Genome-Wide Discovery of Somatic Coding and Regulatory Variants In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Environmental Influences on Endothelial Gene Expression
ENDOTHELIAL CELL GENE EXPRESSION John Matthew Jeff Herbert Supervisors: Prof. Roy Bicknell and Dr. Victoria Heath PhD thesis University of Birmingham August 2012 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT Tumour angiogenesis is a vital process in the pathology of tumour development and metastasis. Targeting markers of tumour endothelium provide a means of targeted destruction of a tumours oxygen and nutrient supply via destruction of tumour vasculature, which in turn ultimately leads to beneficial consequences to patients. Although current anti -angiogenic and vascular targeting strategies help patients, more potently in combination with chemo therapy, there is still a need for more tumour endothelial marker discoveries as current treatments have cardiovascular and other side effects. For the first time, the analyses of in-vivo biotinylation of an embryonic system is performed to obtain putative vascular targets. Also for the first time, deep sequencing is applied to freshly isolated tumour and normal endothelial cells from lung, colon and bladder tissues for the identification of pan-vascular-targets. Integration of the proteomic, deep sequencing, public cDNA libraries and microarrays, delivers 5,892 putative vascular targets to the science community. -

Genome-Wide Screen of Cell-Cycle Regulators in Normal and Tumor Cells
bioRxiv preprint doi: https://doi.org/10.1101/060350; this version posted June 23, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Genome-wide screen of cell-cycle regulators in normal and tumor cells identifies a differential response to nucleosome depletion Maria Sokolova1, Mikko Turunen1, Oliver Mortusewicz3, Teemu Kivioja1, Patrick Herr3, Anna Vähärautio1, Mikael Björklund1, Minna Taipale2, Thomas Helleday3 and Jussi Taipale1,2,* 1Genome-Scale Biology Program, P.O. Box 63, FI-00014 University of Helsinki, Finland. 2Science for Life laboratory, Department of Biosciences and Nutrition, Karolinska Institutet, SE- 141 83 Stockholm, Sweden. 3Science for Life laboratory, Division of Translational Medicine and Chemical Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 21 Stockholm, Sweden To identify cell cycle regulators that enable cancer cells to replicate DNA and divide in an unrestricted manner, we performed a parallel genome-wide RNAi screen in normal and cancer cell lines. In addition to many shared regulators, we found that tumor and normal cells are differentially sensitive to loss of the histone genes transcriptional regulator CASP8AP2. In cancer cells, loss of CASP8AP2 leads to a failure to synthesize sufficient amount of histones in the S-phase of the cell cycle, resulting in slowing of individual replication forks. Despite this, DNA replication fails to arrest, and tumor cells progress in an elongated S-phase that lasts several days, finally resulting in death of most of the affected cells. -

Histone-Related Genes Are Hypermethylated in Lung Cancer
Published OnlineFirst October 1, 2019; DOI: 10.1158/0008-5472.CAN-19-1019 Cancer Genome and Epigenome Research Histone-Related Genes Are Hypermethylated in Lung Cancer and Hypermethylated HIST1H4F Could Serve as a Pan-Cancer Biomarker Shihua Dong1,Wei Li1, Lin Wang2, Jie Hu3,Yuanlin Song3, Baolong Zhang1, Xiaoguang Ren1, Shimeng Ji3, Jin Li1, Peng Xu1, Ying Liang1, Gang Chen4, Jia-Tao Lou2, and Wenqiang Yu1 Abstract Lung cancer is the leading cause of cancer-related deaths lated in all 17 tumor types from TCGA datasets (n ¼ 7,344), worldwide. Cytologic examination is the current "gold stan- which was further validated in nine different types of cancer dard" for lung cancer diagnosis, however, this has low sensi- (n ¼ 243). These results demonstrate that HIST1H4F can tivity. Here, we identified a typical methylation signature of function as a universal-cancer-only methylation (UCOM) histone genes in lung cancer by whole-genome DNA methyl- marker, which may aid in understanding general tumorigen- ation analysis, which was validated by The Cancer Genome esis and improve screening for early cancer diagnosis. Atlas (TCGA) lung cancer cohort (n ¼ 907) and was further confirmed in 265 bronchoalveolar lavage fluid samples with Significance: These findings identify a new biomarker for specificity and sensitivity of 96.7% and 87.0%, respectively. cancer detection and show that hypermethylation of histone- More importantly, HIST1H4F was universally hypermethy- related genes seems to persist across cancers. Introduction to its low specificity, LDCT is far from satisfactory as a screening tool for clinical application, similar to other currently used cancer Lung cancer is one of the most common malignant tumors and biomarkers, such as carcinoembryonic antigen (CEA), neuron- the leading cause of cancer-related deaths worldwide (1, 2). -

Transcriptomic Profiles of High and Low Antibody Responders to Smallpox
Genes and Immunity (2013) 14, 277–285 & 2013 Macmillan Publishers Limited All rights reserved 1466-4879/13 www.nature.com/gene ORIGINAL ARTICLE Transcriptomic profiles of high and low antibody responders to smallpox vaccine RB Kennedy1,2, AL Oberg1,3, IG Ovsyannikova1,2, IH Haralambieva1,2, D Grill1,3 and GA Poland1,2 Despite its eradication over 30 years ago, smallpox (as well as other orthopox viruses) remains a pathogen of interest both in terms of biodefense and for its use as a vector for vaccines and immunotherapies. Here we describe the application of mRNA-Seq transcriptome profiling to understanding immune responses in smallpox vaccine recipients. Contrary to other studies examining gene expression in virally infected cell lines, we utilized a mixed population of peripheral blood mononuclear cells in order to capture the essential intercellular interactions that occur in vivo, and would otherwise be lost, using single cell lines or isolated primary cell subsets. In this mixed cell population we were able to detect expression of all annotated vaccinia genes. On the host side, a number of genes encoding cytokines, chemokines, complement factors and intracellular signaling molecules were downregulated upon viral infection, whereas genes encoding histone proteins and the interferon response were upregulated. We also identified a small number of genes that exhibited significantly different expression profiles in subjects with robust humoral immunity compared with those with weaker humoral responses. Our results provide evidence that differential gene regulation patterns may be at work in individuals with robust humoral immunity compared with those with weaker humoral immune responses. Genes and Immunity (2013) 14, 277–285; doi:10.1038/gene.2013.14; published online 18 April 2013 Keywords: Next-generation sequencing; mRNA-Seq; vaccinia virus; smallpox vaccine INTRODUCTION these 44 subjects had two samples (uninfected and vaccinia Vaccinia virus (VACV) is the immunologically cross-protective infected). -

Proquest Dissertations
IMPROVEMENTS TO FLUORESCENT AFFINITY LABELS AND THE RIBONUCLEASE S SYSTEM AND GENE EXPRESSION RESPONSE TO CLINICALLY RELEVANT RIBONUCLEASES by Rex Wayne Watkins A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Biochemistry) at the UNIVERSITY OF WISCONSIN-MADISON 2010 UMI Number: 3437078 All rights reserved INFORMATION TO ALL USERS The quality of this reproduction is dependent upon the quality of the copy submitted. In the unlikely event that the author did not send a complete manuscript and there are missing pages, these will be noted. Also, if material had to be removed, a note will indicate the deletion. UMT Dissertation Publishing UMI 3437078 Copyright 2010 by ProQuest LLC. All rights reserved. This edition of the work is protected against unauthorized copying under Title 17, United States Code. ProQuest LLC 789 East Eisenhower Parkway P.O. Box 1346 Ann Arbor, Ml 48106-1346 IMPROVEMENTS TO FLUORESCENT AFFINITY LABELS AND THE RIBONUCLEASE S SYSTEM AND GENE EXPRESSION RESPONSE TO CLINICALLY RELEVANT RIBONUCLEASES submitted to the Graduate School of the University of Wisconsin-Madison in partial fulfillment of the requirements for the degree of Doctor of Philosophy By Rex Wayne Watkins Date of final oral examination: ] g August 2010 Month and year degree to be awarded: August 2010 The dissertation is approved by the following members of the Final Oral Committee: Ronald T. Raines, Professor, Biochemistry Aseem Z. Ansari, Professor, Biochemistry Laura L. Kiessling, Professor, Biochemistry Douglas B. Weibel, Professor, Biochemistry Jon S. Thorson, Professor, Pharmaceutical Sciences 1 IMPROVEMENTS TO FLUORESCENT AFFINITY LABELS AND THE RIBONUCLEASE S SYSTEM AND GENE EXPRESSION RESPONSE TO CLINICALLY RELEVANT RIBONUCLEASES Rex Wayne Watkins Under the supervision of Professor Ronald T. -

Rap1-Mediated Chromatin and Gene Expression Changes at Senescence
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2019 Rap1-Mediated Chromatin And Gene Expression Changes At Senescence Shufei Song University of Pennsylvania Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Biochemistry Commons, and the Cell Biology Commons Recommended Citation Song, Shufei, "Rap1-Mediated Chromatin And Gene Expression Changes At Senescence" (2019). Publicly Accessible Penn Dissertations. 3557. https://repository.upenn.edu/edissertations/3557 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/3557 For more information, please contact [email protected]. Rap1-Mediated Chromatin And Gene Expression Changes At Senescence Abstract ABSTRACT RAP1-MEDIATED CHROMATIN AND GENE EXPRESSION CHANGES AT SENESCENCE The telomeric protein Rap1 has been extensively studied for its roles as a transcriptional activator and repressor. Indeed, in both yeast and mammals, Rap1 is known to bind throughout the genome to reorganize chromatin and regulate gene transcription. Previously, our lab published evidence that Rap1 plays important roles in cellular senescence. In telomerase-deficient S. cerevisiae, Rap1 relocalizes from telomeres and subtelomeres to new Rap1 target at senescence (NRTS). This leads to two types of histone loss: Rap1 lowers global histone levels by repressing histone gene transcription and it also results in local nucleosome displacement at the promoters of the activated NRTS. Here, I examine mechanisms of site-specific histone loss by presenting evidence that Rap1 can directly interact with histone tetramers H3/H4, and map this interaction to a three-amino-acid-patch within the DNA binding domain. Functional studies are performed in vivo using a mutant form of Rap1 with weakened histone interactions, and deficient promoter clearance as well as blunted gene activation is observed, indicating that direct Rap1-H3/H4 interactions are involved in nucleosome displacement. -

HIST1H2AK (HIST1H2AM) (NM 003514) Human Recombinant Protein Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TP760612 HIST1H2AK (HIST1H2AM) (NM_003514) Human Recombinant Protein Product data: Product Type: Recombinant Proteins Description: Purified recombinant protein of Human histone cluster 1, H2am (HIST1H2AM), full length, with N-terminal HIS tag, expressed in E.Coli, 50ug Species: Human Expression Host: E. coli Tag: N-His Predicted MW: 13.9 kDa Concentration: >50 ug/mL as determined by microplate BCA method Purity: > 80% as determined by SDS-PAGE and Coomassie blue staining Buffer: 50mM Tris,8M Urea,pH8.0. Storage: Store at -80°C. Stability: Stable for 12 months from the date of receipt of the product under proper storage and handling conditions. Avoid repeated freeze-thaw cycles. RefSeq: NP_003505 Locus ID: 8336 UniProt ID: P0C0S8, A4FTV9 RefSeq Size: 487 Cytogenetics: 6p22.1 RefSeq ORF: 390 Synonyms: dJ193B12.1; H2A.1; H2A/n; H2AC11; H2AC13; H2AC15; H2AC16; H2AFN; HIST1H2AM This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 HIST1H2AK (HIST1H2AM) (NM_003514) Human Recombinant Protein – TP760612 Summary: Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Two molecules of each of the four core histones (H2A, H2B, H3, and H4) form an octamer, around which approximately 146 bp of DNA is wrapped in repeating units, called nucleosomes. -
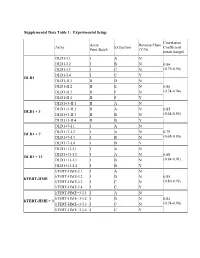
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Download Special Issue
BioMed Research International Novel Bioinformatics Approaches for Analysis of High-Throughput Biological Data Guest Editors: Julia Tzu-Ya Weng, Li-Ching Wu, Wen-Chi Chang, Tzu-Hao Chang, Tatsuya Akutsu, and Tzong-Yi Lee Novel Bioinformatics Approaches for Analysis of High-Throughput Biological Data BioMed Research International Novel Bioinformatics Approaches for Analysis of High-Throughput Biological Data Guest Editors: Julia Tzu-Ya Weng, Li-Ching Wu, Wen-Chi Chang, Tzu-Hao Chang, Tatsuya Akutsu, and Tzong-Yi Lee Copyright © 2014 Hindawi Publishing Corporation. All rights reserved. This is a special issue published in “BioMed Research International.” All articles are open access articles distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Contents Novel Bioinformatics Approaches for Analysis of High-Throughput Biological Data,JuliaTzu-YaWeng, Li-Ching Wu, Wen-Chi Chang, Tzu-Hao Chang, Tatsuya Akutsu, and Tzong-Yi Lee Volume2014,ArticleID814092,3pages Evolution of Network Biomarkers from Early to Late Stage Bladder Cancer Samples,Yung-HaoWong, Cheng-Wei Li, and Bor-Sen Chen Volume 2014, Article ID 159078, 23 pages MicroRNA Expression Profiling Altered by Variant Dosage of Radiation Exposure,Kuei-FangLee, Yi-Cheng Chen, Paul Wei-Che Hsu, Ingrid Y. Liu, and Lawrence Shih-Hsin Wu Volume2014,ArticleID456323,10pages EXIA2: Web Server of Accurate and Rapid Protein Catalytic Residue Prediction, Chih-Hao Lu, Chin-Sheng -

PDF Download
Histone H2A (Phospho Ser129) Polyclonal Antibody Catalog No : YM3277 Reactivity : Human,Mouse,Rat Applications : WB Gene Name : HIST1H2AG/HIST1H2AI/HIST1H2AK/HIST1H2AL/HIST1H2AM/HIST2H2AA3 /HIST2H2AA4/HIST3H2A Protein Name : Histone H2A type 1/Histone H2A type 2/Histone H2A type 3 Human Gene Id : 8329/8330/8332/8336/8969/723790/8337/92815 Human Swiss Prot P0C0S8/Q6FI13/Q7L7L0 No : Mouse Gene Id : 319164/15267/319162 Rat Gene Id : 365877/64646 Rat Swiss Prot No : P02262/P0CC09/Q4FZT6 Immunogen : Synthetic Peptide of Histone H2A (Phospho Ser129) Specificity : The antibody detects endogenous Histone H2A (Phospho Ser129) protein. Formulation : PBS, pH 7.4, containing 0.5%BSA, 0.02% sodium azide as Preservative and 50% Glycerol. Source : Rabbit Dilution : WB: 1:1000-2000 Purification : The antibody was affinity-purified from rabbit antiserum by affinity- chromatography using specific immunogen. Storage Stability : -20°C/1 year Molecularweight : 14091/14095/14121 1 / 3 Observed Band : 14 Cell Pathway : Systemic lupus erythematosus, Background : histone cluster 1 H2A family member i(HIST1H2AI) Homo sapiens Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Two molecules of each of the four core histones (H2A, H2B, H3, and H4) form an octamer, around which approximately 146 bp of DNA is wrapped in repeating units, called nucleosomes. The linker histone, H1, interacts with linker DNA between nucleosomes and functions in the compaction of chromatin into higher order structures. This gene is intronless and encodes a replication-dependent histone that is a member of the histone H2A family. Transcripts from this gene lack polyA tails but instead contain a palindromic termination element. -

The Changing Chromatome As a Driver of Disease: a Panoramic View from Different Methodologies
The changing chromatome as a driver of disease: A panoramic view from different methodologies Isabel Espejo1, Luciano Di Croce,1,2,3 and Sergi Aranda1 1. Centre for Genomic Regulation (CRG), Barcelona Institute of Science and Technology, Dr. Aiguader 88, Barcelona 08003, Spain 2. Universitat Pompeu Fabra (UPF), Barcelona, Spain 3. ICREA, Pg. Lluis Companys 23, Barcelona 08010, Spain *Corresponding authors: Luciano Di Croce ([email protected]) Sergi Aranda ([email protected]) 1 GRAPHICAL ABSTRACT Chromatin-bound proteins regulate gene expression, replicate and repair DNA, and transmit epigenetic information. Several human diseases are highly influenced by alterations in the chromatin- bound proteome. Thus, biochemical approaches for the systematic characterization of the chromatome could contribute to identifying new regulators of cellular functionality, including those that are relevant to human disorders. 2 SUMMARY Chromatin-bound proteins underlie several fundamental cellular functions, such as control of gene expression and the faithful transmission of genetic and epigenetic information. Components of the chromatin proteome (the “chromatome”) are essential in human life, and mutations in chromatin-bound proteins are frequently drivers of human diseases, such as cancer. Proteomic characterization of chromatin and de novo identification of chromatin interactors could thus reveal important and perhaps unexpected players implicated in human physiology and disease. Recently, intensive research efforts have focused on developing strategies to characterize the chromatome composition. In this review, we provide an overview of the dynamic composition of the chromatome, highlight the importance of its alterations as a driving force in human disease (and particularly in cancer), and discuss the different approaches to systematically characterize the chromatin-bound proteome in a global manner. -

HIST1H2AK (HIST1H2AM) (NM 003514) Human Untagged Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for SC117915 HIST1H2AK (HIST1H2AM) (NM_003514) Human Untagged Clone Product data: Product Type: Expression Plasmids Product Name: HIST1H2AK (HIST1H2AM) (NM_003514) Human Untagged Clone Tag: Tag Free Symbol: H2AC17 Synonyms: dJ193B12.1; H2A.1; H2A/n; H2AC11; H2AC13; H2AC15; H2AC16; H2AFN; HIST1H2AM Vector: pCMV6-XL5 E. coli Selection: Ampicillin (100 ug/mL) Cell Selection: None Fully Sequenced ORF: >NCBI ORF sequence for NM_003514, the custom clone sequence may differ by one or more nucleotides ATGTCTGGACGTGGCAAGCAGGGCGGCAAGGCTCGCGCCAAGGCCAAAACCCGCTCCTCTAGAGCTGGGC TCCAATTTCCTGTAGGACGAGTGCACCGCCTGCTCCGCAAGGGCAACTACGCTGAGCGGGTCGGGGCCGG CGCGCCGGTTTACCTGGCGGCGGTGCTGGAGTACCTAACTGCCGAGATCCTGGAGCTGGCGGGCAACGCA GCCCGCGACAACAAAAAGACCCGCATCATCCCGCGCCACTTGCAGCTGGCCATCCGCAACGACGAGGAGC TCAACAAGCTGCTTGGTAAAGTTACCATCGCTCAGGGCGGTGTTCTGCCTAACATCCAGGCCGTACTGCT CCCCAAGAAGACTGAGAGCCACCACAAAGCTAAGGGCAAGTAA 3' Read Nucleotide >OriGene 3' read for NM_003514 unedited Sequence: CGTCCCTANCTGATGNACCGCGGCCCGCTTTTCTANGANCGAGTTTTTTTTTTTTTTTTT TGGGGATCATAAATACTTTTAATATCAGATAAATCATTAAGAAATTGCATTCTGTACTTG ATGACCACACGGGAACCTTGCTAGAGTCAAGAGAACTTGTCACTAGTAATTATGAAGACA CCTTTACGGTGAGCGTTATTAAAACCCTACTAGAGGTTTTGGGTGGGACTCAAGAGCAAG GGGTGGCCACCTGTGGACGAGGGTTCCCTGTTGTTAACAGAACACGTTGCCCACCTCGCA AGTATGCAGCCCAATCAGTCCCCAGGGTCTCGGTTCCCGTTGCGCCCTTCCCCATGGCCA CTGCGCTCATTCATGAGCCTAGGGTGATCAGGCCTCCGGCCTATAAAGAACTGCAGGATC