Correlation Between the Oral Microbiome and Brain Resting State Connectivity in Smokers
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Mysterious Orphans of Mycoplasmataceae
The mysterious orphans of Mycoplasmataceae Tatiana V. Tatarinova1,2*, Inna Lysnyansky3, Yuri V. Nikolsky4,5,6, and Alexander Bolshoy7* 1 Children’s Hospital Los Angeles, Keck School of Medicine, University of Southern California, Los Angeles, 90027, California, USA 2 Spatial Science Institute, University of Southern California, Los Angeles, 90089, California, USA 3 Mycoplasma Unit, Division of Avian and Aquatic Diseases, Kimron Veterinary Institute, POB 12, Beit Dagan, 50250, Israel 4 School of Systems Biology, George Mason University, 10900 University Blvd, MSN 5B3, Manassas, VA 20110, USA 5 Biomedical Cluster, Skolkovo Foundation, 4 Lugovaya str., Skolkovo Innovation Centre, Mozhajskij region, Moscow, 143026, Russian Federation 6 Vavilov Institute of General Genetics, Moscow, Russian Federation 7 Department of Evolutionary and Environmental Biology and Institute of Evolution, University of Haifa, Israel 1,2 [email protected] 3 [email protected] 4-6 [email protected] 7 [email protected] 1 Abstract Background: The length of a protein sequence is largely determined by its function, i.e. each functional group is associated with an optimal size. However, comparative genomics revealed that proteins’ length may be affected by additional factors. In 2002 it was shown that in bacterium Escherichia coli and the archaeon Archaeoglobus fulgidus, protein sequences with no homologs are, on average, shorter than those with homologs [1]. Most experts now agree that the length distributions are distinctly different between protein sequences with and without homologs in bacterial and archaeal genomes. In this study, we examine this postulate by a comprehensive analysis of all annotated prokaryotic genomes and focusing on certain exceptions. -

A Case Study of Salivary Microbiome in Smokers and Non-Smokers in Hungary: Analysis by Shotgun Metagenome Sequencing
Journal of Oral Microbiology ISSN: (Print) (Online) Journal homepage: https://www.tandfonline.com/loi/zjom20 A case study of salivary microbiome in smokers and non-smokers in Hungary: analysis by shotgun metagenome sequencing Roland Wirth , Gergely Maróti , Róbert Mihók , Donát Simon-Fiala , Márk Antal , Bernadett Pap , Anett Demcsák , Janos Minarovits & Kornél L. Kovács To cite this article: Roland Wirth , Gergely Maróti , Róbert Mihók , Donát Simon-Fiala , Márk Antal , Bernadett Pap , Anett Demcsák , Janos Minarovits & Kornél L. Kovács (2020) A case study of salivary microbiome in smokers and non-smokers in Hungary: analysis by shotgun metagenome sequencing, Journal of Oral Microbiology, 12:1, 1773067, DOI: 10.1080/20002297.2020.1773067 To link to this article: https://doi.org/10.1080/20002297.2020.1773067 © 2020 The Author(s). Published by Informa View supplementary material UK Limited, trading as Taylor & Francis Group. Published online: 07 Jun 2020. Submit your article to this journal Article views: 84 View related articles View Crossmark data Full Terms & Conditions of access and use can be found at https://www.tandfonline.com/action/journalInformation?journalCode=zjom20 JOURNAL OF ORAL MICROBIOLOGY 2020, VOL. 12, 1773067 https://doi.org/10.1080/20002297.2020.1773067 A case study of salivary microbiome in smokers and non-smokers in Hungary: analysis by shotgun metagenome sequencing Roland Wirtha, Gergely Marótib, Róbert Mihókc, Donát Simon-Fialac, Márk Antalc, Bernadett Papb, Anett Demcsákd, Janos Minarovitsd and Kornél L. Kovács -

Product Sheet Info
Product Information Sheet for HM-13 Oribacterium sinus, Strain F0268 2% yeast extract (ATCC medium 1834) or equivalent Wilkins-Chalgren anaerobe agar with 5% defibrinated sheep blood or equivalent Catalog No. HM-13 Incubation: Temperature: 37°C For research use only. Not for human use. Atmosphere: Anaerobic (80% N2:10% CO2:10% H2) Propagation: Contributor: 1. Keep vial frozen until ready for use, then thaw. Jacques Izard, Assistant Member of the Staff, Department of 2. Transfer the entire thawed aliquot into a single tube of Molecular Genetics, The Forsyth Institute, Boston, broth. Massachusetts 3. Use several drops of the suspension to inoculate an agar slant and/or plate. Manufacturer: 4. Incubate the tube, slant and/or plate at 37°C for 48 to BEI Resources 72 hours. Product Description: Citation: Bacteria Classification: Lachnospiraceae, Oribacterium Acknowledgment for publications should read “The following Species: Oribacterium sinus reagent was obtained through BEI Resources, NIAID, NIH as Strain: F0268 part of the Human Microbiome Project: Oribacterium sinus, Original Source: Oribacterium sinus (O. sinus), strain F0268 Strain F0268, HM-13.” was isolated in June 1980 from the subgingival plaque of a 27-year-old white female patient with severe periodontitis in Biosafety Level: 1 the United States.1,2 Appropriate safety procedures should always be used with Comments: O. sinus, strain F0268 (HMP ID 6123) is a this material. Laboratory safety is discussed in the following reference genome for The Human Microbiome Project publication: U.S. Department of Health and Human Services, (HMP). HMP is an initiative to identify and characterize Public Health Service, Centers for Disease Control and human microbial flora. -

Bacterial Diversity and Functional Analysis of Severe Early Childhood
www.nature.com/scientificreports OPEN Bacterial diversity and functional analysis of severe early childhood caries and recurrence in India Balakrishnan Kalpana1,3, Puniethaa Prabhu3, Ashaq Hussain Bhat3, Arunsaikiran Senthilkumar3, Raj Pranap Arun1, Sharath Asokan4, Sachin S. Gunthe2 & Rama S. Verma1,5* Dental caries is the most prevalent oral disease afecting nearly 70% of children in India and elsewhere. Micro-ecological niche based acidifcation due to dysbiosis in oral microbiome are crucial for caries onset and progression. Here we report the tooth bacteriome diversity compared in Indian children with caries free (CF), severe early childhood caries (SC) and recurrent caries (RC). High quality V3–V4 amplicon sequencing revealed that SC exhibited high bacterial diversity with unique combination and interrelationship. Gracillibacteria_GN02 and TM7 were unique in CF and SC respectively, while Bacteroidetes, Fusobacteria were signifcantly high in RC. Interestingly, we found Streptococcus oralis subsp. tigurinus clade 071 in all groups with signifcant abundance in SC and RC. Positive correlation between low and high abundant bacteria as well as with TCS, PTS and ABC transporters were seen from co-occurrence network analysis. This could lead to persistence of SC niche resulting in RC. Comparative in vitro assessment of bioflm formation showed that the standard culture of S. oralis and its phylogenetically similar clinical isolates showed profound bioflm formation and augmented the growth and enhanced bioflm formation in S. mutans in both dual and multispecies cultures. Interaction among more than 700 species of microbiota under diferent micro-ecological niches of the human oral cavity1,2 acts as a primary defense against various pathogens. Tis has been observed to play a signifcant role in child’s oral and general health. -

Characterization of Antibiotic Resistance Genes in the Species of the Rumen Microbiota
ARTICLE https://doi.org/10.1038/s41467-019-13118-0 OPEN Characterization of antibiotic resistance genes in the species of the rumen microbiota Yasmin Neves Vieira Sabino1, Mateus Ferreira Santana1, Linda Boniface Oyama2, Fernanda Godoy Santos2, Ana Júlia Silva Moreira1, Sharon Ann Huws2* & Hilário Cuquetto Mantovani 1* Infections caused by multidrug resistant bacteria represent a therapeutic challenge both in clinical settings and in livestock production, but the prevalence of antibiotic resistance genes 1234567890():,; among the species of bacteria that colonize the gastrointestinal tract of ruminants is not well characterized. Here, we investigate the resistome of 435 ruminal microbial genomes in silico and confirm representative phenotypes in vitro. We find a high abundance of genes encoding tetracycline resistance and evidence that the tet(W) gene is under positive selective pres- sure. Our findings reveal that tet(W) is located in a novel integrative and conjugative element in several ruminal bacterial genomes. Analyses of rumen microbial metatranscriptomes confirm the expression of the most abundant antibiotic resistance genes. Our data provide insight into antibiotic resistange gene profiles of the main species of ruminal bacteria and reveal the potential role of mobile genetic elements in shaping the resistome of the rumen microbiome, with implications for human and animal health. 1 Departamento de Microbiologia, Universidade Federal de Viçosa, Viçosa, Minas Gerais, Brazil. 2 Institute for Global Food Security, School of Biological -

Oral Microbiome Composition Reflects Prospective Risk for Esophageal Cancers
Cancer Prevention and Epidemiology Research Oral Microbiome Composition Reflects Prospective Risk for Esophageal Cancers Brandilyn A. Peters1, Jing Wu1,2, Zhiheng Pei2,3,4, Liying Yang5, Mark P. Purdue6, Neal D. Freedman6, Eric J. Jacobs7, Susan M. Gapstur7, Richard B. Hayes1,2, and Jiyoung Ahn1,2 Abstract Bacteria may play a role in esophageal adenocarcinoma (EAC) conditional logistic regression adjusting for BMI, smoking, and and esophageal squamous cell carcinoma (ESCC), although alcohol. We found the periodontal pathogen Tannerella forsythia evidence is limited to cross-sectional studies. In this study, we to be associated with higher risk of EAC. Furthermore, we found examined the relationship of oral microbiota with EAC and ESCC that depletion of the commensal genus Neisseria and the species risk in a prospective study nested in two cohorts. Oral bacteria Streptococcus pneumoniae was associated with lower EAC risk. were assessed using 16S rRNA gene sequencing in prediagnostic Bacterial biosynthesis of carotenoids was also associated with mouthwash samples from n ¼ 81/160 EAC and n ¼ 25/50 ESCC protection against EAC. Finally, the abundance of the periodontal cases/matched controls. Findings were largely consistent across pathogen Porphyromonas gingivalis trended with higher risk of ESCC. both cohorts. Metagenome content was predicted using PiCRUST. Overall, our findings have potential implications for the early We examined associations between centered log-ratio trans- detection and prevention of EAC and ESCC. Cancer Res; 77(23); formed taxon or functional pathway abundances and risk using 6777–87. Ó2017 AACR. Introduction intake, and smoking for EAC, and alcohol drinking, low fruit/ vegetable intake, and smoking for ESCC (4), but the etiology Esophageal cancer is the eighth most common cancer and sixth of these diseases cannot be fully explained by these factors. -

Microbiomes in Supragingival Bio Lms and Saliva of Adolescent Patients
Microbiomes in Supragingival Biolms and Saliva of Adolescent Patients with Induced and Naturally Occurring Gingivitis Relative to Gingival Health Roland Wirth Szegedi Tudomanyegyetem Gergely Maróti Szegedi Biológiai Központ Lídia Lipták Szegedi Tudomanyegyetem Mónika Mester Szegedi Tudomanyegyetem Alaa Al Ayoubi Szegedi Tudomanyegyetem Bernadett Pap Magyar Tudomanyos Akademia Szegedi Biologiai Kozpont Melinda Madléna Szegedi Tudomanyegyetem Janos Minarovits Szegedi Tudomanyegyetem Kornél L. Kovács ( [email protected] ) Szegedi Tudomanyegyetem Biologia Intezet https://orcid.org/0000-0002-3926-0497 Research Keywords: Gingivitis, Microbiomes, gingival health, saliva Posted Date: August 6th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-45630/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/27 Abstract Background: Comparison of the microbiomes in supragingival biolm and saliva samples collected from juvenile patients developing induced or spontaneous gingivitis with healthy controls. Results: 36 supragingival biolm samples from 9 adolescent gingivitis patients wearing orthodontic appliances (induced gingivitis), 40 supragingival plaques from 10 patients having spontaneous gingivitis, and 36 control samples from 9 individuals without gingivitis in the same age group were analyzed by 16S rRNA gene amplicon sequencing. Salivary microbiomes of the same persons were characterized by shotgun metagenome sequencing to compare the sessile, i.e. biolm immobilized communities with planktonic microbiota. The amplicon and whole genome data sets were scrutinized using bioinformatics workows designed to minimize systemic biases. RDP and RefSeq reference databases were compared in the identication of microbiome members. The composition and diversity of bacterial communities did not differ extensively between the two groups of gingivitis patients and controls. -
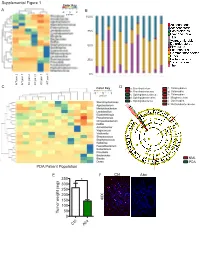
Mental Figure 1 Color Key a -2 0 2 B Z-Score 100%
Supplemental Figure 1 Color Key A -2 0 2 B z-score 100% 75% 50% 25% 0% KC pan 1 WT pan 3 WT KC pan 3 WT pan 2 WT pan 1 WT KC pan 2 C Color Key D a: Brevibacterium f: Chlamydiales b: Brevibacteriaceae g: Chlamydiia -3 0 3 z-score c: Sphingobacteriaceae h: Chlamydiae d: Sphingobacteriales i: Mogibacterium e: Sphingobacteriia j: Oscillospira k: Methylobacteriaceae NML Control Microb.-entrained MΦ PDA PDA Patient Population Control Microb.-entrained MΦ + Myd88i E F Ctrl Abx 350 * 300 250 200 150 40X 100 Tumor weight (mg) 50 0 x Ctrl Ab Supplemental Figure 2 A KC WT B ** * Actinobacteria * ** Bacteroidetes Cyanobacteria Deferribacteres * Firmicutes Proteobacteria % Relative abundance TM7 Others Time(wks) 3 9 13 16 20 24 28 32 36 3 9 13 16 20 24 28 32 36 Alpha Diversity Measure C E 60 KC WT 40 20 B. pseudolongum B. animalis 60 5 KC WT 0 B. adolescentis 40% Rel. abundance 3 9 13 16 20 24 28 32 36 3 9 13 16 20 24 28 32 36 20 Age (weeks) B. pseudolongum B. animalis 5 0 B. adolescentis % Rel. abundance 3 9 13 16 20 24 28 32 36 3 9 13 16 20 24 28 32 36 F Age (weeks) Week 3 Week 9 Week 13 p=0.678 p=0.02 p=0.385 Time(wks) 3 9 24 20 13 16 D 28 32 36 Week 13 KC WT Firmicutes; Ruminococcus Firmicutes; Dehalobacterium Alpha Diversity Measure Firmicutes; Oscillospira Bacteroidates; Odoribacter Axis.2 [12.7%] Actinobacteria; Bifidobacterium Axis.2 [23.8%] Axis.2 [24.7%] Week 16 Bacteroidetes; Bacteroidales Axis.1 [80.8%] Axis.1 [65.4%] Axis.1 [49.6%] Actinobacteria; Bifidobacterium Week 16 Week 20 Week 24 Week 20 p=0.339 p=0.036 p=0.021 Firmicutes; Dehalobacterium -

Exploring the Cockatiel (Nymphicus Hollandicus) Fecal Microbiome, Bacterial Inhabitants of a Worldwide Pet
Exploring the cockatiel (Nymphicus hollandicus) fecal microbiome, bacterial inhabitants of a worldwide pet Luis David Alcaraz1, Apolinar M. Hernández2 and Mariana Peimbert2 1 Laboratorio Nacional de Ciencias de la Sostenibilidad, Instituto de Ecología, Universidad Nacional Autonóma de México, Mexico City, Mexico 2 Departamento de Ciencias Naturales, Unidad Cuajimalpa, Universidad Autónoma Metropolitana, Mexico City, Mexico ABSTRACT Background. Cockatiels (Nymphicus hollandicus) were originally endemic to Australia; now they are popular pets with a global distribution. It is now possible to conduct detailed molecular studies on cultivable and uncultivable bacteria that are part of the intestinal microbiome of healthy animals. These studies show that bacteria are an essential part of the metabolic capacity of animals. There are few studies on bird microbiomes and, to the best of our knowledge, this is the first report on the cockatiel microbiome. Methods. In this paper, we analyzed the gut microbiome from fecal samples of three healthy adult cockatiels by massive sequencing of the 16S rRNA gene. Additionally, we compared the cockatiel fecal microbiomes with those of other bird species, including poultry and wild birds. Results. The vast majority of the bacteria found in cockatiels were Firmicutes, while Proteobacteria and Bacteroidetes were poorly represented. A total of 19,280 different OTUs were detected, of which 8,072 belonged to the Erysipelotrichaceae family. Discussion. It is relevant to study cockatiel the microbiomes of cockatiels owing to their wide geographic distribution and close human contact. This study serves as a reference for cockatiel bacterial diversity. Despite the large OTU numbers, the diversity is not even Submitted 14 July 2016 Accepted 28 November 2016 and is dominated by Firmicutes of the Erysipelotrichaceae family. -

To Split Or Not to Split: an Opinion on Dividing the Genus Burkholderia
Ann Microbiol (2016) 66:1303–1314 DOI 10.1007/s13213-015-1183-1 REVIEW ARTICLE To split or not to split: an opinion on dividing the genus Burkholderia Paulina Estrada-de los Santos 1 & Fernando Uriel Rojas-Rojas1 & Erika Yanet Tapia-García1 & María Soledad Vásquez-Murrieta1 & Ann M. Hirsch2,3 Received: 27 April 2015 /Accepted: 24 November 2015 /Published online: 23 December 2015 # Springer-Verlag Berlin Heidelberg and the University of Milan 2015 Abstract The genus Burkholderia is a large group of species features, and their relationship with plants as either associative of bacteria that inhabit a wide range of environments. We nitrogen-fixers or legume-nodulating/nitrogen-fixing bacteria. previously recommended, based on multilocus sequence anal- We also propose that a concerted and coordinated effort be ysis, that the genus be separated into two distinct groups—one made by researchers on Burkholderia to determine if a defin- that consists predominantly of human, plant, and animal path- itive taxonomic split of this very large genus is justified, es- ogens, including several opportunistic pathogens, and a sec- pecially now as we describe here for the first time intermediate ond, much larger group of species comprising plant-associated groups based upon their 16S rRNA sequences. We need to beneficial and environmental species that are primarily known learn more about the plant-associated Burkholderia strains not to be pathogenic. This second group of species is found regarding their potential for pathogenicity, especially in those mainly in soils, frequently in association with plants as plant strains intermediate between the two groups, and to discover growth-promoting bacteria. -

Prevalence of Ureaplasma Urealyticum, Mycoplasma Hominis and Chlamydia Trachomatis in Patients with Uncomplicated Recurrent Urin
Nephrology and Renal Diseases Research Article ISSN: 2399-908X Prevalence of Ureaplasma urealyticum, Mycoplasma hominis and Chlamydia trachomatis in patients with uncomplicated recurrent urinary tract infections Jadranka Vlasic-Matas1*, Hrvoje Raos2, Marijana Vuckovic2, Stjepan Radic2 and Vesna Capkun3 1Polyclinic Nephrology Department, Split, Croatia 2School of Medicine, University of Split, Split, Croatia 3Department of Nuclear Medicine, Split University Hospital Center, Split, Croatia Abstract Aim: To assess the prevalence of Ureaplasma urealyticum, Mycoplasma hominis and Chlamydia trachomatis in patients with chronic urinary tract infections (UTIs) and its correlation with leukocyturia and symptoms. Methods: The study included 220 patients (130 women and 90 men) presenting with chronic voiding symptoms and sterile leukocyturia. Urine, urethral swabs and cervical swabs (for women patients) were taken to determine the presence of these pathogens. Patients were treated by tetracycline and followed up three and six months after initial therapy. Results: In 186 (85%) out of 220 patients, U. urealyticum was found, while C. trachomatis was present in 34 patients (15%). In majority of female patients (112 out of 130; 86%) U. urealyticum was found. In addition to ureaplasma, in eight patients M. hominis was found. C. trachomatis was identified in 18 female patients (14%). In 74 out of 90 (82%) male patients U. urealyticum was detected while in six of them M. hominis was also found. C. trachomatis was identified in 16 male patients (18%). U. urealyticum was significantly related to leukocyturia, as opposed to C. trachomatis (p<0,001). Women had more frequent symptomatology (p = 0,015) and higer leukocyturia (p<0.001). Conclusion: Leukocyturia is more common find in U. -

Avian Mycoplasmosis (Mycoplasma Gallisepticum)
Avian Importance Mycoplasma gallisepticum is an economically significant pathogen that can Mycoplasmosis cause significant losses in chickens, turkeys and game birds from chronic respiratory disease, reduced feed efficiency, decreased growth and lower egg (Mycoplasma production. In addition, the carcasses of birds sent to slaughter may be downgraded. Many countries with modern poultry operations have eradicated this organism from gallisepticum) commercial chicken and turkey breeding flocks; however, it can still be an issue in other poultry operations, such as multi-age layer flocks, game bird raising facilities Pleuropneumonia–like Organism and backyard birds. Since 1994, conjunctivitis caused by one lineage of M. (PPLO) Infection, Chronic gallisepticum has become a significant disease in wild birds in North America. Respiratory Disease of Chickens, Although other wild birds can be affected, the major impact has been on house Infectious Sinusitis of Turkeys, finches (Carpodacus mexicanus), which have experienced major population House Finch Conjunctivitis declines in some areas. Etiology Last Updated: November 2018 Mycoplasma gallisepticum, a member of the family Mycoplasmataceae (class Mollicutes, order Mycoplasmatales), is one of the most important agents of mycoplasmosis in terrestrial poultry. There are multiple strains of this organism, which can differ in virulence and may also have different host preferences. The house finch lineage is a distinct lineage that has diverged significantly from poultry strains and has become established