A High-Resolution 3D Epigenomic Map Reveals Insights Into the Creation of the Prostate Cancer Transcriptome
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Chromosome 14 Transfer and Functional Studies Identify a Candidate Tumor Suppressor Gene, Mirror Image Polydactyly 1, in Nasopharyngeal Carcinoma
Chromosome 14 transfer and functional studies identify a candidate tumor suppressor gene, Mirror image polydactyly 1, in nasopharyngeal carcinoma Arthur Kwok Leung Cheunga, Hong Lok Lungb, Josephine Mun Yee Kob, Yue Chenga,c, Eric J. Stanbridged, Eugene R. Zabarovskye, John M. Nichollsf, Daniel Chuab, Sai Wah Tsaog, Xin-Yuan Guanb, and Maria Li Lungb,1 aDepartment of Biology and Center for Cancer Research, Hong Kong University of Science and Technology, Clear Water Bay, Kowloon, Hong Kong (SAR), People’s Republic of China; bDepartment of Clinical Oncology, University of Hong Kong, Pokfulam, Hong Kong (SAR), People’s Republic of China; cDepartment of Biology, City of Hope, Beckman Research Institute, Duarte,CA 91010; dDepartment of Microbiology and Molecular Genetics, University of California, Irvine, CA 92697; eDepartment of Microbiology, Tumor and Cell Biology, Karolinska Institute, Stockholm, Sweden; and Departments of fPathology and gAnatomy, University of Hong Kong, Pokfulam, Hong Kong (SAR), People’s Republic of China Edited by George Klein, Karolinska Institutet, Stockholm, Sweden, and approved July 7, 2009 (received for review January 7, 2009) Chromosome 14 allelic loss is common in nasopharyngeal carcinoma particular interest to us that chromosome 14 loss is associated with (NPC) and may reflect essential tumor suppressor gene loss in tumor- cancer metastasis in breast tumors (15) and with poor clinical prognosis igenesis. An intact chromosome 14 was transferred to an NPC cell line for other head and neck cancers (16). Thus, it is possible that a using a microcell-mediated chromosome transfer approach. Microcell chromosome 14 TSG may be a useful prognostic marker in NPC. hybrids (MCHs) containing intact exogenously transferred chromo- In this study, we obtained functional evidence showing definitively some 14 were tumor suppressive in athymic mice, demonstrating that that chromosome 14 is tumor suppressive in NPC. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

(P -Value<0.05, Fold Change≥1.4), 4 Vs. 0 Gy Irradiation
Table S1: Significant differentially expressed genes (P -Value<0.05, Fold Change≥1.4), 4 vs. 0 Gy irradiation Genbank Fold Change P -Value Gene Symbol Description Accession Q9F8M7_CARHY (Q9F8M7) DTDP-glucose 4,6-dehydratase (Fragment), partial (9%) 6.70 0.017399678 THC2699065 [THC2719287] 5.53 0.003379195 BC013657 BC013657 Homo sapiens cDNA clone IMAGE:4152983, partial cds. [BC013657] 5.10 0.024641735 THC2750781 Ciliary dynein heavy chain 5 (Axonemal beta dynein heavy chain 5) (HL1). 4.07 0.04353262 DNAH5 [Source:Uniprot/SWISSPROT;Acc:Q8TE73] [ENST00000382416] 3.81 0.002855909 NM_145263 SPATA18 Homo sapiens spermatogenesis associated 18 homolog (rat) (SPATA18), mRNA [NM_145263] AA418814 zw01a02.s1 Soares_NhHMPu_S1 Homo sapiens cDNA clone IMAGE:767978 3', 3.69 0.03203913 AA418814 AA418814 mRNA sequence [AA418814] AL356953 leucine-rich repeat-containing G protein-coupled receptor 6 {Homo sapiens} (exp=0; 3.63 0.0277936 THC2705989 wgp=1; cg=0), partial (4%) [THC2752981] AA484677 ne64a07.s1 NCI_CGAP_Alv1 Homo sapiens cDNA clone IMAGE:909012, mRNA 3.63 0.027098073 AA484677 AA484677 sequence [AA484677] oe06h09.s1 NCI_CGAP_Ov2 Homo sapiens cDNA clone IMAGE:1385153, mRNA sequence 3.48 0.04468495 AA837799 AA837799 [AA837799] Homo sapiens hypothetical protein LOC340109, mRNA (cDNA clone IMAGE:5578073), partial 3.27 0.031178378 BC039509 LOC643401 cds. [BC039509] Homo sapiens Fas (TNF receptor superfamily, member 6) (FAS), transcript variant 1, mRNA 3.24 0.022156298 NM_000043 FAS [NM_000043] 3.20 0.021043295 A_32_P125056 BF803942 CM2-CI0135-021100-477-g08 CI0135 Homo sapiens cDNA, mRNA sequence 3.04 0.043389246 BF803942 BF803942 [BF803942] 3.03 0.002430239 NM_015920 RPS27L Homo sapiens ribosomal protein S27-like (RPS27L), mRNA [NM_015920] Homo sapiens tumor necrosis factor receptor superfamily, member 10c, decoy without an 2.98 0.021202829 NM_003841 TNFRSF10C intracellular domain (TNFRSF10C), mRNA [NM_003841] 2.97 0.03243901 AB002384 C6orf32 Homo sapiens mRNA for KIAA0386 gene, partial cds. -

A Dissertation Entitled the Androgen Receptor
A Dissertation entitled The Androgen Receptor as a Transcriptional Co-activator: Implications in the Growth and Progression of Prostate Cancer By Mesfin Gonit Submitted to the Graduate Faculty as partial fulfillment of the requirements for the PhD Degree in Biomedical science Dr. Manohar Ratnam, Committee Chair Dr. Lirim Shemshedini, Committee Member Dr. Robert Trumbly, Committee Member Dr. Edwin Sanchez, Committee Member Dr. Beata Lecka -Czernik, Committee Member Dr. Patricia R. Komuniecki, Dean College of Graduate Studies The University of Toledo August 2011 Copyright 2011, Mesfin Gonit This document is copyrighted material. Under copyright law, no parts of this document may be reproduced without the expressed permission of the author. An Abstract of The Androgen Receptor as a Transcriptional Co-activator: Implications in the Growth and Progression of Prostate Cancer By Mesfin Gonit As partial fulfillment of the requirements for the PhD Degree in Biomedical science The University of Toledo August 2011 Prostate cancer depends on the androgen receptor (AR) for growth and survival even in the absence of androgen. In the classical models of gene activation by AR, ligand activated AR signals through binding to the androgen response elements (AREs) in the target gene promoter/enhancer. In the present study the role of AREs in the androgen- independent transcriptional signaling was investigated using LP50 cells, derived from parental LNCaP cells through extended passage in vitro. LP50 cells reflected the signature gene overexpression profile of advanced clinical prostate tumors. The growth of LP50 cells was profoundly dependent on nuclear localized AR but was independent of androgen. Nevertheless, in these cells AR was unable to bind to AREs in the absence of androgen. -

A Genetic Screen Implicates a CWC16/Yju2/CCDC130 Protein and SMU1 in Alternative Splicing in Arabidopsis Thaliana
Downloaded from rnajournal.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press A genetic screen implicates a CWC16/Yju2/CCDC130 protein and SMU1 in alternative splicing in Arabidopsis thaliana TATSUO KANNO, WEN-DAR LIN, JASON L. FU, ANTONIUS J.M. MATZKE, and MARJORI MATZKE Institute of Plant and Microbial Biology, Academia Sinica, Taipei 115, Taiwan ABSTRACT To identify regulators of pre-mRNA splicing in plants, we developed a forward genetic screen based on an alternatively spliced GFP reporter gene in Arabidopsis thaliana. In wild-type plants, three major splice variants issue from the GFP gene but only one represents a translatable GFP mRNA. Compared to wild-type seedlings, which exhibit an intermediate level of GFP expression, mutants identified in the screen feature either a “GFP-weak” or “Hyper-GFP” phenotype depending on the ratio of the three splice variants. GFP-weak mutants, including previously identified prp8 and rtf2, contain a higher proportion of unspliced transcript or canonically spliced transcript, neither of which is translatable into GFP protein. In contrast, the coilin- deficient hyper-gfp1 (hgf1) mutant displays a higher proportion of translatable GFP mRNA, which arises from enhanced splicing of a U2-type intron with noncanonical AT–AC splice sites. Here we report three new hgf mutants that are defective, respectively, in spliceosome-associated proteins SMU1, SmF, and CWC16, an Yju2/CCDC130-related protein that has not yet been described in plants. The smu1 and cwc16 mutants have substantially increased levels of translatable GFP transcript owing to preferential splicing of the U2-type AT–AC intron, suggesting that SMU1 and CWC16 influence splice site selection in GFP pre-mRNA. -

Comparing Spatial Expression Dynamics of Bovine
Comparing spatial expression dynamics of bovine blastocyst under three different procedures : in-vivo, in-vitro derived, Title and somatic cell nuclear transfer embryos Nagatomo, Hiroaki; Akizawa, Hiroki; Sada, Ayari; Kishi, Yasunori; Yamanaka, Ken-ichi; Takuma, Tetsuya; Sasaki, Author(s) Keisuke; Yamauchi, Nobuhiko; Yanagawa, Yojiro; Nagano, Masashi; Kono, Tomohiro; Takahashi, Masashi; Kawahara, Manabu Citation Japanese Journal of Veterinary Research, 63(4), 159-171 Issue Date 2015-11 DOI 10.14943/jjvr.63.4.159 Doc URL http://hdl.handle.net/2115/60304 Type bulletin (article) Additional Information There are other files related to this item in HUSCAP. Check the above URL. File Information JJVR63-4 p.159-171 Suppl. Table 3.pdf (Supplemental Table 3) Instructions for use Hokkaido University Collection of Scholarly and Academic Papers : HUSCAP Supplemental table 3. Genes that were differentially expressed in the ICM relative to the TE in in SCNT blastocyst (SCNT list). Gene Symbol Probe Set ID Regulation Fold change ([ICM] / [TE]) Gene Title EEF1A1 AFFX-Bt-ef1a-3_at UP 1.2092365 eukaryotic translation elongation factor 1 alpha 1 IGFBP3 Bt.422.1.S2_at UP 2.5323892 insulin-like growth factor binding protein 3 IGFBP3 Bt.422.1.S1_at UP 3.7850845 insulin-like growth factor binding protein 3 SULT1A1 Bt.3537.1.S1_at UP 2.7092714 sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 SPP1 Bt.2632.1.S1_at UP 5.6928325 secreted phosphoprotein 1 SCARB1 Bt.4520.1.S1_at UP 3.106944 scavenger receptor class B, member 1 TSPO Bt.3988.1.S1_at -
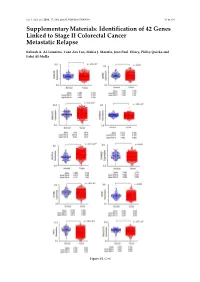
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

A Method for Chromatin Interactome Analysis of Solid Cancer
bioRxiv preprint doi: https://doi.org/10.1101/2021.01.11.426176; this version posted January 11, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Biop‐C: A Method for Chromatin Interactome Analysis of Solid Cancer Needle Biopsy Samples Sambhavi Animesh 1*, Ruchi Choudhary 1, 2*, Xin Yi Ng 3, Joshua Kai Xun Tay 4, Wan‐Qin Chong 3, Boon Cher Goh 1, 3, 5, Melissa Jane Fullwood 1, 2^ 1 Cancer Science Institute of Singapore, Centre for Translational Medicine, National University of Singapore, Singapore 2School of Biological Sciences, Nanyang Technological University, Singapore 3Department of Haematology‐Oncology, National University Cancer Institute of Singapore, National University Health System, Singapore 4Department of Otolaryngology ‐ Head & Neck Surgery, National University Hospital, Singapore 5Department of Pharmacology, Yong Loo Lin School of Medicine, National University Health System, Singapore, Singapore *Equal contribution (Co‐first authors) ^Corresponding author Abstract A major challenge in understanding the 3D genome organization of cancer samples is the lack of a method adapted to solid cancer needle biopsy samples. Here we developed Biop‐C, a modified in situ Hi‐C method, and applied it to characterize three nasopharyngeal cancer patient samples. We identified Topologically‐Associated Domains (TADs), chromatin interaction loops, and Frequently Interacting regions (FIREs) at key oncogenes in nasopharyngeal cancer from Biop‐C heat maps. Our results demonstrate the utility of our Biop‐C method in investigating the 3D genome organization in solid cancers, and the importance of 3D genome organization in regulating oncogenes in nasopharyngeal cancer. -

Facial Genetics: a Brief Overview
fgene-09-00462 October 16, 2018 Time: 12:3 # 1 REVIEW published: 16 October 2018 doi: 10.3389/fgene.2018.00462 Facial Genetics: A Brief Overview Stephen Richmond1*, Laurence J. Howe2,3, Sarah Lewis2,4, Evie Stergiakouli2,4 and Alexei Zhurov1 1 Applied Clinical Research and Public Health, School of Dentistry, College of Biomedical and Life Sciences, Cardiff University, Cardiff, United Kingdom, 2 MRC Integrative Epidemiology Unit, Population Health Sciences, University of Bristol, Bristol, United Kingdom, 3 Institute of Cardiovascular Science, University College London, London, United Kingdom, 4 School of Oral and Dental Sciences, University of Bristol, Bristol, United Kingdom Historically, craniofacial genetic research has understandably focused on identifying the causes of craniofacial anomalies and it has only been within the last 10 years, that there has been a drive to detail the biological basis of normal-range facial variation. This initiative has been facilitated by the availability of low-cost hi-resolution three- dimensional systems which have the ability to capture the facial details of thousands of individuals quickly and accurately. Simultaneous advances in genotyping technology have enabled the exploration of genetic influences on facial phenotypes, both in the Edited by: present day and across human history. Peter Claes, KU Leuven, Belgium There are several important reasons for exploring the genetics of normal-range variation Reviewed by: in facial morphology. Hui-Qi Qu, Children’s Hospital of Philadelphia, - Disentangling