Topic:1. Endosymbiotic Theory History- Konstantin Mereschkowski's 1905 Tree-Of-Life Diagram, Showing the Origin of Complex Life
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Pinpointing the Origin of Mitochondria Zhang Wang Hanchuan, Hubei
Pinpointing the origin of mitochondria Zhang Wang Hanchuan, Hubei, China B.S., Wuhan University, 2009 A Dissertation presented to the Graduate Faculty of the University of Virginia in Candidacy for the Degree of Doctor of Philosophy Department of Biology University of Virginia August, 2014 ii Abstract The explosive growth of genomic data presents both opportunities and challenges for the study of evolutionary biology, ecology and diversity. Genome-scale phylogenetic analysis (known as phylogenomics) has demonstrated its power in resolving the evolutionary tree of life and deciphering various fascinating questions regarding the origin and evolution of earth’s contemporary organisms. One of the most fundamental events in the earth’s history of life regards the origin of mitochondria. Overwhelming evidence supports the endosymbiotic theory that mitochondria originated once from a free-living α-proteobacterium that was engulfed by its host probably 2 billion years ago. However, its exact position in the tree of life remains highly debated. In particular, systematic errors including sparse taxonomic sampling, high evolutionary rate and sequence composition bias have long plagued the mitochondrial phylogenetics. This dissertation employs an integrated phylogenomic approach toward pinpointing the origin of mitochondria. By strategically sequencing 18 phylogenetically novel α-proteobacterial genomes, using a set of “well-behaved” phylogenetic markers with lower evolutionary rates and less composition bias, and applying more realistic phylogenetic models that better account for the systematic errors, the presented phylogenomic study for the first time placed the mitochondria unequivocally within the Rickettsiales order of α- proteobacteria, as a sister clade to the Rickettsiaceae and Anaplasmataceae families, all subtended by the Holosporaceae family. -

Acanthophora Dendroides Harvey (Rhodomelaceae), a New Record for the Atlantic and Pacific Oceans
15 3 NOTES ON GEOGRAPHIC DISTRIBUTION Check List 15 (3): 509–514 https://doi.org/10.15560/15.3.509 Acanthophora dendroides Harvey (Rhodomelaceae), a new record for the Atlantic and Pacific oceans Gabriela C. García-Soto, Juan M. Lopez-Bautista The University of Alabama, Department of Biological Sciences, Science and Engineering Complex, 1325 Hackberry Ln, Tuscaloosa, AL 35401, USA. Corresponding author: Gabriela García-Soto, [email protected] Abstract We record Acanthophora dendroides Harvey for the first time in the Atlantic Ocean. Two specimens from the Philip- pines were resolved as conspecific to the Atlantic A. dendroides in molecular analyses extending its geographic range to the Philippines. In light of new evidence provided by field-collected specimens ofAcanthophora spicifera (M.Vahl) Børgesen (generitype) from Florida and Venezuela, the flattened species A. pacifica(Setchell) Kraft, showed no affin- ity to Acanthophora sensu stricto, suggesting that the genus should be restricted to cylindrical species only. Key words Atlantic Ocean, Philippines, taxonomy. Academic editor: Luciane Fontana da Silva | Received 8 October 2018 | Accepted 21 January 2019 | Published 21 June 2019 Citation: García-Soto GC, Lopez-Bautista JM (2019) Acanthophora dendroides Harvey (Rhodomelaceae), a new record for the Atlantic and Pacific oceans. Check List 15 (3): 509–514. https://doi.org/10.15560/15.3.509 Introduction Kraft are restricted to the Pacific Ocean (Guiry and Guiry 2018), A. dendroides Harvey to the Indian Ocean The genus Acanthophora J.V. Lamouroux 1813 is a (Silva et al. 1996) and A. ramulosa Lindenb. ex. Kutz- member of the tribe Chondrieae and it is distinguished from other genera of the tribe by the presence of spirally ing appears to be confined to the Gulf of Guinea in West arranged acute spines (Gordon-Mills and Womersley Africa (Steentoft 1967). -

Origin and Evolution of Eukaryotic Compartmentalization
TESI DOCTORAL UPF 2016 Origin and evolution of eukaryotic compartmentalization Alexandros Pittis Director Dr. Toni Gabaldon´ Comparative Genomics Group Bioinformatics and Genomics Department Centre for Genomic Regulation (CRG) To my father Stavros for the PhD he never did Acknowledgments Looking back at the years of my PhD in the Comparative Genomics group at CRG, I feel it was equally a process of scientific, as well as personal development. All this time I was very privileged to be surrounded by people that offered me their generous support in both fronts. And they were available in the very moments that I was fighting more myself than the unsolvable (anyway) comparative genomics puzzles. I hope that in the future I will have many times the chance to express them my appreciation, way beyond these few words. First, to Toni, my supervisor, for all his support, and patience, and confidence, and respect, especially at the moments that things did not seem that promising. He offered me freedom, to try, to think, to fail, to learn, to achieve that few enjoy during their PhD, and I am very thankful to him. Then, to my good friends and colleagues, those that I found in the group already, and others that joined after me. A very special thanks to Marinita and Jaime, for all their valuable time, and guidance and paradigm, which nevertheless I never managed to follow. They have both marked my phylogenomics path so far and I cannot escape. To Les and Salvi, my fellow students and pals at the time for all that we shared; to Gab and Fran and Dam, for -

Ribosomal Dna Phylogeny of the Bangiophycidae (Rhodophyta) and the Origin of Secondary Plastids1
American Journal of Botany 88(8): 1390±1400. 2001. RIBOSOMAL DNA PHYLOGENY OF THE BANGIOPHYCIDAE (RHODOPHYTA) AND THE ORIGIN OF SECONDARY PLASTIDS1 KIRSTEN M. MUÈ LLER,2,6 MARIANA C. OLIVEIRA,3 ROBERT G. SHEATH,2 AND DEBASHISH BHATTACHARYA4,5 2Department of Botany and Dean's Of®ce, University of Guelph, Guelph, Ontario, Canada, N1G 2W1; 3Department of Botany, Institute of Biosciences, University of SaÄo Paulo, R. MataÄo, Travessa 14, N. 321, SaÄo Paulo, SP, Brazil, CEP 05508-900; and 4Department of Biological Sciences, University of Iowa, 239 Biology Building, Iowa City, Iowa 52242-3110 USA We sequenced the nuclear small subunit ribosomal DNA coding region from 20 members of the Bangiophycidae and from two members of the Florideophycidae to gain insights into red algal evolution. A combined alignment of nuclear and plastid small subunit rDNA and a data set of Rubisco protein sequences were also studied to complement the understanding of bangiophyte phylogeny and to address red algal secondary symbiosis. Our results are consistent with a monophyletic origin of the Florideophycidae, which form a sister-group to the Bangiales. Bangiales monophyly is strongly supported, although Porphyra is polyphyletic within Bangia. Ban- giophycidae orders such as the Porphyridiales are distributed over three independent red algal lineages. The Compsopogonales sensu stricto, consisting of two freshwater families, Compsopogonaceae and Boldiaceae, forms a well-supported monophyletic grouping. The single taxon within the Rhodochaetales, Rhodochaete parvula, is positioned within a cluster containing members of the Erythropelti- dales. Analyses of Rubisco sequences show that the plastids of the heterokonts are most closely related to members of the Cyanidiales and are not directly related to cryptophyte and haptophyte plastid genomes. -

Refinements for the Amplification and Sequencing of Red Algal DNA Barcode and Redtol Phylogenetic Markers: a Summary of Current Primers, Profiles and Strategies
Review Algae 2013, 28(1): 31-43 http://dx.doi.org/10.4490/algae.2013.28.1.031 Open Access Refinements for the amplification and sequencing of red algal DNA barcode and RedToL phylogenetic markers: a summary of current primers, profiles and strategies Gary W. Saunders1,* and Tanya E. Moore1 1Centre for Environmental and Molecular Algal Research, Department of Biology, University of New Brunswick, Frederic- ton, NB E3B 5A3, Canada This review provides a comprehensive summary of the PCR primers and profiles currently in use in our laboratory for red algal DNA barcoding and phylogenetic research. While work focuses on florideophyte taxa, many of the markers have been applied successfully to the Bangiales, as well as other lineages previously assigned to the Bangiophyceae sensu lato. All of the primers currently in use with their respective amplification profiles and strategies are provided, which can include full fragment, overlapping fragments and what might best be called “informed overlapping fragments”, i.e., a fragment for a marker is amplified and sequenced for a taxon and those sequence data are then used to identify the best primers to amplify the remaining fragment(s) for that marker. We extend this strategy for the more variable markers with sequence from the external PCR primers used to “inform” the selection of internal sequencing primers. This summary will hopefully serve as a useful resource to systematists in the red algal community. Key Words: DNA barcode; Florideophyceae; molecular markers; phylogeny; polymerase chain reaction; primers; RedToL INTRODUCTION There can be little question that molecular techniques redtol/) (Vis et al. 2010). -

Halymeniaceae, Rhodophyta) in Bermuda, Western Atlantic Ocean, Including Five New Species and C
Trinity College Trinity College Digital Repository Faculty Scholarship 6-2018 A Revision of the Genus Cryptonemia (Halymeniaceae, Rhodophyta) in Bermuda, Western Atlantic Ocean, Including Five New Species and C. bermudensis (Collins & M. Howe) comb. nov [post-print] Craig W. Schneider Trinity College, [email protected] Christopher E. Lane University of Rhode Island Gary W. Saunders Follow this and additional works at: https://digitalrepository.trincoll.edu/facpub Part of the Biology Commons TEJP-2017-0083.R1 _____________________________________________________________________________ A revision of the genus Cryptonemia (Halymeniaceae, Rhodophyta) in Bermuda, western Atlantic Ocean, including five new species and C. bermudensis (Collins et M. Howe) comb. nov. Craig W. Schneidera, Christopher E. Laneb and Gary W. Saundersc aDepartment of Biology, Trinity College, Hartford, CT 06106, USA; bDepartment of Biological Sciences, University of Rhode Island, Kingston, RI 02881, USA; cCentre for Environmental & Molecular Algal Research, Department of Biology, University of New Brunswick, Fredericton, NB E3B 5A3, Canada Left Header: C. SCHNEIDER ET AL. _____________________________________________________________________________ CONTACT Craig W. Schneider. E-mail: [email protected] 2 ABSTRACT Cryptonemia specimens collected in Bermuda over the past two decades were analyzed using gene sequences encoding the large subunit of the nuclear ribosomal DNA and the large subunit of RuBisCO as genetic markers to elucidate their phylogenetic positions. They were additionally subjected to morphological assessment and compared with historical collections from the islands. Six species are presently found in the flora including C. bermudensis comb. nov., based on Halymenia bermudensis, and the following five new species: C. abyssalis, C. antricola, C. atrocostalis, C. lacunicola and C. perparva. Of the eight species known in the western Atlantic flora prior to this study, none is found in Bermuda. -

Red and Green Algal Monophyly and Extensive Gene
Please cite this article in press as: Chan et al., Red and Green Algal Monophyly and Extensive Gene Sharing Found in a Rich Reper- toire of Red Algal Genes, Current Biology (2011), doi:10.1016/j.cub.2011.01.037 Current Biology 21, 1–6, February 22, 2011 ª2011 Elsevier Ltd All rights reserved DOI 10.1016/j.cub.2011.01.037 Report Red and Green Algal Monophyly and Extensive Gene Sharing Found in a Rich Repertoire of Red Algal Genes Cheong Xin Chan,1,5 Eun Chan Yang,2,5 Titas Banerjee,1 sequences in our local database, in which we included the Hwan Su Yoon,2,* Patrick T. Martone,3 Jose´ M. Estevez,4 23,961 predicted proteins from C. tuberculosum (see Table and Debashish Bhattacharya1,* S1 available online). Of these hits, 9,822 proteins (72.1%, 1Department of Ecology, Evolution, and Natural Resources including many P. cruentum paralogs) were present in C. tuber- and Institute of Marine and Coastal Sciences, Rutgers culosum and/or other red algae, 6,392 (46.9%) were shared University, New Brunswick, NJ 08901, USA with C. merolae, and 1,609 were found only in red algae. A total 2Bigelow Laboratory for Ocean Sciences, West Boothbay of 1,409 proteins had hits only to red algae and one other Harbor, ME 04575, USA phylum. Using this repertoire, we adopted a simplified recip- 3Department of Botany, University of British Columbia, 6270 rocal BLAST best-hits approach to study the pattern of exclu- University Boulevard, Vancouver, BC V6T 1Z4, Canada sive gene sharing between red algae and other phyla (see 4Instituto de Fisiologı´a, Biologı´a Molecular y Neurociencias Experimental Procedures). -

Diversity and Evolution of Algae: Primary Endosymbiosis
CHAPTER TWO Diversity and Evolution of Algae: Primary Endosymbiosis Olivier De Clerck1, Kenny A. Bogaert, Frederik Leliaert Phycology Research Group, Biology Department, Ghent University, Krijgslaan 281 S8, 9000 Ghent, Belgium 1Corresponding author: E-mail: [email protected] Contents 1. Introduction 56 1.1. Early Evolution of Oxygenic Photosynthesis 56 1.2. Origin of Plastids: Primary Endosymbiosis 58 2. Red Algae 61 2.1. Red Algae Defined 61 2.2. Cyanidiophytes 63 2.3. Of Nori and Red Seaweed 64 3. Green Plants (Viridiplantae) 66 3.1. Green Plants Defined 66 3.2. Evolutionary History of Green Plants 67 3.3. Chlorophyta 68 3.4. Streptophyta and the Origin of Land Plants 72 4. Glaucophytes 74 5. Archaeplastida Genome Studies 75 Acknowledgements 76 References 76 Abstract Oxygenic photosynthesis, the chemical process whereby light energy powers the conversion of carbon dioxide into organic compounds and oxygen is released as a waste product, evolved in the anoxygenic ancestors of Cyanobacteria. Although there is still uncertainty about when precisely and how this came about, the gradual oxygenation of the Proterozoic oceans and atmosphere opened the path for aerobic organisms and ultimately eukaryotic cells to evolve. There is a general consensus that photosynthesis was acquired by eukaryotes through endosymbiosis, resulting in the enslavement of a cyanobacterium to become a plastid. Here, we give an update of the current understanding of the primary endosymbiotic event that gave rise to the Archaeplastida. In addition, we provide an overview of the diversity in the Rhodophyta, Glaucophyta and the Viridiplantae (excluding the Embryophyta) and highlight how genomic data are enabling us to understand the relationships and characteristics of algae emerging from this primary endosymbiotic event. -

Journal of Pharmacology and Toxicological Studies
e-ISSN: 2319-9873 p-ISSN: 2347-2324 Research & Reviews: Journal of Pharmacology and Toxicological Studies A Short Study on Phylogenetics Indu Sama* Department of plant Pathology and forest genetics, Bharat University, India Review Article ABSTRACT Received: 06/09/2016 Accepted : 09/09/2016 Published: 12/09/2016 In science, phylogenetics is the investigation of transformative *For Correspondence connections among gatherings of life forms (e.g. species,populations), which are found through atomic sequencing information and morphological information grids. The term phylogeneticsderives from the Greek Department of plant Pathology expressions phylé (φυλή) and phylon (φῦλον), meaning "tribe", "faction", and forest genetics, Bharat "race" and the descriptive structure, genetikós(γενετικός), of the word University, India genesis (γένεσις) "source", "source", "conception". Indeed, phylogenesis is the procedure, phylogeny is science on this procedure, and phylogenetics is E-mail: [email protected] phylogeny in view of examination of successions of organic macromolecules (DNA, RNA and proteins, in the first). The consequence of phylogenetic Keywords: Phylogenetics, studies is a speculation about the developmental history of taxonomic Devolopment, Computation gatherings: their phylogeny. INTRODUCTION Development is a procedure whereby populaces are modified over the long run and may part into particular branches, hybridize together, or end by elimination [1 -15]. The transformative expanding procedure may be portrayed as a phylogenetic tree, and the spot of each of the different living beings on the tree is in view of a speculation about the arrangement in which developmental spreading occasions happened. Inhistorical phonetics, comparative ideas are utilized regarding connections in the middle of dialects; and in literary feedback withstemmatics [15-25]. -
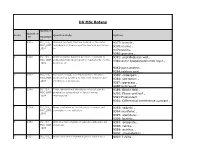
DU Msc Botany
DU MSc Botany Question Question Sr.No Question Body Options Id Descripti on 1 2345 DU_J19_ Channel proteins that are located in the outer 9377: barrels , MSC_BOT membrane of Gram-negative bacteria are known 9378:murins , _Q01 as 9379:porins, 9380:granules , 2 2346 DU_J19_ Gram-negative bacteria are more resistant to 9381: peptidoglycan wall., MSC_BOT antibiotics than Gram-positive bacteria due to the 9382:outer lipopolysaccharide layer., _Q02 presence of 9383:porin protein., 9384:teichoic acid. , 3 2347 DU_J19_ Dormant, tough, non-reproductive structure, 9385: endospore , MSC_BOT produced by bacteria to tide over unfavourable 9386: sclerotium , _Q03 conditions is known as: 9387:sporocarp , 9388:heterocyst , 4 2348 DU_J19_ Three-dimensional structures of a cell can be 9389: Bright field , MSC_BOT studied by using which of the following 9390: Phase contrast , _Q04 microscopes? 9391:Fluorescent , 9392: Differential interference contrast , 5 2349 DU_J19_ Algae that grow at the interface of water and 9393: epipelic , MSC_BOT atmosphere are called as 9394:neustonic , _Q05 9395: planktonic , 9396: benthic, 6 2350 DU_J19_ Orthorhombic crystals of calcium carbonate are 9397: aragonite., MSC_BOT known as 9398: calcite. , _Q06 9399: detritus. , 9400: stromatolites. , 7 2351 DU_J19_ Which one of the following amino acids has a 9401: Lysine , MSC_BOT nonpolar, aliphatic R group? _Q07 7 2351 DU_J19_ Which one of the following amino acids has a MSC_BOT nonpolar, aliphatic R group? 9402: Histidine , _Q07 9403: Arginine , 9404: Glycine, 8 2352 DU_J19_ -

Can We Understand Evolution Without Symbiogenesis?
Can We Understand Evolution Without Symbiogenesis? Francisco Carrapiço …symbiosis is more than a mere casual and isolated biological phenomenon: it is in reality the most fundamental and universal order or law of life. Hermann Reinheimer (1915) Abstract This work is a contribution to the literature and knowledge on evolu- tion that takes into account the biological data obtained on symbiosis and sym- biogenesis. Evolution is traditionally considered a gradual process essentially consisting of natural selection, conducted on minimal phenotypical variations that are the result of mutations and genetic recombinations to form new spe- cies. However, the biological world presents and involves symbiotic associations between different organisms to form consortia, a new structural life dimension and a symbiont-induced speciation. The acknowledgment of this reality implies a new understanding of the natural world, in which symbiogenesis plays an important role as an evolutive mechanism. Within this understanding, symbiosis is the key to the acquisition of new genomes and new metabolic capacities, driving living forms’ evolution and the establishment of biodiversity and complexity on Earth. This chapter provides information on some of the key figures and their major works on symbiosis and symbiogenesis and reinforces the importance of these concepts in our understanding of the natural world and the role they play in the establishing of the evolutionary complexity of living systems. In this context, the concept of the symbiogenic superorganism is also discussed. Keywords Evolution · Symbiogenesis · Symbiosis · Symbiogenic superorgan- ism · New paradigm F. Carrapiço (*) Centre for Ecology Evolution and Environmental Change (CE3C); Centre for Philosophy of Science, Department of Plant Biology, Faculty of science, University of Lisbon, Lisbon, Portugal e-mail: [email protected] © Springer International Publishing Switzerland 2015 81 N. -

Organellar Genome Evolution in Red Algal Parasites: Differences in Adelpho- and Alloparasites
University of Rhode Island DigitalCommons@URI Open Access Dissertations 2017 Organellar Genome Evolution in Red Algal Parasites: Differences in Adelpho- and Alloparasites Eric Salomaki University of Rhode Island, [email protected] Follow this and additional works at: https://digitalcommons.uri.edu/oa_diss Recommended Citation Salomaki, Eric, "Organellar Genome Evolution in Red Algal Parasites: Differences in Adelpho- and Alloparasites" (2017). Open Access Dissertations. Paper 614. https://digitalcommons.uri.edu/oa_diss/614 This Dissertation is brought to you for free and open access by DigitalCommons@URI. It has been accepted for inclusion in Open Access Dissertations by an authorized administrator of DigitalCommons@URI. For more information, please contact [email protected]. ORGANELLAR GENOME EVOLUTION IN RED ALGAL PARASITES: DIFFERENCES IN ADELPHO- AND ALLOPARASITES BY ERIC SALOMAKI A DISSERTATION SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY IN BIOLOGICAL SCIENCES UNIVERSITY OF RHODE ISLAND 2017 DOCTOR OF PHILOSOPHY DISSERTATION OF ERIC SALOMAKI APPROVED: Dissertation Committee: Major Professor Christopher E. Lane Jason Kolbe Tatiana Rynearson Nasser H. Zawia DEAN OF THE GRADUATE SCHOOL UNIVERSITY OF RHODE ISLAND 2017 ABSTRACT Parasitism is a common life strategy throughout the eukaryotic tree of life. Many devastating human pathogens, including the causative agents of malaria and toxoplasmosis, have evolved from a photosynthetic ancestor. However, how an organism transitions from a photosynthetic to a parasitic life history strategy remains mostly unknown. Parasites have independently evolved dozens of times throughout the Florideophyceae (Rhodophyta), and often infect close relatives. This framework enables direct comparisons between autotrophs and parasites to investigate the early stages of parasite evolution.