LETM1 Gene Leucine Zipper and EF-Hand Containing Transmembrane Protein 1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Rise and Fall of the Bovine Corpus Luteum
University of Nebraska Medical Center DigitalCommons@UNMC Theses & Dissertations Graduate Studies Spring 5-6-2017 The Rise and Fall of the Bovine Corpus Luteum Heather Talbott University of Nebraska Medical Center Follow this and additional works at: https://digitalcommons.unmc.edu/etd Part of the Biochemistry Commons, Molecular Biology Commons, and the Obstetrics and Gynecology Commons Recommended Citation Talbott, Heather, "The Rise and Fall of the Bovine Corpus Luteum" (2017). Theses & Dissertations. 207. https://digitalcommons.unmc.edu/etd/207 This Dissertation is brought to you for free and open access by the Graduate Studies at DigitalCommons@UNMC. It has been accepted for inclusion in Theses & Dissertations by an authorized administrator of DigitalCommons@UNMC. For more information, please contact [email protected]. THE RISE AND FALL OF THE BOVINE CORPUS LUTEUM by Heather Talbott A DISSERTATION Presented to the Faculty of the University of Nebraska Graduate College in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Biochemistry and Molecular Biology Graduate Program Under the Supervision of Professor John S. Davis University of Nebraska Medical Center Omaha, Nebraska May, 2017 Supervisory Committee: Carol A. Casey, Ph.D. Andrea S. Cupp, Ph.D. Parmender P. Mehta, Ph.D. Justin L. Mott, Ph.D. i ACKNOWLEDGEMENTS This dissertation was supported by the Agriculture and Food Research Initiative from the USDA National Institute of Food and Agriculture (NIFA) Pre-doctoral award; University of Nebraska Medical Center Graduate Student Assistantship; University of Nebraska Medical Center Exceptional Incoming Graduate Student Award; the VA Nebraska-Western Iowa Health Care System Department of Veterans Affairs; and The Olson Center for Women’s Health, Department of Obstetrics and Gynecology, Nebraska Medical Center. -

What Are Their Roles in Mitochondrial Protein Synthesis?
Characterisation of human mtRF1 and C12orf65: What are their roles in mitochondrial protein synthesis? Aleksandra Pajak M.Res Thesis submitted to Newcastle University in candidature for the degree of Doctor of Philosophy Newcastle University Faculty of Medical Sciences Institute for Ageing and Health Mitochondrial Research Group January 2013 Abstract Mitochondria have their own protein synthesis machinery that synthesises the oxidative phosphorylation components encoded by their mtDNA. This translation process consists of four main phases: initiation, elongation, termination and ribosome recycling. Termination and its control have been the least investigated. Recently, however, the termination factor, mtRF1a, has been characterised as sufficient to release all the nascent proteins from the mitoribosome. Furthermore, bioinformatics has identified three additional members of this mitochondrial release factor family namely, mtRF1, C12orf65 and ICT1. The latter is now known to be incorporated into the mitoribosome but its exact function remains unclear. My project has therefore focussed on characterising the remaining two factors; mtRF1 and C12orf65, and investigating their possible involvement in mitochondrial protein synthesis. It has been demonstrated that protein synthesis is not perfect and bacterial ribosomes not infrequently stall during translation. This can result from limiting amounts of charged tRNAs, stable secondary structures, or truncated/degraded transcripts. Ribosome stalling has been shown to cause growth arrest. In order to prevent that and maintain high efficiency of mitochondrial protein synthesis such stalled complexes need to be rapidly recycled. Bacteria have developed at least three distinct mechanisms by which ribosomes can be rescued. Contrastingly, despite the presence of truncated mRNAs in mitochondria, no such quality control mechanisms have been identified in these organelles. -

Evaluation of Cancer-Derived Myocardial Impairments Using a Mouse Model
www.oncotarget.com Oncotarget, 2020, Vol. 11, (No. 41), pp: 3712-3722 Research Paper Evaluation of cancer-derived myocardial impairments using a mouse model Yoshihiro Miyagawa1, Shota Nukaga1,2, Takuya Mori1, Rina Fujiwara-Tani1, Kiyomu Fujii1, Shiori Mori1, Kei Goto1,3, Shingo Kishi1, Takamitsu Sasaki1, Chie Nakashima1, Hitoshi Ohmori1, Isao Kawahara1,2, Yi Luo4 and Hiroki Kuniyasu1 1Department of Molecular Pathology, Nara Medical University, Kashihara, Nara 634-8521, Japan 2Division of Rehabilitation, Hanna Central Hospital, Ikoma, Nara 630-0243, Japan 3Division of Rehabilitation, Hoshida Minami Hospital, Katano, Osaka 576-0022, Japan 4Key Laboratory of Neuroregeneration of Jiangsu and Ministry of Education, Co-Innovation Center of Neuroregeneration, Nantong University, Nantong, Jiangsu Province 226001, China Correspondence to: Yi Luo, email: [email protected] Hiroki Kuniyasu, email: [email protected] Keywords: cachexia; myocardium; atrophy; mitochondria; oxidative stress Received: June 26, 2020 Accepted: September 10, 2020 Published: October 13, 2020 Copyright: © 2020 Miyagawa et al. This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Myocardial damage in cancer patients is emphasized as a cause of death; however, there are not many murine cachexia models to evaluate cancer-derived heart disorder. Using the mouse cachexia model that we established previously, we investigated myocardial damage in tumor-bearing mice. In cachexic mice, decreased heart weight and myocardial volume, and dilated left ventricular lumen, and atrophied cardiomyocytes were noted. The cardiomyocytes also showed accumulated 8-hydroxydeoxyguanosine, decreased leucine zipper and EF-hand- containing transmembrane protein-1, and increased microtubule-associated protein light chain3-II. -

LETM1 Haploinsufficiency Causes Mitochondrial Defects in Cells From
© 2014. Published by The Company of Biologists Ltd | Disease Models & Mechanisms (2014) 7, 535-545 doi:10.1242/dmm.014464 RESEARCH ARTICLE LETM1 haploinsufficiency causes mitochondrial defects in cells from humans with Wolf-Hirschhorn syndrome: implications for dissecting the underlying pathomechanisms in this condition Lesley Hart1,2, Anita Rauch3, Antony M. Carr2, Joris R. Vermeesch4 and Mark O’Driscoll1,* ABSTRACT abnormalities, a characteristic facial dysmorphology, hypotonia, and Wolf-Hirschhorn syndrome (WHS) represents an archetypical epileptic seizures (Hirschhorn and Cooper, 1961; Hirschhorn et al., example of a contiguous gene deletion disorder – a condition 1965; Wolf et al., 1965). The spectrum and severity of these clinical comprising a complex set of developmental phenotypes with a features typically correlate with deletion size (Battaglia et al., 2008; multigenic origin. Epileptic seizures, intellectual disability, growth Maas et al., 2008; Van Buggenhout et al., 2004; Zollino et al., 2000). restriction, motor delay and hypotonia are major co-morbidities in WHS is generally regarded as a multigenic disorder, although two WHS. Haploinsufficiency of LETM1, which encodes a mitochondrial critical regions have been described: WHSCR1 and WHSCR2, for inner-membrane protein functioning in ion transport, has been WHS critical region 1 and 2, respectively (see Fig. 1). These critical proposed as an underlying pathomechanism, principally for seizures regions are based on the demarcation of the minimum region of but also for other core features of WHS, including growth and motor overlap in individuals exhibiting WHS-like phenotypes. WHSCR1 delay. Growing evidence derived from several model organisms incorporates part of the WHS candidate gene WHSC1 and the entire suggests that reduced LETM1 expression is associated with some WHSC2 gene (White et al., 1995; Wright et al., 1997). -

Stríbrná and Julius Lukes Hassan Hashimi, Lindsay Mcdonald, Eva
Bioenergetics: Trypanosome Letm1 Protein Is Essential for Mitochondrial Potassium Homeostasis Hassan Hashimi, Lindsay McDonald, Eva Stríbrná and Julius Lukes J. Biol. Chem. 2013, 288:26914-26925. doi: 10.1074/jbc.M113.495119 originally published online July 26, 2013 Access the most updated version of this article at doi: 10.1074/jbc.M113.495119 Find articles, minireviews, Reflections and Classics on similar topics on the JBC Affinity Sites. Alerts: • When this article is cited • When a correction for this article is posted Click here to choose from all of JBC's e-mail alerts Supplemental material: http://www.jbc.org/content/suppl/2013/07/26/M113.495119.DC1.html This article cites 60 references, 26 of which can be accessed free at http://www.jbc.org/content/288/37/26914.full.html#ref-list-1 Downloaded from http://www.jbc.org/ at Edinburgh University Library on September 13, 2013 THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 288, NO. 37, pp. 26914–26925, September 13, 2013 © 2013 by The American Society for Biochemistry and Molecular Biology, Inc. Published in the U.S.A. Trypanosome Letm1 Protein Is Essential for Mitochondrial Potassium Homeostasis*□S Received for publication, June 19, 2013, and in revised form, July 23, 2013 Published, JBC Papers in Press, July 26, 2013, DOI 10.1074/jbc.M113.495119 Hassan Hashimi‡§1, Lindsay McDonald‡2, Eva Strˇíbrná‡, and Julius Lukesˇ‡§3 From the ‡Institute of Parasitology, Biology Centre, Czech Academy of Sciences and the §Faculty of Science, University of South Bohemia, 370 05 Cˇeské Budeˇjovice (Budweis), Czech Republic Background: Letm1 is a mitochondrial protein attributed disparate roles, including cation/proton antiport and translation. -

Genome-Wide Investigation of Cellular Functions for Trna Nucleus
Genome-wide Investigation of Cellular Functions for tRNA Nucleus- Cytoplasm Trafficking in the Yeast Saccharomyces cerevisiae DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Hui-Yi Chu Graduate Program in Molecular, Cellular and Developmental Biology The Ohio State University 2012 Dissertation Committee: Anita K. Hopper, Advisor Stephen Osmani Kurt Fredrick Jane Jackman Copyright by Hui-Yi Chu 2012 Abstract In eukaryotic cells tRNAs are transcribed in the nucleus and exported to the cytoplasm for their essential role in protein synthesis. This export event was thought to be unidirectional. Surprisingly, several lines of evidence showed that mature cytoplasmic tRNAs shuttle between nucleus and cytoplasm and their distribution is nutrient-dependent. This newly discovered tRNA retrograde process is conserved from yeast to vertebrates. Although how exactly the tRNA nuclear-cytoplasmic trafficking is regulated is still under investigation, previous studies identified several transporters involved in tRNA subcellular dynamics. At least three members of the β-importin family function in tRNA nuclear-cytoplasmic intracellular movement: (1) Los1 functions in both the tRNA primary export and re-export processes; (2) Mtr10, directly or indirectly, is responsible for the constitutive retrograde import of cytoplasmic tRNA to the nucleus; (3) Msn5 functions solely in the re-export process. In this thesis I focus on the physiological role(s) of the tRNA nuclear retrograde pathway. One possibility is that nuclear accumulation of cytoplasmic tRNA serves to modulate translation of particular transcripts. To test this hypothesis, I compared expression profiles from non-translating mRNAs and polyribosome-bound translating mRNAs collected from msn5Δ and mtr10Δ mutants and wild-type cells, in fed or acute amino acid starvation conditions. -
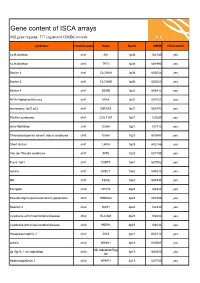
ISCA Disease List
Gene content of ISCA arrays 402 gene regions, 377 registered OMIM records syndrome chromosome Gene Band OMIM ISCA 8x60k 1p36 deletion chr1 SKI 1p36 164780 yes 1p36 deletion chr1 TP73 1p36 601990 yes Bartter 4 chr1 CLCNKA 1p36 602024 yes Bartter 3 chr1 CLCNKB 1p36 602023 yes Bartter 4 chr1 BSND 1p32 606412 yes NFIA Haploinsufficiency chr1 NFIA 1p31 600727 yes monosomy 1p31 p22 chr1 DIRAS3 1p31 605193 yes Stickler syndrome chr1 COL11A1 1p21 120280 yes atrial fibrillation chr1 GJA5 1q21 121013 yes Thrombocytopenia absent radius syndrome chr1 GJA8 1q21 600897 yes Short stature chr1 LHX4 1q25 602146 yes Van der Woude syndrome chr1 IRF6 1q32 607199 yes Fryns 1q41 chr1 DISP1 1q41 607502 yes autism chr1 DISC1 1q42 605210 yes MR chr1 TBCE 1q42 604934 yes Feingold chr2 MYCN 2q24 164840 yes Pseudovaginal perineoscrotal hypospadias chr2 SRD5A2 2p23 607306 yes Noonan 4 chr2 SOS1 2p22 182530 yes Cystinuria with mitochondrial disease chr2 SLC3A1 2p21 104614 yes Cystinuria with mitochondrial disease chr2 PREPL 2p21 104614 yes Holopresencaphly 2 chr2 SIX3 2p21 603714 yes autism chr2 NRXN1 2p16 600565 yes MicrodeletionReg 2p15p16.1 microdeletion chr2 2p15 602559 yes ion Nephronophthsis 1 chr2 NPHP1 2q13 607100 yes Holoprosencephaly 9 chr2 GLI2 2q14 165230 yes visceral heterotaxy chr2 CFC1 2q21 605194 yes Mowat-Wilson syndrome chr2 ZEB2 2q22 605802 yes autism chr2 SLC4A10 2q24 605556 yes SCN1A-related seizures chr2 SCN1A 2q24 182389 yes HYPOMYELINATION, GLOBAL CEREBRAL chr2 SLC25A12 2q31 603667 yes Split/hand foot malformation -5 chr2 DLX1 2q31 600029 yes -

Diverse Entry Gates to Mitochondria Provide Insights Into the Evolution of Eukaryotes
1 Review Peeping at TOMs - Diverse entry gates to mitochondria provide insights into the evolution of eukaryotes Jan Mania, Chris Meisingerb and André Schneidera, 1 aDepartment of Chemistry and Biochemistry, University of Bern, Freiestrasse 3, CH- 3012 Bern, Switzerland bInstitut für Biochemie und Molekularbiologie, ZBMZ and BIOSS Centre for Biological Signalling Studies, Universität Freiburg, 79104 Freiburg, Germany 1 Correspondence to: [email protected] | downloaded: 13.3.2017 http://boris.unibe.ch/92438/ source: 2 Mitochondria are essential for eukaryotic life and more than 95% of their proteins are imported as precursors from the cytosol. The targeting signals for this post-translational import are conserved in all eukaryotes. However, this conservation does not hold true for the protein translocase of the mitochondrial outer membrane that serves as entry gate for essentially all precursor proteins. Only two of its subunits, Tom40 and Tom22, are conserved and thus likely were present in the last eukaryotic common ancestor. Tom7 is found in representatives of all supergroups except the Excavates. This suggests that it was added to the core of the translocase after the Excavates segregated from all other eukaryotes. A comparative analysis of the biochemically and functionally characterized outer membrane translocases of yeast, plants and trypanosomes, which represent three eukaryotic supergroups, shows that the receptors that recognize the conserved import signals differ strongly between the different systems. They present a remarkable example of convergent evolution at the molecular level. The structural diversity of the functionally conserved import receptors therefore provides insight into the early evolutionary history of mitochondria. Protein import distinguishes mitochondria from its endosymbiontic ancestor The origin of eukaryotic cells arguably is the most important transition in evolution besides the origin of life itself. -

C6orf203 Controls OXPHOS Function Through Modulation of Mitochondrial Protein Biosynthesis
bioRxiv preprint doi: https://doi.org/10.1101/704403; this version posted July 17, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. C6orf203 controls OXPHOS function through modulation of mitochondrial protein biosynthesis number of characters excluding Materials and Methods: 40,651 Sara Palacios-Zambrano1,2, Luis Vázquez-Fonseca1,2, Cristina González-Páramos1,2, Laura Mamblona1,2, Laura Sánchez-Caballero3, Leo Nijtmans3, Rafael Garesse1,2 and Miguel Angel Fernández-Moreno1,2,* 1 Departamento de Bioquímica, Instituto de Investigaciones Biomédicas “Alberto Sols” UAM CSIC and Centro de Investigación Biomédica en Red en Enfermedades Raras (CIBERER). Facultad de Medicina, Universidad Autónoma de Madrid. Madrid 28029, Spain. 2 Instituto de Investigación Sanitaria Hospital 12 de Octubre (imas12), Madrid 28041, Spain. 3 Department of Pediatrics, Radboud Center for Mitochondrial Medicine, Radboud University Medical Center, Nijmegen, The Netherlands. * To whom correspondence should be addressed. Tel:+34 91 497 31 29; Email: [email protected] Running title “C6orf203 controls mt-proteins synthesis” bioRxiv preprint doi: https://doi.org/10.1101/704403; this version posted July 17, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. ABSTRACT Mitochondria are essential organelles present in the vast majority of eukaryotic cells. Their central function is to produce cellular energy through the OXPHOS system, and functional alterations provoke so-called mitochondrial OXPHOS diseases. It is estimated that several hundred mitochondrial proteins have unknown functions. Very recently, C6orf203 was described to participate in mitochondrial transcription under induced mitochondrial DNA depletion stress conditions. -

Downloaded on 02/15/2018)
proteomes Article Proteomic Analysis of 3T3-L1 Adipocytes Treated with Insulin and TNF-α 1, 2 1, 3 Hayley Chan y, Ketaki P. Bhide , Aditya Vaidyam z, Victoria Hedrick , Tiago Jose Paschoal Sobreira 3, Thomas G. Sors 4, Ryan W. Grant 5,§ and Uma K. Aryal 3,6,* 1 Department of Computer Science, Purdue University, West Lafayette, IN 47907, USA; [email protected] (H.C.); [email protected] (A.V.) 2 College of Agriculture, Purdue University, West Lafayette, IN 47907, USA; [email protected] 3 Purdue Proteomics Facility, Bindley Bioscience Center, Purdue University, West Lafayette, IN 47907, USA; [email protected] (V.H.); [email protected] (T.J.P.S.) 4 Purdue Institute of Inflammation, Immunology and Infectious Disease, Purdue University, West Lafayette, IN 47907, USA; [email protected] 5 Department of Nutrition Science, Purdue University, West Lafayette, IN 47907, USA; [email protected] 6 Department of Comparative Pathobiology, College of Veterinary Medicine, Purdue University, West Lafayette, IN 47907, USA * Correspondence: [email protected]; Tel.: +(765)-494-4960 Present addresses: Indiana University School of Medicine, West Lafayette, IN 47907, USA. y Present addresses: Division of Digital Psychiatry, Harvard Medical School, Boston, MA 02111, USA. z § Present addresses: Pharmavite, Los Angeles, CA 90089, USA. Received: 29 August 2019; Accepted: 17 October 2019; Published: 20 October 2019 Abstract: Insulin resistance is an indication of early stage Type 2 diabetes (T2D). Insulin resistant adipose tissues contain higher levels of insulin than the physiological level, as well as higher amounts of intracellular tumor necrosis factor-α (TNF-α) and other cytokines. However, the mechanism of insulin resistance remains poorly understood. -

Mitochondrial Calcium Regulation of Redox Signaling in Cancer
cells Review Mitochondrial Calcium Regulation of Redox Signaling in Cancer 1, 2, 3 2 Céline Delierneux y , Sana Kouba y, Santhanam Shanmughapriya , Marie Potier-Cartereau , Mohamed Trebak 1 and Nadine Hempel 1,4,* 1 Department of Cellular and Molecular Physiology, The Pennsylvania State University College of Medicine, 500 University Dr., Hershey, PA 17033, USA; [email protected] (C.D.); [email protected] (M.T.) 2 Inserm U1069 Nutrition, Croissance et Cancer, Université de Tours, 10 Boulevard Tonnellé, 37032 Tours, France; [email protected] (S.K.); [email protected] (M.P.-C.) 3 Department of Medicine, The Pennsylvania State University College of Medicine, 500 University Dr., Hershey, PA 17033, USA; [email protected] 4 Department of Pharmacology, and Obstetrics and Gynecology, The Pennsylvania State University College of Medicine, 500 University Dr., Hershey, PA 17033, USA * Correspondence: [email protected]; Tel.: +717-531-4037 These authors contributed equally to this work. y Received: 8 January 2020; Accepted: 10 February 2020; Published: 12 February 2020 Abstract: Calcium (Ca2+) uptake into the mitochondria shapes cellular Ca2+ signals and acts as a key effector for ATP generation. In addition, mitochondria-derived reactive oxygen species (mROS), produced as a consequence of ATP synthesis at the electron transport chain (ETC), modulate cellular signaling pathways that contribute to many cellular processes. Cancer cells modulate mitochondrial Ca2+ ([Ca2+]m) homeostasis by altering the expression and function of mitochondrial Ca2+ channels and transporters required for the uptake and extrusion of mitochondrial Ca2+. Regulated elevations in [Ca2+]m are required for the activity of several mitochondrial enzymes, and this in turn regulates metabolic flux, mitochondrial ETC function and mROS generation. -

Product Size GOT1 P00504 F CAAGCTGT
Table S1. List of primer sequences for RT-qPCR. Gene Product Uniprot ID F/R Sequence(5’-3’) name size GOT1 P00504 F CAAGCTGTCAAGCTGCTGTC 71 R CGTGGAGGAAAGCTAGCAAC OGDHL E1BTL0 F CCCTTCTCACTTGGAAGCAG 81 R CCTGCAGTATCCCCTCGATA UGT2A1 F1NMB3 F GGAGCAAAGCACTTGAGACC 93 R GGCTGCACAGATGAACAAGA GART P21872 F GGAGATGGCTCGGACATTTA 90 R TTCTGCACATCCTTGAGCAC GSTT1L E1BUB6 F GTGCTACCGAGGAGCTGAAC 105 R CTACGAGGTCTGCCAAGGAG IARS Q5ZKA2 F GACAGGTTTCCTGGCATTGT 148 R GGGCTTGATGAACAACACCT RARS Q5ZM11 F TCATTGCTCACCTGCAAGAC 146 R CAGCACCACACATTGGTAGG GSS F1NLE4 F ACTGGATGTGGGTGAAGAGG 89 R CTCCTTCTCGCTGTGGTTTC CYP2D6 F1NJG4 F AGGAGAAAGGAGGCAGAAGC 113 R TGTTGCTCCAAGATGACAGC GAPDH P00356 F GACGTGCAGCAGGAACACTA 112 R CTTGGACTTTGCCAGAGAGG Table S2. List of differentially expressed proteins during chronic heat stress. score name Description MW PI CC CH Down regulated by chronic heat stress A2M Uncharacterized protein 158 1 0.35 6.62 A2ML4 Uncharacterized protein 163 1 0.09 6.37 ABCA8 Uncharacterized protein 185 1 0.43 7.08 ABCB1 Uncharacterized protein 152 1 0.47 8.43 ACOX2 Cluster of Acyl-coenzyme A oxidase 75 1 0.21 8 ACTN1 Alpha-actinin-1 102 1 0.37 5.55 ALDOC Cluster of Fructose-bisphosphate aldolase 39 1 0.5 6.64 AMDHD1 Cluster of Uncharacterized protein 37 1 0.04 6.76 AMT Aminomethyltransferase, mitochondrial 42 1 0.29 9.14 AP1B1 AP complex subunit beta 103 1 0.15 5.16 APOA1BP NAD(P)H-hydrate epimerase 32 1 0.4 8.62 ARPC1A Actin-related protein 2/3 complex subunit 42 1 0.34 8.31 ASS1 Argininosuccinate synthase 47 1 0.04 6.67 ATP2A2 Cluster of Calcium-transporting