Reconstructing SARS-Cov-2 Response Signaling and Regulatory Networks
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

1 Silencing Branched-Chain Ketoacid Dehydrogenase Or
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.21.960153; this version posted February 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Silencing branched-chain ketoacid dehydrogenase or treatment with branched-chain ketoacids ex vivo inhibits muscle insulin signaling Running title: BCKAs impair insulin signaling Dipsikha Biswas1, PhD, Khoi T. Dao1, BSc, Angella Mercer1, BSc, Andrew Cowie1 , BSc, Luke Duffley1, BSc, Yassine El Hiani2, PhD, Petra C. Kienesberger1, PhD, Thomas Pulinilkunnil1†, PhD 1Department of Biochemistry and Molecular Biology, Dalhousie Medicine New Brunswick, Saint John, New Brunswick, Canada, 2Department of Physiology and Biophysics, Dalhousie University, Halifax, Nova Scotia, Canada. †Correspondence to Thomas Pulinilkunnil, PhD Department of Biochemistry and Molecular Biology, Faculty of Medicine, Dalhousie University Dalhousie Medicine New Brunswick, 100 Tucker Park Road, Saint John E2L4L5, New Brunswick, Canada. Telephone: (506) 636-6973; Fax: (506) 636-6001; email: [email protected]. 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.02.21.960153; this version posted February 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International -

Application of a MYC Degradation
SCIENCE SIGNALING | RESEARCH ARTICLE CANCER Copyright © 2019 The Authors, some rights reserved; Application of a MYC degradation screen identifies exclusive licensee American Association sensitivity to CDK9 inhibitors in KRAS-mutant for the Advancement of Science. No claim pancreatic cancer to original U.S. Devon R. Blake1, Angelina V. Vaseva2, Richard G. Hodge2, McKenzie P. Kline3, Thomas S. K. Gilbert1,4, Government Works Vikas Tyagi5, Daowei Huang5, Gabrielle C. Whiten5, Jacob E. Larson5, Xiaodong Wang2,5, Kenneth H. Pearce5, Laura E. Herring1,4, Lee M. Graves1,2,4, Stephen V. Frye2,5, Michael J. Emanuele1,2, Adrienne D. Cox1,2,6, Channing J. Der1,2* Stabilization of the MYC oncoprotein by KRAS signaling critically promotes the growth of pancreatic ductal adeno- carcinoma (PDAC). Thus, understanding how MYC protein stability is regulated may lead to effective therapies. Here, we used a previously developed, flow cytometry–based assay that screened a library of >800 protein kinase inhibitors and identified compounds that promoted either the stability or degradation of MYC in a KRAS-mutant PDAC cell line. We validated compounds that stabilized or destabilized MYC and then focused on one compound, Downloaded from UNC10112785, that induced the substantial loss of MYC protein in both two-dimensional (2D) and 3D cell cultures. We determined that this compound is a potent CDK9 inhibitor with a previously uncharacterized scaffold, caused MYC loss through both transcriptional and posttranslational mechanisms, and suppresses PDAC anchorage- dependent and anchorage-independent growth. We discovered that CDK9 enhanced MYC protein stability 62 through a previously unknown, KRAS-independent mechanism involving direct phosphorylation of MYC at Ser . -

1 Silencing Branched-Chain Ketoacid Dehydrogenase Or
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.21.960153; this version posted February 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Silencing branched-chain ketoacid dehydrogenase or treatment with branched-chain ketoacids ex vivo inhibits muscle insulin signaling Running title: BCKAs impair insulin signaling Dipsikha Biswas1, PhD, Khoi T. Dao1, BSc, Angella Mercer1, BSc, Andrew Cowie1 , BSc, Luke Duffley1, BSc, Yassine El Hiani2, PhD, Petra C. Kienesberger1, PhD, Thomas Pulinilkunnil1†, PhD 1Department of Biochemistry and Molecular Biology, Dalhousie Medicine New Brunswick, Saint John, New Brunswick, Canada, 2Department of Physiology and Biophysics, Dalhousie University, Halifax, Nova Scotia, Canada. †Correspondence to Thomas Pulinilkunnil, PhD Department of Biochemistry and Molecular Biology, Faculty of Medicine, Dalhousie University Dalhousie Medicine New Brunswick, 100 Tucker Park Road, Saint John E2L4L5, New Brunswick, Canada. Telephone: (506) 636-6973; Fax: (506) 636-6001; email: [email protected]. 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.02.21.960153; this version posted February 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International -
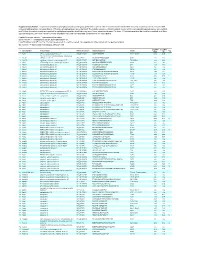
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

EGL Test Description
2460 Mountain Industrial Boulevard | Tucker, Georgia 30084 Phone: 470-378-2200 or 855-831-7447 | Fax: 470-378-2250 eglgenetics.com Autism Spectrum Disorders: Tier 2 Deletion/Duplication Panel Test Code: MD021 Turnaround time: 2 weeks CPT Codes: 81161 x1, 81323 x1, 81403 x1, 81404 x1, 81405 x1, 81406 x1 Condition Description Genetics of Autism Spectrum Disorders Autism spectrum disorders (ASDs) are a group of neurodevelopmental disorders which include autism, pervasive developmental delay-not otherwise specified (PDD-NOS), and Asperger syndrome. ASDs are characterized by impairments in social relationships, variable degrees of language and communication deficits, and repetitive behaviors and/or a narrow range of interests. The age of onset is prior to age 3 with a variable clinical presentation, ranging in severity both amongst individuals as well as amongst the various subtypes of ASDs. Additional clinical features may also be observed in individuals with an ASD, such as intellectual disability (up to ~50%) and seizures (~25%). Known genetic causes of autism include cytogenetically visible chromosome abnormalities (3-5%), copy number variants – which include submicroscopic deletions and duplications (~6-7%), and single gene disorders (~5%). EGL Genetics’s integrated testing strategy allows for a comprehensive cytogenetics, metabolic, and molecular analysis of ASD in your patient. For a summary of autism testing at EGL, please click here. *Please note that some genes on this panel are associated with additional phenotypes. References: Autism and Developmental Disabilities Monitoring Network Surveillance Year 2006 Principal Investigators and the CDC (2009). MMWR Surveill Summ, 58:1-20. Bolton et al. (2011). Br J Psychiatry, 198:289-294. -

Genome-Wide CRISPR Screen for PARKIN Regulators Reveals Transcriptional Repression As a Determinant of Mitophagy
Genome-wide CRISPR screen for PARKIN regulators PNAS PLUS reveals transcriptional repression as a determinant of mitophagy Christoph Pottinga, Christophe Crochemorea, Francesca Morettia, Florian Nigscha, Isabel Schmidta, Carole Mannevillea, Walter Carbonea, Judith Knehra, Rowena DeJesusb, Alicia Lindemanb, Rob Maherb, Carsten Russb, Gregory McAllisterb, John S. Reece-Hoyesb, Gregory R. Hoffmanb, Guglielmo Romaa, Matthias Müllera, Andreas W. Sailera, and Stephen B. Helliwella,1 aNovartis Institutes for BioMedical Research, Basel CH 4002, Switzerland; and bNovartis Institutes for BioMedical Research, Cambridge, MA 02139 Edited by Beth Levine, The University of Texas Southwestern, Dallas, TX, and approved November 29, 2017 (received for review June 20, 2017) PARKIN, an E3 ligase mutated in familial Parkinson’s disease, promotes how these proteins are themselves regulated, is important for mitophagy by ubiquitinating mitochondrial proteins for efficient the further understanding of mitophagy and the pathogenesis engagement of the autophagy machinery. Specifically, PARKIN- of Parkinson’s disease. synthesized ubiquitin chains represent targets for the PINK1 kinase Previous studies identified regulators of PINK1/PARKIN- generating phosphoS65-ubiquitin (pUb), which constitutes the mitoph- mediated mitophagy employing RNAi screens with damage- agy signal. Physiological regulation of PARKIN abundance, however, induced mitochondrial translocation of overexpressed GFP- and the impact on pUb accumulation are poorly understood. Using PARKIN as a mitophagy proxy (12–15). These efforts pro- cells designed to discover physiological regulators of PARKIN abun- foundly advanced the understanding of mitophagy regulation; dance, we performed a pooled genome-wide CRISPR/Cas9 knockout however, little is known about how cells set the threshold for screen. Testing identified genes individually resulted in a list of mitophagy to proceed. -

Phosphorylation of BCKDK of BCAA Catabolism at Y246 by Src Promotes Metastasis of Colorectal Cancer
Oncogene (2020) 39:3980–3996 https://doi.org/10.1038/s41388-020-1262-z ARTICLE Phosphorylation of BCKDK of BCAA catabolism at Y246 by Src promotes metastasis of colorectal cancer 1 1 2 3 1,4 1 1 1 Qin Tian ● Ping Yuan ● Chuntao Quan ● Mingyang Li ● Juanjuan Xiao ● Lu Zhang ● Hui Lu ● Tengfei Ma ● 1 1 5 1 1 1 3 1,4 1 Ling Zou ● Fei Wang ● Peipei Xue ● Xiaofang Ni ● Wei Wang ● Lin Liu ● Zhe Wang ● Feng Zhu ● Qiuhong Duan Received: 24 November 2019 / Revised: 4 March 2020 / Accepted: 6 March 2020 / Published online: 1 April 2020 © The Author(s) 2020. This article is published with open access Abstract Branched-chain α-keto acid dehydrogenase kinase (BCKDK), the key enzyme of branched-chain amino acids (BCAAs) metabolism, has been reported to promote colorectal cancer (CRC) tumorigenesis by upregulating the MEK-ERK signaling pathway. However, the profile of BCKDK in metastatic colorectal cancer (mCRC) remains unknown. Here, we report a novel role of BCKDK in mCRC. BCKDK is upregulated in CRC tissues. Increased BCKDK expression was associated with metastasis and poor clinical prognosis in CRC patients. Knockdown of BCKDK decreased CRC cell migration and invasion ex vivo, and lung metastasis in vivo. BCKDK promoted the epithelial mesenchymal transition (EMT) program, by 1234567890();,: 1234567890();,: decreasing the expression of E-cadherin, epithelial marker, and increasing the expression of N-cadherin and Vimentin, which are mesenchymal markers. Moreover, BCKDK-knockdown experiments in combination with phosphoproteomics analysis revealed the potent role of BCKDK in modulating multiple signal transduction pathways, including EMT and metastasis. -

BCKDK: an Emerging Kinase Target for Metabolic Diseases and Cancer
NEWS & ANALYSIS TARGET WATCH tolerated, with no signs of apparent toxicity, further supporting the therapeutic promise for BCAA- associated metabolic diseases. BCKDK: an emerging kinase target for In patients with colorectal cancer, high expression of BCKDK was associated metabolic diseases and cancer with poor prognosis. In experiments in colorectal cancer cell lines and xenograft The understudied kinase branched-chain The rate- limiting step of BCAA catabolism and models, BCKDK promoted tumorigenesis ketoacid dehydrogenase kinase (BCKDK) has clearance is the irreversible decarboxylation through activation of the MEK/ERK pathway recently been implicated in various human of BCKAs by the branched-chain α- ketoacid (EBioMedicine 20, 50–60; 2017). diseases. In addition to its established role in dehydrogenase complex (BCKDH). BCKDH regulating the catabolic flux of branched-chain activity is inhibited by phosphorylation of the Chemical tools amino acids (BCAAs), BCKDK was also E1α subunit by BCKDK. The literature landscape for BCKDK found to enhance MEK/ERK signalling, Inhibition of BCKDK with the selective, small- molecule inhibitors has focused on a key pathway driving cell growth and allosteric inhibitor BT2 (FIG. 1) was sufficient the development of allosteric inhibitors. proliferation in cancer. So, BCKDK has to enhance BCKDH activity across tissue These inhibitors bind to BCKDK, triggering become an attractive therapeutic target for types in insulin-resistant, diet- induced obese movements within the N-terminal domain BCAA- associated diseases and cancers. mice (Diabetes 68, 1730–1746; 2019). This helix that result in separation of BCKDK from correlated with lower plasma concentrations BCKDH. This separation works in tandem Biological functions of BCAAs and BCKAs. -

Download 20190410); Fragmentation for 20 S
ARTICLE https://doi.org/10.1038/s41467-020-17387-y OPEN Multi-layered proteomic analyses decode compositional and functional effects of cancer mutations on kinase complexes ✉ Martin Mehnert 1 , Rodolfo Ciuffa1, Fabian Frommelt 1, Federico Uliana1, Audrey van Drogen1, ✉ ✉ Kilian Ruminski1,3, Matthias Gstaiger1 & Ruedi Aebersold 1,2 fi 1234567890():,; Rapidly increasing availability of genomic data and ensuing identi cation of disease asso- ciated mutations allows for an unbiased insight into genetic drivers of disease development. However, determination of molecular mechanisms by which individual genomic changes affect biochemical processes remains a major challenge. Here, we develop a multilayered proteomic workflow to explore how genetic lesions modulate the proteome and are trans- lated into molecular phenotypes. Using this workflow we determine how expression of a panel of disease-associated mutations in the Dyrk2 protein kinase alter the composition, topology and activity of this kinase complex as well as the phosphoproteomic state of the cell. The data show that altered protein-protein interactions caused by the mutations are asso- ciated with topological changes and affected phosphorylation of known cancer driver pro- teins, thus linking Dyrk2 mutations with cancer-related biochemical processes. Overall, we discover multiple mutation-specific functionally relevant changes, thus highlighting the extensive plasticity of molecular responses to genetic lesions. 1 Department of Biology, Institute of Molecular Systems Biology, ETH Zurich, -

Genome-Wide Identification and Evolutionary Analysis of Sarcocystis Neurona Protein Kinases
Article Genome-Wide Identification and Evolutionary Analysis of Sarcocystis neurona Protein Kinases Edwin K. Murungi 1,* and Henry M. Kariithi 2 1 Department of Biochemistry and Molecular Biology, Egerton University, P.O. Box 536, 20115 Njoro, Kenya 2 Biotechnology Research Institute, Kenya Agricultural and Livestock Research Organization, P.O. Box 57811, Kaptagat Rd, Loresho, 00200 Nairobi, Kenya; [email protected] * Correspondence: [email protected]; Tel: +254-789-716-059 Academic Editor: Anthony Underwood Received: 6 January 2017; Accepted: 17 March 2017; Published: 21 March 2017 Abstract: The apicomplexan parasite Sarcocystis neurona causes equine protozoal myeloencephalitis (EPM), a degenerative neurological disease of horses. Due to its host range expansion, S. neurona is an emerging threat that requires close monitoring. In apicomplexans, protein kinases (PKs) have been implicated in a myriad of critical functions, such as host cell invasion, cell cycle progression and host immune response evasion. Here, we used various bioinformatics methods to define the kinome of S. neurona and phylogenetic relatedness of its PKs to other apicomplexans. We identified 97 putative PKs clustering within the various eukaryotic kinase groups. Although containing the universally-conserved PKA (AGC group), S. neurona kinome was devoid of PKB and PKC. Moreover, the kinome contains the six-conserved apicomplexan CDPKs (CAMK group). Several OPK atypical kinases, including ROPKs 19A, 27, 30, 33, 35 and 37 were identified. Notably, S. neurona is devoid of the virulence-associated ROPKs 5, 6, 18 and 38, as well as the Alpha and RIO kinases. Two out of the three S. neurona CK1 enzymes had high sequence similarities to Toxoplasma gondii TgCK1-α and TgCK1-β and the Plasmodium PfCK1. -

Gene Symbol Accession Alias/Prev Symbol Official Full Name AAK1 NM 014911.2 KIAA1048, Dkfzp686k16132 AP2 Associated Kinase 1
Gene Symbol Accession Alias/Prev Symbol Official Full Name AAK1 NM_014911.2 KIAA1048, DKFZp686K16132 AP2 associated kinase 1 (AAK1) AATK NM_001080395.2 AATYK, AATYK1, KIAA0641, LMR1, LMTK1, p35BP apoptosis-associated tyrosine kinase (AATK) ABL1 NM_007313.2 ABL, JTK7, c-ABL, p150 v-abl Abelson murine leukemia viral oncogene homolog 1 (ABL1) ABL2 NM_007314.3 ABLL, ARG v-abl Abelson murine leukemia viral oncogene homolog 2 (arg, Abelson-related gene) (ABL2) ACVR1 NM_001105.2 ACVRLK2, SKR1, ALK2, ACVR1A activin A receptor ACVR1B NM_004302.3 ACVRLK4, ALK4, SKR2, ActRIB activin A receptor, type IB (ACVR1B) ACVR1C NM_145259.2 ACVRLK7, ALK7 activin A receptor, type IC (ACVR1C) ACVR2A NM_001616.3 ACVR2, ACTRII activin A receptor ACVR2B NM_001106.2 ActR-IIB activin A receptor ACVRL1 NM_000020.1 ACVRLK1, ORW2, HHT2, ALK1, HHT activin A receptor type II-like 1 (ACVRL1) ADCK1 NM_020421.2 FLJ39600 aarF domain containing kinase 1 (ADCK1) ADCK2 NM_052853.3 MGC20727 aarF domain containing kinase 2 (ADCK2) ADCK3 NM_020247.3 CABC1, COQ8, SCAR9 chaperone, ABC1 activity of bc1 complex like (S. pombe) (CABC1) ADCK4 NM_024876.3 aarF domain containing kinase 4 (ADCK4) ADCK5 NM_174922.3 FLJ35454 aarF domain containing kinase 5 (ADCK5) ADRBK1 NM_001619.2 GRK2, BARK1 adrenergic, beta, receptor kinase 1 (ADRBK1) ADRBK2 NM_005160.2 GRK3, BARK2 adrenergic, beta, receptor kinase 2 (ADRBK2) AKT1 NM_001014431.1 RAC, PKB, PRKBA, AKT v-akt murine thymoma viral oncogene homolog 1 (AKT1) AKT2 NM_001626.2 v-akt murine thymoma viral oncogene homolog 2 (AKT2) AKT3 NM_181690.1