OTX2 Duplication Is Implicated in Hemifacial Microsomia
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

EFA6A, an Exchange Factor for Arf6, Regulates Early Steps in Ciliogenesis
EFA6A, an exchange factor for Arf6, regulates early steps in ciliogenesis Mariagrazia Partisani, Carole Baron, Rania Ghossoub, Racha Fayad, Sophie Pagnotta, Sophie Abélanet, Eric Macia, Frédéric Brau, Sandra Lacas-Gervais, Alexandre Benmerah, et al. To cite this version: Mariagrazia Partisani, Carole Baron, Rania Ghossoub, Racha Fayad, Sophie Pagnotta, et al.. EFA6A, an exchange factor for Arf6, regulates early steps in ciliogenesis. Journal of Cell Science, Company of Biologists, 2021, 134 (2), pp.jcs249565. 10.1242/jcs.249565. hal-03120586 HAL Id: hal-03120586 https://hal.archives-ouvertes.fr/hal-03120586 Submitted on 19 Apr 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. © 2021. Published by The Company of Biologists Ltd | Journal of Cell Science (2021) 134, jcs249565. doi:10.1242/jcs.249565 RESEARCH ARTICLE EFA6A, an exchange factor for Arf6, regulates early steps in ciliogenesis Mariagrazia Partisani1, Carole L. Baron1, Rania Ghossoub2, Racha Fayad1, Sophie Pagnotta3, Sophie Abélanet1, Eric Macia1,Frédéric Brau1, Sandra Lacas-Gervais3, Alexandre Benmerah4,Frédéric Luton1 and Michel Franco1,* ABSTRACT localized to the PC and which play an important role in its assembly Ciliogenesis is a coordinated process initiated by the recruitment and and/or function. -

E-Cadherin Accumulation Within the Lymphovascular Embolus of Inflammatory Breast Cancer Is Due to Altered Trafficking
ANTICANCER RESEARCH 30: 3903-3910 (2010) E-Cadherin Accumulation within the Lymphovascular Embolus of Inflammatory Breast Cancer Is Due to Altered Trafficking YIN YE1, JOSEPH D. TELLEZ1, MARIA DURAZO1, MEAGAN BELCHER1, KURTIS YEARSLEY2 and SANFORD H. BARSKY1,3,4 1Department of Pathology, University of Nevada School of Medicine, Reno, NV 89557, U.S.A.; 2Department of Pathology, Ohio State University, Columbus, OH 43210, U.S.A.; 3Department of Pathology, The Whittemore-Peterson Institute, Reno, NV 89557, U.S.A.; 4Department of Pathology, Nevada Cancer Institute, Las Vegas, NV 89135, U.S.A. Abstract. E-Cadherin functions as a tumor suppressor in of the 95 KD band were observed. These findings suggest some invasive breast carcinomas and metastasis is promoted that it is the altered E-cadherin trafficking that contributes when its expression is lost. It has been observed, however, to its oncogenic rather than suppressive role in IBC. that in one of the most aggressive human breast cancers, inflammatory breast cancer (IBC), E-cadherin is E-Cadherin, an adhesion protein present in normal overexpressed and this accounts for the formation of the epithelial cells within lateral junctions (zona adherens), is lymphovascular embolus, a structure efficient at metastasis thought to function as a tumor suppressor in certain types and resistant to chemotherapy through unknown of invasive breast carcinomas and metastasis is promoted cytoprotective mechanisms. Studies using a human xenograft when its expression is lost by gene mutation, promoter model of IBC, MARY-X, indicate that the mechanism of E- methylation or promoter repression by snail/slug and other cadherin overexpression is not transcriptional but related to mediators of epithelial-mesenchymal transition (EMT) (1- altered protein trafficking. -
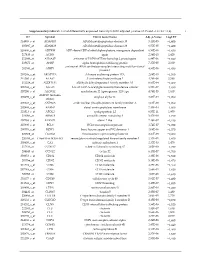
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

A Regulator of Innate Immune Responses
(19) TZZ ¥_T (11) EP 2 942 357 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 11.11.2015 Bulletin 2015/46 C07K 14/47 (2006.01) A61K 38/00 (2006.01) C12N 15/113 (2010.01) (21) Application number: 15169327.2 (22) Date of filing: 04.08.2009 (84) Designated Contracting States: (72) Inventor: Barber, Glen N. AT BE BG CH CY CZ DE DK EE ES FI FR GB GR Palmetto Bay, FL 33157 (US) HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO SE SI SK SM TR (74) Representative: Inspicos A/S Kogle Allé 2 (30) Priority: 04.08.2008 US 129975 P P.O. Box 45 2970 Hørsholm (DK) (62) Document number(s) of the earlier application(s) in accordance with Art. 76 EPC: Remarks: 09805473.7 / 2 324 044 This application was filed on 27-05-2015 as a divisional application to the application mentioned (71) Applicant: Barber, Glen N. under INID code 62. Palmetto Bay, FL 33157 (US) (54) STING (STIMULATOR OF INTEFERON GENES), A REGULATOR OF INNATE IMMUNE RESPONSES (57) Novel molecules termed STING which include STING compositions are useful for the treatment of an nucleic acids, polynucleotides, oligonucleotides, pep- immune-related disorder, including treating and prevent- tides, mutants, variants and active fragments thereof, ing infection by modulating immunity. modulate innate and adaptive immunity in a subject. EP 2 942 357 A1 Printed by Jouve, 75001 PARIS (FR) EP 2 942 357 A1 Description RELATED APPLICATIONS 5 [0001] This application claims priority under 35 USC § 119 to U.S. -

Downloaded Per Proteome Cohort Via the Web- Site Links of Table 1, Also Providing Information on the Deposited Spectral Datasets
www.nature.com/scientificreports OPEN Assessment of a complete and classifed platelet proteome from genome‑wide transcripts of human platelets and megakaryocytes covering platelet functions Jingnan Huang1,2*, Frauke Swieringa1,2,9, Fiorella A. Solari2,9, Isabella Provenzale1, Luigi Grassi3, Ilaria De Simone1, Constance C. F. M. J. Baaten1,4, Rachel Cavill5, Albert Sickmann2,6,7,9, Mattia Frontini3,8,9 & Johan W. M. Heemskerk1,9* Novel platelet and megakaryocyte transcriptome analysis allows prediction of the full or theoretical proteome of a representative human platelet. Here, we integrated the established platelet proteomes from six cohorts of healthy subjects, encompassing 5.2 k proteins, with two novel genome‑wide transcriptomes (57.8 k mRNAs). For 14.8 k protein‑coding transcripts, we assigned the proteins to 21 UniProt‑based classes, based on their preferential intracellular localization and presumed function. This classifed transcriptome‑proteome profle of platelets revealed: (i) Absence of 37.2 k genome‑ wide transcripts. (ii) High quantitative similarity of platelet and megakaryocyte transcriptomes (R = 0.75) for 14.8 k protein‑coding genes, but not for 3.8 k RNA genes or 1.9 k pseudogenes (R = 0.43–0.54), suggesting redistribution of mRNAs upon platelet shedding from megakaryocytes. (iii) Copy numbers of 3.5 k proteins that were restricted in size by the corresponding transcript levels (iv) Near complete coverage of identifed proteins in the relevant transcriptome (log2fpkm > 0.20) except for plasma‑derived secretory proteins, pointing to adhesion and uptake of such proteins. (v) Underrepresentation in the identifed proteome of nuclear‑related, membrane and signaling proteins, as well proteins with low‑level transcripts. -

Regulation of Human Cerebral Cortical Development by EXOC7
ARTICLE Regulation of human cerebral cortical development by EXOC7 and EXOC8, components of the exocyst complex, and roles in neural progenitor cell proliferation and survival Michael E. Coulter, MD, PhD1,2, Damir Musaev, BSc3, Ellen M. DeGennaro, BA1,4, Xiaochang Zhang, PhD1,5, Katrin Henke, PhD6, Kiely N. James, PhD3, Richard S. Smith, PhD1, R. Sean Hill, PhD1, Jennifer N. Partlow, MS1, Muna Al-Saffar, MBChB, MSc1,7, A. Stacy Kamumbu, BA1, Nicole Hatem, BA1, A. James Barkovich, MD8, Jacqueline Aziza, MD9, Nicolas Chassaing, MD, PhD10,11, Maha S. Zaki, MD, PhD12, Tipu Sultan, MD13, Lydie Burglen, MD, PhD14,15, Anna Rajab, MD, PhD16, Lihadh Al-Gazali, MBChB, MSc7, Ganeshwaran H. Mochida, MD, MMSc1,17, Matthew P. Harris, PhD6, Joseph G. Gleeson, MD3 and Christopher A. Walsh, MD, PhD 1 Purpose: The exocyst complex is a conserved protein complex that (LOF) variants in a recessively inherited disorder characterized by mediates fusion of intracellular vesicles to the plasma membrane brain atrophy, seizures, and developmental delay, and in severe and is implicated in processes including cell polarity, cell migration, cases, microcephaly and infantile death. In EXOC8, we found a ciliogenesis, cytokinesis, autophagy, and fusion of secretory vesicles. homozygous truncating variant in a family with a similar clinical The essential role of these genes in human genetic disorders, disorder. We modeled exoc7 deficiency in zebrafish and found the however, is unknown. absence of exoc7 causes microcephaly. Methods: We performed homozygosity mapping and exome Conclusion: Our results highlight the essential role of the exocyst sequencing of consanguineous families with recessively inherited pathway in normal cortical development and how its perturbation brain development disorders. -

BMC Proceedings Biomed Central
BMC Proceedings BioMed Central Research Open Access Pathway results from the chicken data set using GOTM, Pathway Studio and Ingenuity softwares Agnès Bonnet1, Sandrine Lagarrigue2,3, Laurence Liaubet1, Christèle Robert- Granié4, Magali SanCristobal1 and Gwenola Tosser-Klopp*1 Address: 1INRA, UMR444, Laboratoire de Génétique Cellulaire, F-31326 Castanet-Tolosan, France, 2INRA, UMR 598, Génétique Animale, F-35000 Rennes, France, 3Agrocampus Ouest, UMR 598 Génétique Animale, F-35000 Rennes, France and 4INRA, UR631, Station d'Amélioration Génétique des Animaux, F-31326 Castanet-Tolosan, France Email: Agnès Bonnet - [email protected]; Sandrine Lagarrigue - [email protected]; Laurence Liaubet - [email protected]; Christèle Robert-Granié - [email protected]; Magali SanCristobal - [email protected]; Gwenola Tosser-Klopp* - [email protected] * Corresponding author from EADGENE and SABRE Post-analyses Workshop Lelystad, The Netherlands. 12–14 November 2008 Published: 16 July 2009 BMC Proceedings 2009, 3(Suppl 4):S11 doi:10.1186/1753-6561-3-S4-S11 <supplement> <title> <p>EADGENE and SABRE Post-analyses Workshop</p> </title> <editor>Dirk-Jan de Koning</editor> <sponsor> <note>The publication of these proceedings was supported by the EC-funded Network of Excellence EADGENE (EC contract number FOOD-CT-2004-506416).</note> </sponsor> <note>Proceedings</note> <url>http://www.biomedcentral.com/content/pdf/1753-6561-3-S4-info.pdf</url> </supplement> This article is available from: http://www.biomedcentral.com/1753-6561/3/S4/S11 © 2009 Bonnet et al; licensee BioMed Central Ltd. This is an open access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. -

The Role of the Exocyst in Renal Ciliogenesis, Cystogenesis, Tubulogenesis, and Development Joshua H
Review Article KIDNEY RESEARCH Kidney Res Clin Pract 2019;38(3):260-266 AND pISSN: 2211-9132 • eISSN: 2211-9140 CLINICAL PRACTICE https://doi.org/10.23876/j.krcp.19.050 The role of the exocyst in renal ciliogenesis, cystogenesis, tubulogenesis, and development Joshua H. Lipschutz1,2 1Department of Medicine, Medical University of South Carolina, Charleston, SC, USA 2Department of Medicine, Ralph H. Johnson Veterans Affairs Medical Center, Charleston, SC, USA The exocyst is a highly conserved eight-subunit protein complex (EXOC1-8) involved in the targeting and docking of exocytic vesicles translocating from the trans-Golgi network to various sites in renal cells. EXOC5 is a central exocyst component because it connects EXOC6, bound to the vesicles exiting the trans-Golgi network via the small GTPase RAB8, to the rest of the exocyst complex at the plasma membrane. In the kidney, the exocyst complex is involved in primary ciliognesis, cystogenesis, and tubulogenesis. The exocyst, and its regulators, have also been found in urinary extracellular vesicles, and may be centrally involved in urocrine signaling and repair following acute kidney injury. The exocyst is centrally involved in the development of other organs, including the eye, ear, and heart. The exocyst is regulated by many different small GTPases of the RHO, RAL, RAB, and ARF families. The small GTPases, and their guanine nucleotide exchange factors and GTPase-activating proteins, likely give the exocyst specificity of function. The recent development of a floxed Exoc5 mouse line will aid researchers in studying the role of the exocyst in multiple cells and organ types by allowing for tissue-specific knockout, in conjunction with Cre-driver mouse lines. -

Patterns of Sequence Conservation in Presynaptic Neural Genes
University of Pennsylvania ScholarlyCommons Departmental Papers (CIS) Department of Computer & Information Science November 2006 Patterns of Sequence Conservation in Presynaptic Neural Genes Dexter Hadley University of Pennsylvania Tara Murphy University of Pennsylvania Otto Valladares University of Pennsylvania Sridhar Hannenhalli University of Pennsylvania Lyle H. Ungar University of Pennsylvania, [email protected] See next page for additional authors Follow this and additional works at: https://repository.upenn.edu/cis_papers Recommended Citation Dexter Hadley, Tara Murphy, Otto Valladares, Sridhar Hannenhalli, Lyle H. Ungar, Junhyong Kim, and Maja Bucan, "Patterns of Sequence Conservation in Presynaptic Neural Genes", . November 2006. Reprinted from Genome Biology, Volume 7, Issue 11, November 2006, pages R105.1-R105.19. Publisher URL: http://genomebiology.com/2006/7/11/R105 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/cis_papers/282 For more information, please contact [email protected]. Patterns of Sequence Conservation in Presynaptic Neural Genes Abstract Background: The neuronal synapse is a fundamental functional unit in the central nervous system of animals. Because synaptic function is evolutionarily conserved, we reasoned that functional sequences of genes and related genomic elements known to play important roles in neurotransmitter release would also be conserved. Results: Evolutionary rate analysis revealed that presynaptic proteins evolve slowly, although some members of large gene families exhibit accelerated evolutionary rates relative to other family members. Comparative sequence analysis of 46 megabases spanning 150 presynaptic genes identified more than 26,000 elements that are highly conserved in eight vertebrate species, as well as a small subset of sequences (6%) that are shared among unrelated presynaptic genes. -

Prion Characterization Using Cell Based Approaches
University of Kentucky UKnowledge Theses and Dissertations--Microbiology, Microbiology, Immunology, and Molecular Immunology, and Molecular Genetics Genetics 2012 PRION CHARACTERIZATION USING CELL BASED APPROACHES Vadim Khaychuk University of Kentucky, [email protected] Right click to open a feedback form in a new tab to let us know how this document benefits ou.y Recommended Citation Khaychuk, Vadim, "PRION CHARACTERIZATION USING CELL BASED APPROACHES" (2012). Theses and Dissertations--Microbiology, Immunology, and Molecular Genetics. 2. https://uknowledge.uky.edu/microbio_etds/2 This Doctoral Dissertation is brought to you for free and open access by the Microbiology, Immunology, and Molecular Genetics at UKnowledge. It has been accepted for inclusion in Theses and Dissertations--Microbiology, Immunology, and Molecular Genetics by an authorized administrator of UKnowledge. For more information, please contact [email protected]. STUDENT AGREEMENT: I represent that my thesis or dissertation and abstract are my original work. Proper attribution has been given to all outside sources. I understand that I am solely responsible for obtaining any needed copyright permissions. I have obtained and attached hereto needed written permission statements(s) from the owner(s) of each third-party copyrighted matter to be included in my work, allowing electronic distribution (if such use is not permitted by the fair use doctrine). I hereby grant to The University of Kentucky and its agents the non-exclusive license to archive and make accessible my work in whole or in part in all forms of media, now or hereafter known. I agree that the document mentioned above may be made available immediately for worldwide access unless a preapproved embargo applies. -

Novel Skin Phenotypes Revealed by a Genome-Wide Mouse Reverse Genetic Screen
ARTICLE Received 9 Jan 2014 | Accepted 4 Mar 2014 | Published 11 Apr 2014 DOI: 10.1038/ncomms4540 OPEN Novel skin phenotypes revealed by a genome-wide mouse reverse genetic screen Kifayathullah Liakath-Ali1,2,3, Valerie E. Vancollie4, Emma Heath1, Damian P. Smedley4, Jeanne Estabel4, David Sunter4, Tia DiTommaso5,w, Jacqueline K. White4, Ramiro Ramirez-Solis4, Ian Smyth5, Karen P. Steel4,6 & Fiona M. Watt1 Permanent stop-and-shop large-scale mouse mutant resources provide an excellent platform to decipher tissue phenogenomics. Here we analyse skin from 538 knockout mouse mutants generated by the Sanger Institute Mouse Genetics Project. We optimize immunolabelling of tail epidermal wholemounts to allow systematic annotation of hair follicle, sebaceous gland and interfollicular epidermal abnormalities using ontology terms from the Mammalian Phenotype Ontology. Of the 50 mutants with an epidermal phenotype, 9 map to human genetic conditions with skin abnormalities. Some mutant genes are expressed in the skin, whereas others are not, indicating systemic effects. One phenotype is affected by diet and several are incompletely penetrant. In-depth analysis of three mutants, Krt76, Myo5a (a model of human Griscelli syndrome) and Mysm1, provides validation of the screen. Our study is the first large-scale genome-wide tissue phenotype screen from the International Knockout Mouse Consortium and provides an open access resource for the scientific community. 1 Centre for Stem Cells and Regenerative Medicine, King’s College London, Guy’s Hospital, London SE1 9RT, UK. 2 Department of Biochemistry, University of Cambridge, Tennis Court Road, Cambridge CB2 1QW, UK. 3 Wellcome Trust—Medical Research Council Stem Cell Institute, University of Cambridge, Tennis Court Road, Cambridge CB2 1QR, UK. -
Whole-Genome Sequencing of European Autochthonous and Commercial Pig Breeds Allows the Detection of Signatures of Selection
Whole-genome sequencing of European autochthonous and commercial pig breeds allows the detection of signatures of selection for adaptation of genetic resources to different breeding and production systems Samuele Bovo, Anisa Ribani, Maria Muñoz, Estefania Alves, Jose P. Araujo, Riccardo Bozzi, Marjeta Čandek-Potokar, Rui Charneca, Federica Di Palma, Graham Etherington, et al. To cite this version: Samuele Bovo, Anisa Ribani, Maria Muñoz, Estefania Alves, Jose P. Araujo, et al.. Whole-genome sequencing of European autochthonous and commercial pig breeds allows the detection of signatures of selection for adaptation of genetic resources to different breeding and production systems. Genetics Se- lection Evolution, BioMed Central, 2020, 52 (1), pp.33. 10.1186/s12711-020-00553-7. hal-02883278 HAL Id: hal-02883278 https://hal.archives-ouvertes.fr/hal-02883278 Submitted on 29 Jun 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Bovo et al. Genet Sel Evol (2020) 52:33 https://doi.org/10.1186/s12711-020-00553-7 Genetics Selection Evolution RESEARCH ARTICLE Open Access Whole-genome sequencing of European autochthonous and commercial pig breeds allows the detection of signatures of selection for adaptation of genetic resources to diferent breeding and production systems Samuele Bovo1, Anisa Ribani1, Maria Muñoz2, Estefania Alves2, Jose P.