ABSTRACT Studies on Bovine Γ-Glutamylamine Cyclotransferase
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Journal of Chromatography
aphy & S r ep og a t r a a t m i o o r n Lilla et al., J Chromatograph Separat Techniq 2012, 3:2 h T e C c f Journal of Chromatography h DOI: 10.4172/2157-7064.1000122 o n l i a q ISSN:n 2157-7064 u r e u s o J Separation Techniques Research Article OpenOpen Access Access Structural Characterization of Transglutaminase-Catalyzed Casein Cross- Linking Sergio Lilla1,2, Gianfranco Mamone2, Maria Adalgisa Nicolai1, Lina Chianese1, Gianluca Picariello2, Simonetta Caira2 and Francesco Addeo1,2* 1Dipartimento di Scienza degli Alimenti, University of Naples “Federico II”, Parco Gussone, Portici 80055, Italy 2Istituto di Scienze dell’Alimentazione (ISA) – CNR, Via Roma 64, 83100 Avellino, Italy Abstract Microbial transglutaminase is used in the food industry to improve texture by catalyzing protein cross-linking. Casein is a well-known transglutaminase substrate, but the complete role of glutamine (Q) and lysine (K) residues in its cross-linking is not fully understood. In this study, we describe the characterization of microbial Transglutaminase -modified casein using a combination of immunological and proteomic techniques. Using 5-(biotinamido)pentylamine as an acyl acceptor probe, three Q residues of β-casein and one of αs1-casein were found to participate as acyl donors. However, no Q-residues were involved in network formation with κ-casein or αs2-casein. Q and K residues in the ε-(γ-glutamyl)lysine-isopeptide bonds β-casein were identified by nanoelectrospray tandem mass spectrometry of the proteolytic digests. This work reports our progress toward a better understanding of the function and mechanism of action of microbial transglutaminase-mediated proteins. -

Amidoligases with ATP-Grasp, Glutamine Synthetase-Like and Acetyltransferase-Like Domains: Synthesis of Novel Metabolites and Peptide Modifications of Proteinswz
View Article Online / Journal Homepage / Table of Contents for this issue Molecular BioSystems This article was published as part of the Computational and Systems Biology themed issue Please take a look at the full table of contents to access the other papers in this issue. Open Access Article. Published on 13 October 2009. Downloaded 9/27/2021 9:23:51 AM. View Article Online PAPER www.rsc.org/molecularbiosystems | Molecular BioSystems Amidoligases with ATP-grasp, glutamine synthetase-like and acetyltransferase-like domains: synthesis of novel metabolites and peptide modifications of proteinswz Lakshminarayan M. Iyer,a Saraswathi Abhiman,a A. Maxwell Burroughsb and L. Aravind*a Received 28th August 2009, Accepted 28th August 2009 First published as an Advance Article on the web 13th October 2009 DOI: 10.1039/b917682a Recent studies have shown that the ubiquitin system had its origins in ancient cofactor/amino acid biosynthesis pathways. Preliminary studies also indicated that conjugation systems for other peptide tags on proteins, such as pupylation, have evolutionary links to cofactor/amino acid biosynthesis pathways. Following up on these observations, we systematically investigated the non-ribosomal amidoligases of the ATP-grasp, glutamine synthetase-like and acetyltransferase folds by classifying the known members and identifying novel versions. We then established their contextual connections using information from domain architectures and conserved gene neighborhoods. This showed remarkable, previously uncharacterized functional links between diverse peptide ligases, several peptidases of unrelated folds and enzymes involved in synthesis of modified amino acids. Using the network of contextual connections we were able to predict numerous novel pathways for peptide synthesis and modification, amine-utilization, secondary metabolite synthesis and potential peptide-tagging systems. -

Computational and Systems Biology Themed Issue
View Article Online / Journal Homepage / Table of Contents for this issue Molecular BioSystems This article was published as part of the Computational and Systems Biology themed issue Please take a look at the full table of contents to access the other papers in this issue. Open Access Article. Published on 13 October 2009. Downloaded 9/23/2021 6:41:00 PM. View Article Online PAPER www.rsc.org/molecularbiosystems | Molecular BioSystems Amidoligases with ATP-grasp, glutamine synthetase-like and acetyltransferase-like domains: synthesis of novel metabolites and peptide modifications of proteinswz Lakshminarayan M. Iyer,a Saraswathi Abhiman,a A. Maxwell Burroughsb and L. Aravind*a Received 28th August 2009, Accepted 28th August 2009 First published as an Advance Article on the web 13th October 2009 DOI: 10.1039/b917682a Recent studies have shown that the ubiquitin system had its origins in ancient cofactor/amino acid biosynthesis pathways. Preliminary studies also indicated that conjugation systems for other peptide tags on proteins, such as pupylation, have evolutionary links to cofactor/amino acid biosynthesis pathways. Following up on these observations, we systematically investigated the non-ribosomal amidoligases of the ATP-grasp, glutamine synthetase-like and acetyltransferase folds by classifying the known members and identifying novel versions. We then established their contextual connections using information from domain architectures and conserved gene neighborhoods. This showed remarkable, previously uncharacterized functional links between diverse peptide ligases, several peptidases of unrelated folds and enzymes involved in synthesis of modified amino acids. Using the network of contextual connections we were able to predict numerous novel pathways for peptide synthesis and modification, amine-utilization, secondary metabolite synthesis and potential peptide-tagging systems. -

Role of Transglutaminase 2 in Cell Death, Survival, and Fibrosis
cells Review Role of Transglutaminase 2 in Cell Death, Survival, and Fibrosis Hideki Tatsukawa * and Kiyotaka Hitomi Cellular Biochemistry Laboratory, Graduate School of Pharmaceutical Sciences, Nagoya University, Tokai National Higher Education and Research System, Nagoya 464-8601, Aichi, Japan; [email protected] * Correspondence: [email protected]; Tel.: +81-52-747-6808 Abstract: Transglutaminase 2 (TG2) is a ubiquitously expressed enzyme catalyzing the crosslink- ing between Gln and Lys residues and involved in various pathophysiological events. Besides this crosslinking activity, TG2 functions as a deamidase, GTPase, isopeptidase, adapter/scaffold, protein disulfide isomerase, and kinase. It also plays a role in the regulation of hypusination and serotonylation. Through these activities, TG2 is involved in cell growth, differentiation, cell death, inflammation, tissue repair, and fibrosis. Depending on the cell type and stimulus, TG2 changes its subcellular localization and biological activity, leading to cell death or survival. In normal unstressed cells, intracellular TG2 exhibits a GTP-bound closed conformation, exerting prosurvival functions. However, upon cell stimulation with Ca2+ or other factors, TG2 adopts a Ca2+-bound open confor- mation, demonstrating a transamidase activity involved in cell death or survival. These functional discrepancies of TG2 open form might be caused by its multifunctional nature, the existence of splicing variants, the cell type and stimulus, and the genetic backgrounds and variations of the mouse models used. TG2 is also involved in the phagocytosis of dead cells by macrophages and in fibrosis during tissue repair. Here, we summarize and discuss the multifunctional and controversial Citation: Tatsukawa, H.; Hitomi, K. roles of TG2, focusing on cell death/survival and fibrosis. -

1114 Tissue Transglutaminase (TG2) and Mitochondrial Function And
[Frontiers In Bioscience, Landmark, 22, 1114-1137, March 1, 2017] Tissue transglutaminase (TG2) and mitochondrial function and dysfunction Thung-S. Lai 1, Cheng-Jui Lin 2,3, Yu-Ting Wu4, Chih-Jen Wu2,5,6 1Institute of Biomedical Science, Mackay Medical College, New Taipei City, Taiwan, ROC, 2Nephrology/ Department of Internal Medicine, Mackay Memorial Hospital, Taipei, Taiwan, ROC, 3Nursing and Management, Mackay Junior College of Medicine, Taipei, Taiwan, ROC, 4Institute of Biochemistry and Molecular Biology, National Yang-Ming University, Taipei, Taiwan, 5Department of Medicine, Mackay Medical College, New Taipei City, Taiwan, ROC, 6Graduate Institute of Medical Science, Taipei Medical University, Taipei, Taiwan, ROC TABLE OF CONTENTS 1. Abstract 2. Introduction 3. TG2: a multifunctional enzyme. 3.1. Transamidation Reaction (TGase function) 3.1.1. Inter- or intra-molecular crosslinking 3.1.2. Aminylation 3.1.3. Deamidation 3.2. Isopeptidase activity 3.3. Protein Disulfide Isomerase (PDI) activity 3.4. GTP/ATP hydrolysis activity 4. Structure and function of TG2 4.1. TGase active site 4.2. GTP and ATP binding site 5. Regulation of in vivo TGase activity by GTP, redox, and nitric oxide (NO) 5.1. Regulation of in vivo TGase activity by GTP 5.2. Regulation of in vivo TGase activity by redox 5.3. Regulation of in vivo TGase activity by NO 6. Regulation of TG2 expression 6.1. NFkB regulates the expression of TG2 6.2. Hypoxia regulates the expression of TG2 6.3. TGFb regulates the expression of TG2 6.4. Oxidative stress and EGF up-regulate the expression of TG2. 7. TG2 is localized in mitochondria and several other locations 8. -
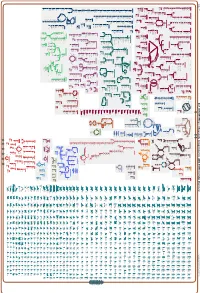
Generate Metabolic Map Poster
Authors: Pallavi Subhraveti Ron Caspi Peter Midford Peter D Karp An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Ingrid Keseler Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. Gcf_003855395Cyc: Shewanella livingstonensis LMG 19866 Cellular Overview Connections between pathways are omitted for legibility. -

Mitochondria Targeting As an Effective Strategy for Cancer Therapy
International Journal of Molecular Sciences Review Mitochondria Targeting as an Effective Strategy for Cancer Therapy Poorva Ghosh , Chantal Vidal, Sanchareeka Dey and Li Zhang * Department of Biological Sciences, the University of Texas at Dallas, Richardson, TX 75080, USA; [email protected] (P.G.); [email protected] (C.V.); [email protected] (S.D.) * Correspondence: [email protected]; Tel.: +972-883-5757 Received: 25 February 2020; Accepted: 6 May 2020; Published: 9 May 2020 Abstract: Mitochondria are well known for their role in ATP production and biosynthesis of macromolecules. Importantly, increasing experimental evidence points to the roles of mitochondrial bioenergetics, dynamics, and signaling in tumorigenesis. Recent studies have shown that many types of cancer cells, including metastatic tumor cells, therapy-resistant tumor cells, and cancer stem cells, are reliant on mitochondrial respiration, and upregulate oxidative phosphorylation (OXPHOS) activity to fuel tumorigenesis. Mitochondrial metabolism is crucial for tumor proliferation, tumor survival, and metastasis. Mitochondrial OXPHOS dependency of cancer has been shown to underlie the development of resistance to chemotherapy and radiotherapy. Furthermore, recent studies have demonstrated that elevated heme synthesis and uptake leads to intensified mitochondrial respiration and ATP generation, thereby promoting tumorigenic functions in non-small cell lung cancer (NSCLC) cells. Also, lowering heme uptake/synthesis inhibits mitochondrial OXPHOS and effectively reduces oxygen consumption, thereby inhibiting cancer cell proliferation, migration, and tumor growth in NSCLC. Besides metabolic changes, mitochondrial dynamics such as fission and fusion are also altered in cancer cells. These alterations render mitochondria a vulnerable target for cancer therapy. This review summarizes recent advances in the understanding of mitochondrial alterations in cancer cells that contribute to tumorigenesis and the development of drug resistance. -

Associated Gene Expression Profiles in Multiple Sclerosis Jeroen Melief1, Marie Orre2, Koen Bossers3, Corbert G
Melief et al. Acta Neuropathologica Communications (2019) 7:60 https://doi.org/10.1186/s40478-019-0705-7 RESEARCH Open Access Transcriptome analysis of normal-appearing white matter reveals cortisol- and disease- associated gene expression profiles in multiple sclerosis Jeroen Melief1, Marie Orre2, Koen Bossers3, Corbert G. van Eden1, Karianne G. Schuurman1, Matthew R. J. Mason1,3, Joost Verhaagen3, Jörg Hamann1,4 and Inge Huitinga1* Abstract Inter-individual differences in cortisol production by the hypothalamus–pituitary–adrenal (HPA) axis are thought to contribute to clinical and pathological heterogeneity of multiple sclerosis (MS). At the same time, accumulating evidence indicates that MS pathogenesis may originate in the normal-appearing white matter (NAWM). Therefore, we performed a genome-wide transcriptional analysis, by Agilent microarray, of post-mortem NAWM of 9 control subjects and 18 MS patients to investigate to what extent gene expression reflects disease heterogeneity and HPA- axis activity. Activity of the HPA axis was determined by cortisol levels in cerebrospinal fluid and by numbers of corticotropin-releasing neurons in the hypothalamus, while duration of MS and time to EDSS6 served as indicator of disease severity. Applying weighted gene co-expression network analysis led to the identification of a range of gene modules with highly similar co-expression patterns that strongly correlated with various indicators of HPA-axis activity and/or severity of MS. Interestingly, molecular profiles associated with relatively -

Celiac Disease in the Pediatric Patient
Laboratory Services Celiac Disease in the Pediatric Patient What is Celiac Disease? Celiac disease is a common chronic condition that affects about 1% of the general population. It occurs in genetically predisposed individuals and is triggered by the ingestion of products that contain wheat, barley or rye, collectively known as “gluten”. Ingestion of gluten initiates an inflammatory cascade in the intestines that causes progressive destruction of the small intestinal villi. Symptoms of Celiac Disease Symptoms of celiac disease are highly variable and may be gastrointestinal or non-gastrointestinal as shown in the table. • Symptoms may occur singly or in combination • Age of onset can be any time from infancy to late in adulthood • Young children tend to have predominantly gastrointestinal symptoms while older children and adults are more likely to have non-gastrointestinal manifestations initially Because of the variable manifestations, a high index of suspicion for the disease and liberal use of screening tests is needed to avoid unnecessary delays in diagnosis. Associated Conditions. Some people are at increased risk for celiac disease because of an associated condition. These individuals may have no symptoms or very minor complaints but when investigated are found to have the intestinal damage. Groups at increased risk for celiac disease include: • First degree relatives of an index case • Turner syndrome • Type 1 diabetics • Selective IgA deficiency • Down Syndrome • Other autoimmune diseases (autoimmune hepatitis, thyroiditis) Diagnosing Celiac Disease Blood tests looking for specific antibodies that are found when the disease is active can be used to screen for celiac disease. Recommended tests include: • Tissue transglutaminase antibody (TTG) • Endomysium antibody (EMA) • Deamidated gliadin peptide antibodies (DGP) The TTG-IgA antibody provides the most cost effective and reliable test to screen for celiac disease. -

The Human Gamma-Glutamyltransferase Gene Family
Hum Genet (2008) 123:321–332 DOI 10.1007/s00439-008-0487-7 REVIEW The human gamma-glutamyltransferase gene family Nora Heisterkamp · John GroVen · David Warburton · Tam P. Sneddon Received: 9 November 2007 / Accepted: 6 March 2008 / Published online: 21 March 2008 © Springer-Verlag 2008 Abstract Assays for gamma-glutamyl transferase related genes or sequences. These sequences were given (GGT1, EC 2.3.2.2) activity in blood are widely used in a multiple diVerent names, leading to inconsistencies and clinical setting to measure tissue damage. The well-charac- confusion. Here we systematically evaluated all human terized GGT1 is an extracellular enzyme that is anchored to sequences related to GGT1 using genomic and cDNA data- the plasma membrane of cells. There, it hydrolyzes and base searches and identiWed thirteen genes belonging to the transfers -glutamyl moieties from glutathione and other extended GGT family, of which at least six appear to be -glutamyl compounds to acceptors. As such, it has a critical active. In collaboration with the HUGO Gene Nomencla- function in the metabolism of glutathione and in the con- ture Committee (HGNC) we have designated possible version of the leukotriene LTC4 to LTD4. GGT deWciency active genes with nucleotide or amino acid sequence simi- in man is rare and for the few patients reported to date, larity to GGT1, as GGT5 (formerly GGL, GGTLA1/GGT- mutations in GGT1 have not been described. These patients rel), GGT6 (formerly rat ggt6 homologue) and GGT7 (for- do secrete glutathione in urine and fail to metabolize LTC4. merly GGTL3, GGT4). Two loci have the potential to Earlier pre-genome investigations had indicated that encode only the light chain portion of GGT and have now besides GGT1, the human genome contains additional been designated GGTLC1 (formerly GGTL6, GGTLA4) and GGTLC2. -

The Uniqueness of Tryptophan in Biology: Properties, Metabolism, Interactions and Localization in Proteins
International Journal of Molecular Sciences Review The Uniqueness of Tryptophan in Biology: Properties, Metabolism, Interactions and Localization in Proteins Sailen Barik 3780 Pelham Drive, Mobile, AL 36619, USA; [email protected] Received: 2 November 2020; Accepted: 17 November 2020; Published: 20 November 2020 Abstract: Tryptophan (Trp) holds a unique place in biology for a multitude of reasons. It is the largest of all twenty amino acids in the translational toolbox. Its side chain is indole, which is aromatic with a binuclear ring structure, whereas those of Phe, Tyr, and His are single-ring aromatics. In part due to these elaborate structural features, the biosynthetic pathway of Trp is the most complex and the most energy-consuming among all amino acids. Essential in the animal diet, Trp is also the least abundant amino acid in the cell, and one of the rarest in the proteome. In most eukaryotes, Trp is the only amino acid besides Met, which is coded for by a single codon, namely UGG. Due to the large and hydrophobic π-electron surface area, its aromatic side chain interacts with multiple other side chains in the protein, befitting its strategic locations in the protein structure. Finally, several Trp derivatives, namely tryptophylquinone, oxitriptan, serotonin, melatonin, and tryptophol, have specialized functions. Overall, Trp is a scarce and precious amino acid in the cell, such that nature uses it parsimoniously, for multiple but selective functions. Here, the various aspects of the uniqueness of Trp are presented in molecular terms. Keywords: tryptophan; indole; virus; immunity; serotonin; kynurenine; codon 1. Introduction Tryptophan (Trp, W) is one of three aromatic amino acids that minimally contain a six-membered benzene ring in their side chains, the other two being phenylalanine (Phe, F) and tyrosine (Tyr, Y). -

Setd1 Histone 3 Lysine 4 Methyltransferase Complex Components in Epigenetic Regulation
SETD1 HISTONE 3 LYSINE 4 METHYLTRANSFERASE COMPLEX COMPONENTS IN EPIGENETIC REGULATION Patricia A. Pick-Franke Submitted to the faculty of the University Graduate School in partial fulfillment of the requirements for the degree Master of Science in the Department of Biochemistry and Molecular Biology Indiana University December 2010 Accepted by the Faculty of Indiana University, in partial fulfillment of the requirements for the degree of Master of Science. _____________________________________ David Skalnik, Ph.D., Chair _____________________________________ Kristin Chun, Ph.D. Master’s Thesis Committee _____________________________________ Simon Rhodes, Ph.D. ii DEDICATION This thesis is dedicated to my sons, Zachary and Zephaniah who give me great joy, hope and continuous inspiration. I can only hope that I successfully set a good example demonstrating that one can truly accomplish anything, if you never give up and reach for your dreams. iii ACKNOWLEDGEMENTS I would like to thank my committee members Dr. Skalnik, Dr. Chun and Dr. Rhodes for allowing me to complete this dissertation. They have been incredibly generous with their flexibility. I must make a special thank you to Jeanette McClintock, who willingly gave her expertise in statistical analysis with the Cfp1 microarray data along with encouragement, support and guidance to complete this work. I would like to thank Courtney Tate for her ceaseless willingness to share ideas, and her methods and materials, and Erika Dolbrota for her generous instruction as well as the name of a good doctor. I would also like to acknowledge the superb mentorship of Dr. Jeon Heong Lee, PhD and the contagious passion and excitement for the life of science of Dr.