to the vial. Vortex thoroughly and spin Reconstitution down. For further dilution, see Application Protocol. Shipping Shipped at ambient temperature Storage Lyophilized product shall be stored at -20°C. See COA for expiry date. Reconstituted product can be stored at -20°C for up to 4 weeks. Avoid repeated freeze-thaw cycles. Notes For research use only Product of Sweden. -

Predicted Coronavirus Nsp5 Protease Cleavage Sites in The
bioRxiv preprint doi: https://doi.org/10.1101/2021.06.08.447224; this version posted June 8, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Predicted Coronavirus Nsp5 Protease Cleavage Sites in the 2 Human Proteome: A Resource for SARS-CoV-2 Research 3 Benjamin M. Scott1,2*, Vincent Lacasse3, Ditte G. Blom4, Peter D. Tonner5, Nikolaj S. Blom6 4 1 Associate, Biosystems and Biomaterials Division, National Institute of Standards and Technology, 5 Gaithersburg, Maryland, USA. 6 2 Department of Chemistry and Biochemistry, University of Maryland, College Park, Maryland, USA. 7 3 Segal Cancer Proteomics Centre, Lady Davis Institute, Jewish General Hospital, McGill University, Montreal, 8 Quebec, Canada 9 4 Department of Applied Mathematics and Computer Science, Technical University of Denmark, Lyngby, 10 Denmark. 11 5 Statistical Engineering Division, National Institute of Standards and Technology, Gaithersburg, Maryland, USA 12 6 Department of Bioengineering, Kongens Lyngby, Technical University of Denmark 13 14 *Corresponding author, [email protected] 15 16 BMS current affiliation: Concordia University, Centre for Applied Synthetic Biology, Montreal, Quebec, Canada 17 18 19 Abstract 20 Background: The coronavirus nonstructural protein 5 (Nsp5) is a cysteine protease required for 21 processing the viral polyprotein and is therefore crucial for viral replication. Nsp5 from several 22 coronaviruses have also been found to cleave host proteins, disrupting molecular pathways 23 involved in innate immunity. Nsp5 from the recently emerged SARS-CoV-2 virus interacts with 24 and can cleave human proteins, which may be relevant to the pathogenesis of COVID-19. -

Idep: an Integrated Web Application for Differential Expression and Pathway Analysis of RNA-Seq Data Steven Xijin Ge* , Eun Wo Son and Runan Yao
Ge et al. BMC Bioinformatics (2018) 19:534 https://doi.org/10.1186/s12859-018-2486-6 METHODOLOGY ARTICLE Open Access iDEP: an integrated web application for differential expression and pathway analysis of RNA-Seq data Steven Xijin Ge* , Eun Wo Son and Runan Yao Abstract Background: RNA-seq is widely used for transcriptomic profiling, but the bioinformatics analysis of resultant data can be time-consuming and challenging, especially for biologists. We aim to streamline the bioinformatic analyses of gene-level data by developing a user-friendly, interactive web application for exploratory data analysis, differential expression, and pathway analysis. Results: iDEP (integrated Differential Expression and Pathway analysis) seamlessly connects 63 R/Bioconductor packages, 2 web services, and comprehensive annotation and pathway databases for 220 plant and animal species. The workflow can be reproduced by downloading customized R code and related pathway files. As an example, we analyzed an RNA-Seq dataset of lung fibroblasts with Hoxa1 knockdown and revealed the possible roles of SP1 and E2F1 and their target genes, including microRNAs, in blocking G1/S transition. In another example, our analysis shows that in mouse B cells without functional p53, ionizing radiation activates the MYC pathway and its downstream genes involved in cell proliferation, ribosome biogenesis, and non-coding RNA metabolism. In wildtype B cells, radiation induces p53-mediated apoptosis and DNA repair while suppressing the target genes of MYC and E2F1, and leads to growth and cell cycle arrest. iDEP helps unveil the multifaceted functions of p53 and the possible involvement of several microRNAs such as miR-92a, miR-504, and miR-30a. -

Identification of Endogenous Adenomatous Polyposis Coli Interaction Partners and Β-Catenin−Independent Targets by Proteomics
Published OnlineFirst June 3, 2019; DOI: 10.1158/1541-7786.MCR-18-1154 Cancer Genes and Networks Molecular Cancer Research Identification of Endogenous Adenomatous Polyposis Coli Interaction Partners and b-Catenin– Independent Targets by Proteomics Olesja Popow1,2,3,Joao~ A. Paulo2, Michael H. Tatham4, Melanie S. Volk3, Alejandro Rojas-Fernandez5, Nicolas Loyer3, Ian P. Newton3, Jens Januschke3, Kevin M. Haigis1,6, and Inke Nathke€ 3 Abstract Adenomatous Polyposis Coli (APC) is the most frequently tion between endogenous MINK1 and APC and further mutated gene in colorectal cancer. APC negatively regulates confirmed the negative, and b-catenin–independent, regu- the Wnt signaling pathway by promoting the degradation of lation of MINK1 by APC. Increased Mink1/Msn levels were b-catenin, but the extent to which APC exerts Wnt/b-cate- also observed in mouse intestinal tissue and Drosophila nin–independent tumor-suppressive activity is unclear. To follicular cells expressing mutant Apc/APC when compared identify interaction partners and b-catenin–independent with wild-type tissue/cells. Collectively, our results highlight targets of endogenous, full-length APC, we applied label- the extent and importance of Wnt-independent APC func- free and multiplexed tandem mass tag-based mass spec- tions in epithelial biology and disease. trometry. Affinity enrichment-mass spectrometry identified more than 150 previously unidentified APC interaction Implications: The tumor-suppressive function of APC, the partners. Moreover, our global proteomic analysis revealed most frequently mutated gene in colorectal cancer, is mainly that roughly half of the protein expression changes that attributed to its role in b-catenin/Wnt signaling. Our study occur in response to APC loss are independent of b-catenin.