Ribosome•Rela Structures Reveal the Mechanism of Stringent Response Activation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Hns Gene of Escherichia Coli Is Transcriptionally Down-Regulated by (P)Ppgpp
microorganisms Article The hns Gene of Escherichia coli Is Transcriptionally Down-Regulated by (p)ppGpp Anna Brandi, Mara Giangrossi, Attilio Fabbretti and Maurizio Falconi * School of Biosciences and Veterinary Medicine, University of Camerino, 62032 Camerino, Italy; [email protected] (A.B.); [email protected] (M.G.); [email protected] (A.F.) * Correspondence: [email protected]; Tel.: +39-0737-403274 Received: 25 August 2020; Accepted: 8 October 2020; Published: 10 October 2020 Abstract: Second messenger nucleotides, such as guanosine penta- or tetra-phosphate, commonly referred to as (p)ppGpp, are powerful signaling molecules, used by all bacteria to fine-tune cellular metabolism in response to nutrient availability. Indeed, under nutritional starvation, accumulation of (p)ppGpp reduces cell growth, inhibits stable RNAs synthesis, and selectively up- or down- regulates the expression of a large number of genes. Here, we show that the E. coli hns promoter responds to intracellular level of (p)ppGpp. hns encodes the DNA binding protein H-NS, one of the major components of bacterial nucleoid. Currently, H-NS is viewed as a global regulator of transcription in an environment-dependent mode. Combining results from relA (ppGpp synthetase) and spoT (ppGpp synthetase/hydrolase) null mutants with those from an inducible plasmid encoded RelA system, we have found that hns expression is inversely correlated with the intracellular concentration of (p)ppGpp, particularly in exponential phase of growth. Furthermore, we have reproduced in an in vitro system the observed in vivo (p)ppGpp-mediated transcriptional repression of hns promoter. Electrophoretic mobility shift assays clearly demonstrated that this unusual nucleotide negatively affects the stability of RNA polymerase-hns promoter complex. -

Développement D'hybrides Aptamère-Peptide
UNIVERSITÉ DU QUÉBEC INSTITUT NATIONAL DE LA RECHERCHE SCIENTIFIQUE INSTITUT ARMAND-FRAPPIER DÉVELOPPEMENT D’HYBRIDES APTAMÈRE-PEPTIDE ANTIMICROBIEN UN MODÈLE POUR CIBLER DES BACTÉRIES PATHOGÈNES Par Amal Thamri Mémoire présenté pour l’obtention du grade de Maître en sciences (M.Sc.) en microbiologie appliquée Jury d’évaluation Examinateur externe: Dr. Roger.C Lévesque, Université de Laval Examinateur interne: Dr. Frédéric Veyrier, INRS-IAF Directeur de recherche: Dr. Jonathan Perreault, INRS-IAF Co-directrice de recherche: Dre. Annie Castonguay, INRS-IAF Remerciements Cette maîtrise a été réalisée dans le cadre d’une coopération entre INRS et le ministère d’enseignement supérieur tunisien. Les recherches objets de ce mémoire ont été réalisées dans différents laboratoires de l’institut Armand-Frappier: Laboratoires des professeurs: Jonathan Perreault, Annie Castonguay, Eric Déziel, David Chatenet. Je tiens à remercier toutes les personnes qui ont participé de près ou de loin au bon déroulement et à l’avancement des travaux de cette maîtrise. Je remercie spécialement mon directeur de recherche Jonathan Perreault pour avoir cru en mes capacités d’intégrer sa jeune équipe et travailler sur un projet novateur. Je remercie très sincèrement ma co-directrice Annie Castonguay pour ses encouragements tout au long de ces deux années. Je les remercie également d'avoir bien assuré la direction et l'encadrement de mes travaux de recherche. Merci au professeur Eric Déziel pour sa compréhension et ses conseils précieux. J’adresse aussi mes remerciements au professeur David Chatenet pour sa disponibilité, ses directions bénéfiques et ses remarques pertinentes. J’adresse mes sentiments de respect et de gratitude à Myriam Letourneau et Marie- Christine Groleau pour leur générosité et leur aide à mettre en place et améliorer plusieurs expériences. -

The Link Between Purine Metabolism and Production of Antibiotics in Streptomyces
antibiotics Review The Link between Purine Metabolism and Production of Antibiotics in Streptomyces Smitha Sivapragasam and Anne Grove * Department of Biological Sciences, Louisiana State University, Baton Rouge, LA 70803, USA; [email protected] * Correspondence: [email protected] Received: 10 May 2019; Accepted: 3 June 2019; Published: 6 June 2019 Abstract: Stress and starvation causes bacterial cells to activate the stringent response. This results in down-regulation of energy-requiring processes related to growth, as well as an upregulation of genes associated with survival and stress responses. Guanosine tetra- and pentaphosphates (collectively referred to as (p)ppGpp) are critical for this process. In Gram-positive bacteria, a main function of (p)ppGpp is to limit cellular levels of GTP, one consequence of which is reduced transcription of genes that require GTP as the initiating nucleotide, such as rRNA genes. In Streptomycetes, the stringent response is also linked to complex morphological differentiation and to production of secondary metabolites, including antibiotics. These processes are also influenced by the second messenger c-di-GMP. Since GTP is a substrate for both (p)ppGpp and c-di-GMP, a finely tuned regulation of cellular GTP levels is required to ensure adequate synthesis of these guanosine derivatives. Here, we discuss mechanisms that operate to control guanosine metabolism and how they impinge on the production of antibiotics in Streptomyces species. Keywords: c-di-GMP; guanosine and (p)ppGpp; purine salvage; secondary metabolism; Streptomycetes; stringent response 1. Introduction Bacteria experience constant challenges, either in the environment or when infecting a host. They utilize various mechanisms to survive such stresses, which may include changes in temperature, pH, or oxygen content as well as limited access to carbon or nitrogen sources. -

Quantification of Guanosine Triphosphate and Tetraphosphate In
Quantification of guanosine triphosphate and tetraphosphate in plants and algae using stable isotope-labelled internal standards Julia Bartoli, Sylvie Citerne, Gregory Mouille, Emmanuelle Bouveret, Ben Field To cite this version: Julia Bartoli, Sylvie Citerne, Gregory Mouille, Emmanuelle Bouveret, Ben Field. Quantification of guanosine triphosphate and tetraphosphate in plants and algae using stable isotope-labelled internal standards. Talanta, Elsevier, 2020, 219, pp.121261. 10.1016/j.talanta.2020.121261. cea-02961710 HAL Id: cea-02961710 https://hal-cea.archives-ouvertes.fr/cea-02961710 Submitted on 19 Nov 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Quantification of guanosine triphosphate and tetraphosphate in plants and algae using stable isotope- labelled internal standards. Julia Bartoli1, Sylvie Citerne2, Gregory Mouille2, Emmanuelle Bouveret3 and Ben Field4* 1Aix Marseille Univ CNRS, LISM, UMR 7255, IMM FR 3479, 31 Chemin Joseph Aiguier, 13009, Marseille, France 2 Institut Jean-Pierre Bourgin, INRAE, AgroParisTech, Université Paris-Saclay, -

Guanosine Pentaphosphate Phosphohydrolase of Escherichia Coli Is a Long-Chain Exopolyphosphatase J
Proc. Natl. Acad. Sci. USA Vol. 90, pp. 7029-7033, August 1993 Biochemistry Guanosine pentaphosphate phosphohydrolase of Escherichia coli is a long-chain exopolyphosphatase J. D. KEASLING*, LEROY BERTSCHt, AND ARTHUR KORNBERGtI *Department of Chemical Engineering, University of California, Berkeley, CA 94720-9989; and tDepartment of Biochemistry, Stanford University School of Medicine, Stanford, CA 94305-5307 Contributed by Arthur Kornberg, April 14, 1993 ABSTRACT An exopolyphosphatase [exopoly(P)ase; EC MATERIALS AND METHODS 3.6.1.11] activity has recently been purified to homogeneity from a mutant strain of Escherichia coi which lacks the Reagents and Proteins. Sources were as follows: ATP, principal exopoly(P)ase. The second exopoly(P)ase has now ADP, nonradiolabeled nucleotides, poly(P)s, bovine serum been identified as guanosine pentaphosphate phosphohydro- albumin, and ovalbumin from Sigma; [y-32P]ATP at 6000 lase (GPP; EC 3.6.1.40) by three lines of evidence: (i) the Ci/mmol (1 Ci = 37 GBq) and [y-32P]GTP at 6000 Ci/mmol sequences of five btptic digestion fragments of the purified from ICN; Q-Sepharose fast flow, catalase, aldolase, Super- protein are found in the translated gppA gene, (u) the size ofthe ose-12 fast protein liquid chromatography (FPLC) column, protein (100 kDa) agrees with published values for GPP, and and Chromatofocusing column and reagents from Pharmacia (iu) the ratio of exopoly(P)ase activity to GPP activity remains LKB; DEAE-Fractogel, Pll phosphocellulose, and DE52 constant throughout a 300-fold purification in the last steps of DEAE-cellulose from Whatman; protein standards for SDS/ the procedure. -

Insights Into Anaerobic Degradation of Benzene and Naphthalene
TECHNISCHE UNIVERSITÄT MÜNCHEN Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt Lehrstuhl für Siedlungswasserwirtschaft Insights into anaerobic degradation of benzene and naphthalene Xiyang Dong Vollständiger Abdruck der von der Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt der Technischen Universität München zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften genehmigten Dissertation. Vorsitzende(r): Prof. Dr. Wolfgang Liebl Prüfer der Dissertation: 1. Priv.-Doz. Dr. Tillmann Lueders 2. Prof. Dr. Siegfried Scherer 3. Prof. Dr. Rainer Meckenstock Die Dissertation wurde am 03.07.2017 bei der Technischen Universität München eingereicht und durch die Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt am 18.08.2017 angenommen. 天道酬勤 God rewards those who work hard Abstract Aromatic hydrocarbons, e.g. benzene and naphthalene, have toxic, mutagenic and/or carcinogenic properties. Fortunately, these compounds can be degraded in environmental systems by indigenous microorganisms. Especially under anaerobic conditions, the physiology and ecology of the microbes involved are still poorly understood. In this thesis, important knowledge gaps are addressed in this field using microbiological approaches combined with “omics tools” (metagenomics and metaproteomics). A novel “reverse stable isotope labelling” approach is also introduced to investigate biodegradation activities. Firstly, the enrichment culture BPL was studied, which can degrade benzene coupled with sulfate reduction. It is dominated by an organism of the genus Pelotomaculum. Members of this genus are usually known to be fermenters, undergoing syntrophy with anaerobic respiring microorganisms or methanogens. It remains unclear if Pelotomaculum identified here (namely, Pelotomaculum candidate BPL) could perform both benzene degradation and sulfate reduction. By using a metagenomic approach, a high-quality genome was reconstructed for it. -

Arxiv:Q-Bio.QM/0511042 V2 25 Nov 2005 Nomto Ae Lseig Upeetr Material Supplementary Clustering: Based Information .THE I
Information based clustering: Supplementary material Noam Slonim, Gurinder Singh Atwal, Gaˇsper Tkaˇcik, and William Bialek Joseph Henry Laboratories of Physics, and Lewis–Sigler Institute for Integrative Genomics Princeton University, Princeton, New Jersey 08544 USA (Dated: December 4, 2005) This technical report provides the supplementary material for a paper entitled “Information based clustering,” to appear shortly in Proceedings of the National Academy of Sciences (USA). In Section I we present in detail the iterative clustering algorithm used in our experiments and in Section II we describe the validation scheme used to determine the statistical significance of our results. Then in subsequent sections we provide all the experimental results for three very different applications: the response of gene expression in yeast to different forms of environmental stress, the dynamics of stock prices in the Standard and Poor’s 500, and viewer ratings of popular movies. In particular, we highlight some of the results that seem to deserve special attention. All the experimental results and relevant code, including a freely available web application, can be found at http://www.genomics.princeton.edu/biophysics-theory . Contents many choices at different levels of the analysis. In recent work we suggest that some generality can be achieved I. The Iclust algorithm 1 through the use of information theory (1). Here we re- view this formulation briefly and then proceed to the II. Evaluating clusters’ coherence 3 technical details of its implementation that were left out III. First application: The yeast ESR data 3 of Ref (1). A. Description of the data 3 We formulate clustering as a tradeoff between maxi- B. -
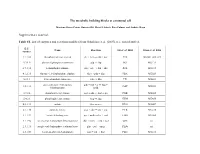
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

334429671.Pdf
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by DSpace at Tartu University Library Talanta 205 (2019) 120161 Contents lists available at ScienceDirect Talanta journal homepage: www.elsevier.com/locate/talanta Analysis of nucleotide pools in bacteria using HPLC-MS in HILIC mode T Eva Zborníkováa,b, Zdeněk Knejzlíka, Vasili Hauryliukc,d,e, Libor Krásnýf, Dominik Rejmana,* a Institute of Organic Chemistry and Biochemistry, Academy of Sciences of the Czech Republic, v.v.i., Flemingovonam. 2, CZ-166 10, Prague 6, Czech Republic b Charles University, Faculty of Science, Department of Analytical Chemistry, Hlavova 8, 128 43, Prague 2, Czech Republic c Department of Molecular Biology, Umeå University, Building 6K, 6L University Hospital Area, SE-901 87, Umeå, Sweden d Laboratory for Molecular Infection Medicine Sweden (MIMS), Umeå University, Building 6K and 6L, University Hospital Area, 90187, Umeå, Sweden e University of Tartu, Institute of Technology, 50411, Tartu, Estonia f Institute of Microbiology, Czech Academy of Sciences v.v.i., Vídeňská 1083, 142 20, Prague 4, Czech Republic ARTICLE INFO ABSTRACT Keywords: Nucleotides, nucleosides and their derivatives are present in all cells at varying concentrations that change with Nucleotide the nutritional, and energetic status of the cell. Precise measurement of the concentrations of these molecules is HPLC-MS instrumental for understanding their regulatory effects. Such measurement is challenging due to the inherent HILIC instability of these molecules and, despite many decades of research, the reported values differ widely. Here, we ppGpp present a comprehensive and easy-to-use approach for determination of the intracellular concentrations of > 25 Stringent response target molecular species. -

(P)Ppgpp in Plants and Algae Ben Field
Green magic: regulation of the chloroplast stress response by (p)ppGpp in plants and algae Ben Field To cite this version: Ben Field. Green magic: regulation of the chloroplast stress response by (p)ppGpp in plants and algae. Journal of Experimental Botany, Oxford University Press (OUP), 2018, 69 (11), pp.2797-2807. 10.1093/jxb/erx485. hal-01855838 HAL Id: hal-01855838 https://hal-amu.archives-ouvertes.fr/hal-01855838 Submitted on 8 Aug 2018 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Green magic: regulation of the chloroplast stress response by (p)ppGpp in plants and algae Ben Field Aix Marseille Univ, CEA, CNRS, France Correspondence: [email protected] Abstract The hyperphosphorylated nucleotides guanosine pentaphosphate and tetraphosphate [together referred to as (p)ppGpp, or ‘magic spot’] orchestrate a signalling cascade in bacteria that controls growth under optimal conditions and in response to environmental stress. (p)ppGpp is also found in the chloroplasts of plants and algae where it has also been shown to accumulate in response to abiotic stress. Recent studies suggest that (p)ppGpp is a potent inhibitor of chloroplast gene expression in vivo, and is a significant regulator of chloroplast function that can influence both the growth and the development of plants. -

The Nucleotide Pgpp Acts As a Third Alarmone in Bacillus, with Functions Distinct from Those of (P)Ppgpp
ARTICLE https://doi.org/10.1038/s41467-020-19166-1 OPEN The nucleotide pGpp acts as a third alarmone in Bacillus, with functions distinct from those of (p)ppGpp Jin Yang 1, Brent W. Anderson 1, Asan Turdiev2, Husan Turdiev2, David M. Stevenson1, ✉ ✉ Daniel Amador-Noguez 1, Vincent T. Lee 2 & Jue D. Wang 1 1234567890():,; The alarmone nucleotides guanosine tetraphosphate and pentaphosphate, commonly refer- red to as (p)ppGpp, regulate bacterial responses to nutritional and other stresses. There is evidence for potential existence of a third alarmone, guanosine-5′-monophosphate-3′- diphosphate (pGpp), with less-clear functions. Here, we demonstrate the presence of pGpp in bacterial cells, and perform a comprehensive screening to identify proteins that interact respectively with pGpp, ppGpp and pppGpp in Bacillus species. Both ppGpp and pppGpp interact with proteins involved in inhibition of purine nucleotide biosynthesis and with GTPases that control ribosome assembly or activity. By contrast, pGpp interacts with purine biosynthesis proteins but not with the GTPases. In addition, we show that hydrolase NahA (also known as YvcI) efficiently produces pGpp by hydrolyzing (p)ppGpp, thus modulating alarmone composition and function. Deletion of nahA leads to reduction of pGpp levels, increased (p)ppGpp levels, slower growth recovery from nutrient downshift, and loss of competitive fitness. Our results support the existence and physiological relevance of pGpp as a third alarmone, with functions that can be distinct from those of (p)ppGpp. 1 Department of Bacteriology, University of Wisconsin, Madison, WI 53706, USA. 2 Department of Cell Biology and Molecular Genetics, University of ✉ Maryland, College Park, MD, USA. -

Occurrence of the Regulatory Nucleotides Ppgpp and Pppgpp Following Induction of the Stringent Response in Staphylococci
JOURNAL OF BACTERIOLOGY, Sept. 1995, p. 5161–5165 Vol. 177, No. 17 0021-9193/95/$04.0010 Copyright q 1995, American Society for Microbiology Occurrence of the Regulatory Nucleotides ppGpp and pppGpp following Induction of the Stringent Response in Staphylococci R. CASSELS,* B. OLIVA,† AND D. KNOWLES Department of Microbial Metabolism and Biochemistry, SmithKline Beecham Pharmaceuticals, Brockham Park, Betchworth, Surrey RH3 7AJ, United Kingdom Received 6 February 1995/Accepted 27 June 1995 The stringent response in Escherichia coli and many other organisms is regulated by the nucleotides ppGpp and pppGpp. We show here for the first time that at least six staphylococcal species also synthesize ppGpp and pppGpp upon induction of the stringent response by mupirocin. Spots corresponding to ppGpp and pppGpp on thin-layer chromatograms suggest that pppGpp is the principal regulatory nucleotide synthesized by staphylococci in response to mupirocin, rather than ppGpp as in E. coli. Bacteria adapt to amino acid or carbon source insufficiency and higher organisms after amino acid starvation or induction by a complex series of regulatory events known as the stringent by a variety of antibiotics (7, 18, 26). However, in halobacteria response (reviewed in reference 4). In Escherichia coli, for (25) and streptococci (21), stringency is not necessarily coupled example, amino acid starvation leads to an accumulation of with (p)ppGpp production. Thus, in Streptococcus pyogenes uncharged cognate tRNA. When the ratio of charged to un- and Streptococcus rattus, chloramphenicol reverses the inhibi- charged tRNA falls below a critical threshold (24), occupation tion of RNA synthesis caused by mupirocin, but there is no of the vacant mRNA codon at the ribosomal A site by un- generation of ppGpp in the presence of mupirocin alone.