Mouse Bag5 Conditional Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Exploring the Relationship Between Gut Microbiota and Major Depressive Disorders
E3S Web of Conferences 271, 03055 (2021) https://doi.org/10.1051/e3sconf/202127103055 ICEPE 2021 Exploring the Relationship between Gut Microbiota and Major Depressive Disorders Catherine Tian1 1Shanghai American School, Shanghai, China Abstract. Major Depressive Disorder (MDD) is a psychiatric disorder accompanied with a high rate of suicide, morbidity and mortality. With the symptom of an increasing or decreasing appetite, there is a possibility that MDD may have certain connections with gut microbiota, the colonies of microbes which reside in the human digestive system. In recent years, more and more studies started to demonstrate the links between MDD and gut microbiota from animal disease models and human metabolism studies. However, this relationship is still largely understudied, but it is very innovative since functional dissection of this relationship would furnish a new train of thought for more effective treatment of MDD. In this study, by using multiple genetic analytic tools including Allen Brain Atlas, genetic function analytical tools, and MicrobiomeAnalyst, I explored the genes that shows both expression in the brain and the digestive system to affirm that there is a connection between gut microbiota and the MDD. My approach finally identified 7 MDD genes likely to be associated with gut microbiota, implicating 3 molecular pathways: (1) Wnt Signaling, (2) citric acid cycle in the aerobic respiration, and (3) extracellular exosome signaling. These findings may shed light on new directions to understand the mechanism of MDD, potentially facilitating the development of probiotics for better psychiatric disorder treatment. 1 Introduction 1.1 Major Depressive Disorder Major Depressive Disorder (MDD) is a mood disorder that will affect the mood, behavior and other physical parts. -

Profiling Haplotype Specific Cpg and Cph Methylation Within A
www.nature.com/scientificreports OPEN Profling haplotype specifc CpG and CpH methylation within a schizophrenia GWAS locus on chromosome 14 in schizophrenia and healthy subjects Margarita Alfmova*, Nikolay Kondratyev, Arkadiy Golov & Vera Golimbet Interrogating DNA methylation within schizophrenia risk loci holds promise to identify mechanisms by which genes infuence the disease. Based on the hypothesis that allele specifc methylation (ASM) of a single CpG, or perhaps CpH, might mediate or mark the efects of genetic variants on disease risk and phenotypes, we explored haplotype specifc methylation levels of individual cytosines within a genomic region harbouring the BAG5, APOPT1 and KLC1 genes in peripheral blood of schizophrenia patients and healthy controls. Three DNA fragments located in promoter, intronic and intergenic areas were studied by single-molecule real-time bisulfte sequencing enabling the analysis of long reads of DNA with base-pair resolution and the determination of haplotypes directly from sequencing data. Among 1,012 cytosines studied, we did not fnd any site where methylation correlated with the disease or cognitive defcits after correction for multiple testing. At the same time, we determined the methylation profle associated with the schizophrenia risk haplotype within the KLC1 fourth intron and confrmed ASM for cytosines located in the vicinity of rs67899457. These genetically associated DNA methylation variations may be related to the pathophysiological mechanism diferentiating the risk and non-risk haplotypes and merit further investigation. Schizophrenia is a common, highly heritable disorder characterized by positive, negative, and cognitive symp- toms. Large genome-wide association studies (GWAS) of the Psychiatric Genomics Consortium (PGC) have identifed more than 100 genomic regions that are signifcantly associated with schizophrenia1,2. -

Occurrence and Identification of Emeria Species in Broiler Rearing
Tran et al. Journal of Animal Science and Technology (2015) 57:44 DOI 10.1186/s40781-015-0077-x RESEARCH Open Access In silico characterisation, homology modelling and structure-based functional annotation of blunt snout bream (Megalobrama amblycephala) Hsp70 and Hsc70 proteins Ngoc Tuan Tran1,3, Ivan Jakovlić1 and Wei-Min Wang1,2* Abstract Background: Heat shock proteins play an important role in protection from stress stimuli and metabolic insults in almost all organisms. Methods: In this study, computational tools were used to deeply analyse the physicochemical characteristics and, using homology modelling, reliably predict the tertiary structure of the blunt snout bream (Ma-) Hsp70 and Hsc70 proteins. Derived three-dimensional models were then used to predict the function of the proteins. Results: Previously published predictions regarding the protein length, molecular weight, theoretical isoelectric point and total number of positive and negative residues were corroborated. Among the new findings are: the extinction coefficient (33725/33350 and 35090/34840 - Ma-Hsp70/ Ma-Hsc70, respectively), instability index (33.68/35.56 – both stable), aliphatic index (83.44/80.23 – both very stable), half-life estimates (both relatively stable), grand average of hydropathicity (−0.431/-0.473 – both hydrophilic) and amino acid composition (alanine-lysine-glycine/glycine-lysine-aspartic acid were the most abundant, no disulphide bonds, the N-terminal of both proteins was methionine). Homology modelling was performed by SWISS-MODEL program and the proposed model was evaluated as highly reliable based on PROCHECK’s Ramachandran plot, ERRAT, PROVE, Verify 3D, ProQ and ProSA analyses. Conclusions: The research revealed a high structural similarity to Hsp70 and Hsc70 proteins from several taxonomically distant animal species, corroborating a remarkably high level of evolutionary conservation among the members of this protein family. -
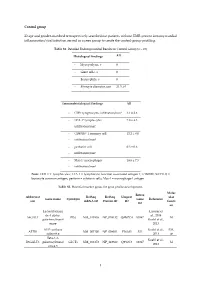
1 Control Group 23 Age and Gender-Matched
Control group 23 age and gender-matched retrospectively searched out patients without EMB-proven intramyocardial inflammation/viral infection served as a peer group to create the control-group profiling. Table S1. Detailed Endomyocardial Results in Control Group (n = 23) Histological findings All - Myocytolysis, n 0 - Giant cells, n 0 - Eosinophils, n 0 - Myocyte diameter, µm 21.3 ± 0 Immunohistological findings All - CD3+ lymphocytes infiltration/mm² 3.1 ± 2.6 - LFA-1+ lymphocytes 7.8 ± 4.3 infiltration/mm² - CD45R0+ T memory cell 12.2 ± 8.0 - infiltration/mm² - perforin+ cell 0.3 ± 0.6 - infiltration/mm² - Mac-1+ macrophages 18.6 ± 7.5 - infiltration/mm² Note: CD3 = T lymphocytes; LFA-1 = lymphocyte function-associated antigen-1; CD45R0 (UCHL1) = leucocyte common antigen; perforin = cytotoxic cells; Mac-1 = macrophage-1 antigen. Table S2. Potential marker genes for gene profile development. Molec Entrez Abbreviat RefSeq RefSeq Uniprot ular Gene name Synonym Gene Reference ion mRNA ID Protein ID ID functi ID on Lactosylcerami Lassner et de 4-alpha- al., 2014 A4GALT P(k) NM_017436 NP_059132 Q9NPC4 53947 M galactosyltransf Kuehl et al., erase 2013 ATP synthase Kuehl et al., EM, ATP5I NM_007100 NP_009031 P56385 521 subunit g 2013 ce Beta-1,3- Kuehl et al., B3GALT5 galactosyltransf GLCT5 NM_033172 NP_149362 Q9Y2C3 10317 M 2013 erase 5 1 BAG family molecular Kuehl et al., BAG5 NM_004873 NP_004864 Q9UL15 9529 CC chaperone 2013 regulator 5 Lassner et CC-chemokine al., 2014 CCL20 ST38 NM_004591 NP_004582 P78556 6364 IR ligand 20 Kuehl et al., -

Prioritization of Candidate Disease Genes by Combining Topological Similarity and Semantic Similarity ⇑ Bin Liu, Min Jin , Pan Zeng
Journal of Biomedical Informatics 57 (2015) 1–5 Contents lists available at ScienceDirect Journal of Biomedical Informatics journal homepage: www.elsevier.com/locate/yjbin Prioritization of candidate disease genes by combining topological similarity and semantic similarity ⇑ Bin Liu, Min Jin , Pan Zeng College of Computer Science and Electronic Engineering, Hunan University, Changsha 410082, China article info abstract Article history: The identification of gene–phenotype relationships is very important for the treatment of human Received 26 December 2014 diseases. Studies have shown that genes causing the same or similar phenotypes tend to interact with Revised 1 July 2015 each other in a protein–protein interaction (PPI) network. Thus, many identification methods based on Accepted 6 July 2015 the PPI network model have achieved good results. However, in the PPI network, some interactions Available online 11 July 2015 between the proteins encoded by candidate gene and the proteins encoded by known disease genes are very weak. Therefore, some studies have combined the PPI network with other genomic information Keywords: and reported good predictive performances. However, we believe that the results could be further Disease genes improved. In this paper, we propose a new method that uses the semantic similarity between the candi- Random walk Topological similarity date gene and known disease genes to set the initial probability vector of a random walk with a restart Semantic similarity algorithm in a human PPI network. The effectiveness of our method was demonstrated by leave-one-out cross-validation, and the experimental results indicated that our method outperformed other methods. Additionally, our method can predict new causative genes of multifactor diseases, including Parkinson’s disease, breast cancer and obesity. -

A SARS-Cov-2 Protein Interaction Map Reveals Targets for Drug Repurposing
Article A SARS-CoV-2 protein interaction map reveals targets for drug repurposing https://doi.org/10.1038/s41586-020-2286-9 A list of authors and affiliations appears at the end of the paper Received: 23 March 2020 Accepted: 22 April 2020 A newly described coronavirus named severe acute respiratory syndrome Published online: 30 April 2020 coronavirus 2 (SARS-CoV-2), which is the causative agent of coronavirus disease 2019 (COVID-19), has infected over 2.3 million people, led to the death of more than Check for updates 160,000 individuals and caused worldwide social and economic disruption1,2. There are no antiviral drugs with proven clinical efcacy for the treatment of COVID-19, nor are there any vaccines that prevent infection with SARS-CoV-2, and eforts to develop drugs and vaccines are hampered by the limited knowledge of the molecular details of how SARS-CoV-2 infects cells. Here we cloned, tagged and expressed 26 of the 29 SARS-CoV-2 proteins in human cells and identifed the human proteins that physically associated with each of the SARS-CoV-2 proteins using afnity-purifcation mass spectrometry, identifying 332 high-confdence protein–protein interactions between SARS-CoV-2 and human proteins. Among these, we identify 66 druggable human proteins or host factors targeted by 69 compounds (of which, 29 drugs are approved by the US Food and Drug Administration, 12 are in clinical trials and 28 are preclinical compounds). We screened a subset of these in multiple viral assays and found two sets of pharmacological agents that displayed antiviral activity: inhibitors of mRNA translation and predicted regulators of the sigma-1 and sigma-2 receptors. -

Cam/BAG5/Hsc70 Signaling Complex Dynamically Regulates Leaf
www.nature.com/scientificreports OPEN CaM/BAG5/Hsc70 signaling complex dynamically regulates leaf senescence Received: 27 June 2016 Luhua Li1,2, Yangfei Xing1,2, Dong Chang1,2, Shasha Fang2, Boyang Cui2, Qi Li2, Xuejie Wang2, Accepted: 28 July 2016 Shang Guo1, Xue Yang1, Shuzhen Men2 & Yuequan Shen1,2,3 Published: 19 August 2016 Calcium signaling plays an essential role in plant cell physiology, and chaperone-mediated protein folding directly regulates plant programmed cell death. The Arabidopsis thaliana protein AtBAG5 (Bcl-2-associated athanogene 5) is unique in that it contains both a BAG domain capable of binding Hsc70 (Heat shock cognate protein 70) and a characteristic IQ motif that is specific for Ca2+-free CaM (Calmodulin) binding and hence acts as a hub linking calcium signaling and the chaperone system. Here, we determined crystal structures of AtBAG5 alone and in complex with Ca2+-free CaM. Structural and biochemical studies revealed that Ca2+-free CaM and Hsc70 bind AtBAG5 independently, whereas Ca2+-saturated CaM and Hsc70 bind AtBAG5 with negative cooperativity. Further in vivo studies confirmed that AtBAG5 localizes to mitochondria and that its overexpression leads to leaf senescence symptoms including decreased chlorophyll retention and massive ROS production in dark-induced plants. Mutants interfering the CaM/AtBAG5/Hsc70 complex formation leads to different phenotype of leaf senescence. Collectively, we propose that the CaM/AtBAG5/Hsc70 signaling complex plays an important role in regulating plant senescence. Senescence occurs at the final stage of leaf development in plants and is a critical process for the recycling of nutrients from senescent leaves to other parts of the plant. -

Qt8b18n6c6 Nosplash Caf9e24
Copyright 2014 by Christopher K. Fuller ii For Kai and Lena iii Acknowledgements This work would not have been possible without the support of my research advisor Hao Li. His broad base of interests gave me the freedom to search for something that so well matches my inclinations. I appreciate his insight, enthusiasm, skepticism, and overall encouragement of this effort. I wish to thank my committee, Saunak Sen and Kathleen Giacomini, for their suggestions and support. I wish to thank postdoctoral scholar Xin He for catalyzing the research that became the focus of my work. In addition, I wish to thank postdoctoral scholar Jiashun Zheng for his numerous suggestions and support. I wish to thank the thousands of anonymous individuals who make integrative genomics research possible by providing samples and the researchers dedicated to making these widely available. In addition, I wish to thank the MuTHER and DIAGRAM consortia for providing access to their full summary results. I wish to thank my wife, Sharoni, for her support and for permitting me to encroach on her domain of expertise. You are still the real biologist in the family. Finally, I wish to thank my Mom and Dad ... who took me to the library. iv Abstract Genome-wide association studies (GWAS) have linked various complex diseases to many dozens, sometimes hundreds, of individual genomic loci. Since these are generally of small effect and may lack both functional annotations and an obvious relation to other disease-associated regions, they are difficult to place in a functional context that advances our understanding of the disease. -

HSP70 Multi-Functionality in Cancer
cells Review HSP70 Multi-Functionality in Cancer Zarema Albakova 1,2,* , Grigoriy A. Armeev 1, Leonid M. Kanevskiy 2, Elena I. Kovalenko 2 and Alexander M. Sapozhnikov 1,2 1 Department of Biology, Lomonosov Moscow State University, 119192 Moscow, Russia; [email protected] (G.A.A.); [email protected] (A.M.S.) 2 Department of Immunology, Shemyakin and Ovchinnikov Institute of Bioorganic Chemistry of the Russian Academy of Sciences, 117997 Moscow, Russia; [email protected] (L.M.K.); [email protected] (E.I.K.) * Correspondence: [email protected] Received: 30 January 2020; Accepted: 28 February 2020; Published: 2 March 2020 Abstract: The 70-kDa heat shock proteins (HSP70s) are abundantly present in cancer, providing malignant cells selective advantage by suppressing multiple apoptotic pathways, regulating necrosis, bypassing cellular senescence program, interfering with tumor immunity, promoting angiogenesis and supporting metastasis. This direct involvement of HSP70 in most of the cancer hallmarks explains the phenomenon of cancer “addiction” to HSP70, tightly linking tumor survival and growth to the HSP70 expression. HSP70 operates in different states through its catalytic cycle, suggesting that it can multi-function in malignant cells in any of these states. Clinically, tumor cells intensively release HSP70 in extracellular microenvironment, resulting in diverse outcomes for patient survival. Given its clinical significance, small molecule inhibitors were developed to target different sites of the HSP70 machinery. Furthermore, several HSP70-based immunotherapy approaches were assessed in clinical trials. This review will explore different roles of HSP70 on cancer progression and emphasize the importance of understanding the flexibility of HSP70 nature for future development of anti-cancer therapies. -

Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name BCL2-associated athanogene 5 Gene Symbol BAG5 Organism Human Gene Summary The protein encoded by this gene is a member of the BAG1-related protein family. BAG1 is an anti-apoptotic protein that functions through interactions with a variety of cell apoptosis and growth related proteins including BCL-2 Raf-protein kinase steroid hormone receptors growth factor receptors and members of the heat shock protein 70 kDa family. This protein contains a BAG domain near the C-terminus which could bind and inhibit the chaperone activity of Hsc70/Hsp70. Three transcript variants encoding two different isoforms have been found for this gene. Gene Aliases BAG-5 RefSeq Accession No. NC_000014.8, NT_026437.12 UniGene ID Hs.5443 Ensembl Gene ID ENSG00000166170 Entrez Gene ID 9529 Assay Information Unique Assay ID qHsaCED0001492 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000445922, ENST00000337322, ENST00000557666, ENST00000299204 Amplicon Context Sequence GACGTTCTGTCTCCTGTGCTGCCCGCTTCCTAGCTTGCTGAATATCTCCTTTTCC TTCAGTATCTACAGAGTCTATTTCAAAAAGCTGTTTTGTTAGAATCCTCTCCAGTT TCTTGTAATTCTTGTCATCTGACAGACCACTGAAGC Amplicon Length (bp) 117 Chromosome Location 14:104027270-104027416 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 96 R2 0.9998 cDNA Cq 19.78 cDNA Tm (Celsius) 78.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 23.47 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation -

Multilevel SNP Analysis for Salmonella Response in Young Chicks Jason Robert Hasenstein Iowa State University
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 2007 Single, several, saturated: multilevel SNP analysis for Salmonella response in young chicks Jason Robert Hasenstein Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Genetics and Genomics Commons Recommended Citation Hasenstein, Jason Robert, "Single, several, saturated: multilevel SNP analysis for Salmonella response in young chicks" (2007). Retrospective Theses and Dissertations. 15994. https://lib.dr.iastate.edu/rtd/15994 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. i Single, several, saturated: Multilevel SNP a nalysis for Salmonella response in young chicks by Jason Robert Hasenstein A dissertation submitted to the graduate faculty in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY Major: Genetics Program of Study Committee: Susan J. Lamont, Major Professor Jack C. M. Dekkers Max F. Rothschild D. L. ( Hank ) Harris Jean -Luc Jannink Iowa State University Ames, Iowa 2007 Copyright © Jason Robert Hasenstein, 2007. All Rights Reserv ed. UMI Number: 3274823 UMI Microform 3274823 Copyright 2007 by ProQuest Information and -

Biochemical Characterization of Binding Partners of Two Hsp70 Co-Chaperones in Saccharomyces Cerevisiae
The Texas Medical Center Library DigitalCommons@TMC The University of Texas MD Anderson Cancer Center UTHealth Graduate School of The University of Texas MD Anderson Cancer Biomedical Sciences Dissertations and Theses Center UTHealth Graduate School of (Open Access) Biomedical Sciences 12-2012 BIOCHEMICAL CHARACTERIZATION OF BINDING PARTNERS OF TWO HSP70 CO-CHAPERONES IN SACCHAROMYCES CEREVISIAE Jacob Verghese Follow this and additional works at: https://digitalcommons.library.tmc.edu/utgsbs_dissertations Part of the Biochemistry Commons, Cell Biology Commons, Medicine and Health Sciences Commons, and the Microbiology Commons Recommended Citation Verghese, Jacob, "BIOCHEMICAL CHARACTERIZATION OF BINDING PARTNERS OF TWO HSP70 CO- CHAPERONES IN SACCHAROMYCES CEREVISIAE" (2012). The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access). 301. https://digitalcommons.library.tmc.edu/utgsbs_dissertations/301 This Dissertation (PhD) is brought to you for free and open access by the The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences at DigitalCommons@TMC. It has been accepted for inclusion in The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access) by an authorized administrator of DigitalCommons@TMC. For more information, please contact [email protected]. BIOCHEMICAL CHARACTERIZATION OF BINDING PARTNERS OF TWO HSP70 CO-CHAPERONES IN SACCHAROMYCES CEREVISIAE A DISSERTATION Presented to the Faculty of The University of Texas Health Science Center at Houston and The University of Texas M.D. Anderson Cancer Center Graduate School of Biomedical Sciences in Partial Fulfillment of the Requirements for the Degree of DOCTOR OF PHILOSOPHY by Jacob Verghese, B.Sc., M.S.