Interpreting the Molecular Mechanisms of Yinchenhao Decoction on Hepatocellular Carcinoma Through Absorbed Components Based on Network Pharmacology
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Information to Mammadova-Bach Et Al., “Laminin Α1 Orchestrates VEGFA Functions in the Ecosystem of Colorectal Carcinogenesis”
Supplemental information to Mammadova-Bach et al., “Laminin α1 orchestrates VEGFA functions in the ecosystem of colorectal carcinogenesis” Supplemental material and methods Cloning of the villin-LMα1 vector The plasmid pBS-villin-promoter containing the 3.5 Kb of the murine villin promoter, the first non coding exon, 5.5 kb of the first intron and 15 nucleotides of the second villin exon, was generated by S. Robine (Institut Curie, Paris, France). The EcoRI site in the multi cloning site was destroyed by fill in ligation with T4 polymerase according to the manufacturer`s instructions (New England Biolabs, Ozyme, Saint Quentin en Yvelines, France). Site directed mutagenesis (GeneEditor in vitro Site-Directed Mutagenesis system, Promega, Charbonnières-les-Bains, France) was then used to introduce a BsiWI site before the start codon of the villin coding sequence using the 5’ phosphorylated primer: 5’CCTTCTCCTCTAGGCTCGCGTACGATGACGTCGGACTTGCGG3’. A double strand annealed oligonucleotide, 5’GGCCGGACGCGTGAATTCGTCGACGC3’ and 5’GGCCGCGTCGACGAATTCACGC GTCC3’ containing restriction site for MluI, EcoRI and SalI were inserted in the NotI site (present in the multi cloning site), generating the plasmid pBS-villin-promoter-MES. The SV40 polyA region of the pEGFP plasmid (Clontech, Ozyme, Saint Quentin Yvelines, France) was amplified by PCR using primers 5’GGCGCCTCTAGATCATAATCAGCCATA3’ and 5’GGCGCCCTTAAGATACATTGATGAGTT3’ before subcloning into the pGEMTeasy vector (Promega, Charbonnières-les-Bains, France). After EcoRI digestion, the SV40 polyA fragment was purified with the NucleoSpin Extract II kit (Machery-Nagel, Hoerdt, France) and then subcloned into the EcoRI site of the plasmid pBS-villin-promoter-MES. Site directed mutagenesis was used to introduce a BsiWI site (5’ phosphorylated AGCGCAGGGAGCGGCGGCCGTACGATGCGCGGCAGCGGCACG3’) before the initiation codon and a MluI site (5’ phosphorylated 1 CCCGGGCCTGAGCCCTAAACGCGTGCCAGCCTCTGCCCTTGG3’) after the stop codon in the full length cDNA coding for the mouse LMα1 in the pCIS vector (kindly provided by P. -

Association Analyses of Known Genetic Variants with Gene
ASSOCIATION ANALYSES OF KNOWN GENETIC VARIANTS WITH GENE EXPRESSION IN BRAIN by Viktoriya Strumba A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Bioinformatics) in The University of Michigan 2009 Doctoral Committee: Professor Margit Burmeister, Chair Professor Huda Akil Professor Brian D. Athey Assistant Professor Zhaohui S. Qin Research Statistician Thomas Blackwell To Sam and Valentina Dmitriy and Elizabeth ii ACKNOWLEDGEMENTS I would like to thank my advisor Professor Margit Burmeister, who tirelessly guided me though seemingly impassable corridors of graduate work. Throughout my thesis writing period she provided sound advice, encouragement and inspiration. Leading by example, her enthusiasm and dedication have been instrumental in my path to becoming a better scientist. I also would like to thank my co-advisor Tom Blackwell. His careful prodding always kept me on my toes and looking for answers, which taught me the depth of careful statistical analysis. His diligence and dedication have been irreplaceable in most difficult of projects. I also would like to thank my other committee members: Huda Akil, Brian Athey and Steve Qin as well as David States. You did not make it easy for me, but I thank you for believing and not giving up. Huda’s eloquence in every subject matter she explained have been particularly inspiring, while both Huda’s and Brian’s valuable advice made the completion of this dissertation possible. I would also like to thank all the members of the Burmeister lab, both past and present: Sandra Villafuerte, Kristine Ito, Cindy Schoen, Karen Majczenko, Ellen Schmidt, Randi Burns, Gang Su, Nan Xiang and Ana Progovac. -

Recombinant Human Carbonic Anhydrase XIII Protein Catalog Number: ATGP3059
Recombinant human Carbonic Anhydrase XIII protein Catalog Number: ATGP3059 PRODUCT INPORMATION Expression system E.coli Domain 1-262aa UniProt No. Q8N1Q1 NCBI Accession No. NP_940986.1 Alternative Names Carbonic anhydrase 13, CAXIII PRODUCT SPECIFICATION Molecular Weight 31.8 kDa (285aa) confirmed by MALDI-TOF Concentration 0.5mg/ml (determined by Bradford assay) Formulation Liquid in. Phosphate-Buffered Saline (pH 7.4) containing 10% glycerol, 1mM DTT Purity > 90% by SDS-PAGE Biological Activity Specific activity is > 2,500pmol/min/ug, and is defined as the amount of enzyme that hydrolyze 1.0pmole of 4- nitrophenyl acetate to 4-nitrophenol per minute at pH 7.5 at 37C. Tag His-Tag Application SDS-PAGE, Enzyme Activity Storage Condition Can be stored at +2C to +8C for 1 week. For long term storage, aliquot and store at -20C to -80C. Avoid repeated freezing and thawing cycles. BACKGROUND Description CA13 also known as carbonic anhydrase 13 belongs to the alpha-carbonic anhydrase family. The carbonic anhydrase from a family of enzymes that catalyze the rapid interconversion of carbon dioxide and water to bicarbonate and protons, a reversible reaction that occurs relatively slowly in the absence of catalyst. The active 1 Recombinant human Carbonic Anhydrase XIII protein Catalog Number: ATGP3059 site of most carbonic anhydrases contains a zinc ion; they are claasified as metalloenzymes. There are at least five distinct CA families (alpha, beta, gamma, delta, and epsilon). These families have no significant amino acid sequence similarity and in most cases are thought to be an example of convergent evolution. The alpha-CAs are found in humans. -

Table 2. Significant
Table 2. Significant (Q < 0.05 and |d | > 0.5) transcripts from the meta-analysis Gene Chr Mb Gene Name Affy ProbeSet cDNA_IDs d HAP/LAP d HAP/LAP d d IS Average d Ztest P values Q-value Symbol ID (study #5) 1 2 STS B2m 2 122 beta-2 microglobulin 1452428_a_at AI848245 1.75334941 4 3.2 4 3.2316485 1.07398E-09 5.69E-08 Man2b1 8 84.4 mannosidase 2, alpha B1 1416340_a_at H4049B01 3.75722111 3.87309653 2.1 1.6 2.84852656 5.32443E-07 1.58E-05 1110032A03Rik 9 50.9 RIKEN cDNA 1110032A03 gene 1417211_a_at H4035E05 4 1.66015788 4 1.7 2.82772795 2.94266E-05 0.000527 NA 9 48.5 --- 1456111_at 3.43701477 1.85785922 4 2 2.8237185 9.97969E-08 3.48E-06 Scn4b 9 45.3 Sodium channel, type IV, beta 1434008_at AI844796 3.79536664 1.63774235 3.3 2.3 2.75319499 1.48057E-08 6.21E-07 polypeptide Gadd45gip1 8 84.1 RIKEN cDNA 2310040G17 gene 1417619_at 4 3.38875643 1.4 2 2.69163229 8.84279E-06 0.0001904 BC056474 15 12.1 Mus musculus cDNA clone 1424117_at H3030A06 3.95752801 2.42838452 1.9 2.2 2.62132809 1.3344E-08 5.66E-07 MGC:67360 IMAGE:6823629, complete cds NA 4 153 guanine nucleotide binding protein, 1454696_at -3.46081884 -4 -1.3 -1.6 -2.6026947 8.58458E-05 0.0012617 beta 1 Gnb1 4 153 guanine nucleotide binding protein, 1417432_a_at H3094D02 -3.13334396 -4 -1.6 -1.7 -2.5946297 1.04542E-05 0.0002202 beta 1 Gadd45gip1 8 84.1 RAD23a homolog (S. -

Steroid-Dependent Regulation of the Oviduct: a Cross-Species Transcriptomal Analysis
University of Kentucky UKnowledge Theses and Dissertations--Animal and Food Sciences Animal and Food Sciences 2015 Steroid-dependent regulation of the oviduct: A cross-species transcriptomal analysis Katheryn L. Cerny University of Kentucky, [email protected] Right click to open a feedback form in a new tab to let us know how this document benefits ou.y Recommended Citation Cerny, Katheryn L., "Steroid-dependent regulation of the oviduct: A cross-species transcriptomal analysis" (2015). Theses and Dissertations--Animal and Food Sciences. 49. https://uknowledge.uky.edu/animalsci_etds/49 This Doctoral Dissertation is brought to you for free and open access by the Animal and Food Sciences at UKnowledge. It has been accepted for inclusion in Theses and Dissertations--Animal and Food Sciences by an authorized administrator of UKnowledge. For more information, please contact [email protected]. STUDENT AGREEMENT: I represent that my thesis or dissertation and abstract are my original work. Proper attribution has been given to all outside sources. I understand that I am solely responsible for obtaining any needed copyright permissions. I have obtained needed written permission statement(s) from the owner(s) of each third-party copyrighted matter to be included in my work, allowing electronic distribution (if such use is not permitted by the fair use doctrine) which will be submitted to UKnowledge as Additional File. I hereby grant to The University of Kentucky and its agents the irrevocable, non-exclusive, and royalty-free license to archive and make accessible my work in whole or in part in all forms of media, now or hereafter known. -

P-Glycoprotein-Mediated Chemoresistance Is Reversed by Carbonic Anhydrase XII Inhibitors
www.impactjournals.com/oncotarget/ Oncotarget, Advance Publications 2016 P-glycoprotein-mediated chemoresistance is reversed by carbonic anhydrase XII inhibitors Joanna Kopecka1, Gregory M. Rankin2, Iris C. Salaroglio1, Sally-Ann Poulsen2,*, Chiara Riganti1,* 1Department of Oncology, University of Torino, 10126 Torino, Italy 2Eskitis Institute for Drug Discovery, Griffith University, Brisbane, Nathan, Queensland, 4111, Australia *These authors contributed equally to this work Correspondence to: Sally-Ann Poulsen, email: [email protected] Chiara Riganti, email: [email protected] Keywords: carbonic anhydrase XII, P-glycoprotein, doxorubicin, chemoresistance, intracellular pH Received: August 26, 2016 Accepted: October 28, 2016 Published: November 03, 2016 ABSTRACT Carbonic anhydrase XII (CAXII) is a membrane enzyme that maintains pH homeostasis and sustains optimum P-glycoprotein (Pgp) efflux activity in cancer cells. Here, we investigated a panel of eight CAXII inhibitors (compounds 1–8), for their potential to reverse Pgp mediated tumor cell chemoresistance. Inhibitors (5 nM) were screened in human and murine cancer cells (colon, lung, breast, bone) with different expression levels of CAXII and Pgp. We identified three CAXII inhibitors (compounds 1, 2 and 4) that significantly (≥ 2 fold) increased the intracellular retention of the Pgp-substrate and chemotherapeutic doxorubicin, and restored its cytotoxic activity. The inhibitors lowered intracellular pH to indirectly impair Pgp activity. Ca12-knockout assays confirmed that the chemosensitizing property of the compounds was dependent on active CAXII. Furthermore, in a preclinical model of drug-resistant breast tumors compound 1 (1900 ng/kg) restored the efficacy of doxorubicin to the same extent as the direct Pgp inhibitor tariquidar. The expression of carbonic anhydrase IX had no effect on the intracellular doxorubicin accumulation. -

High-Throughput Screening Studies of Inhibition of Human Carbonic Anhydrase II and Bacterial Flagella Antimicrobial Activity
Western Michigan University ScholarWorks at WMU Dissertations Graduate College 5-2010 High-Throughput Screening Studies of Inhibition of Human Carbonic Anhydrase II and Bacterial Flagella Antimicrobial Activity Albert A. Barrese III Western Michigan University Follow this and additional works at: https://scholarworks.wmich.edu/dissertations Part of the Biochemistry, Biophysics, and Structural Biology Commons, and the Biology Commons Recommended Citation Barrese, Albert A. III, "High-Throughput Screening Studies of Inhibition of Human Carbonic Anhydrase II and Bacterial Flagella Antimicrobial Activity" (2010). Dissertations. 500. https://scholarworks.wmich.edu/dissertations/500 This Dissertation-Open Access is brought to you for free and open access by the Graduate College at ScholarWorks at WMU. It has been accepted for inclusion in Dissertations by an authorized administrator of ScholarWorks at WMU. For more information, please contact [email protected]. HIGH-THROUGHPUT SCREENING STUDIES OF INHIBITION OF HUMAN CARBONIC ANHYDRASE II AND BACTERIAL FLAGELLA ANTIMICROBIAL ACTIVITY by Albert A. Barrese III A Dissertation Submitted to the Faculty of The Graduate College in partial fulfillment of the requirements for the Degree of Doctor of Philosophy Department of Biological Sciences Advisor: Brian C. Tripp, Ph.D. Western Michigan University Kalamazoo, Michigan May 2010 UMI Number: 3410393 All rights reserved INFORMATION TO ALL USERS The quality of this reproduction is dependent upon the quality of the copy submitted. In the unlikely event that the author did not send a complete manuscript and there are missing pages, these will be noted. Also, if material had to be removed, a note will indicate the deletion. UMT Dissertation Publishing UMI 3410393 Copyright 2010 by ProQuest LLC. -
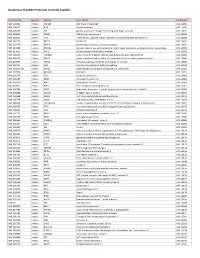
Quantigene Flowrna Probe Sets Currently Available
QuantiGene FlowRNA Probe Sets Currently Available Accession No. Species Symbol Gene Name Catalog No. NM_003452 Human ZNF189 zinc finger protein 189 VA1-10009 NM_000057 Human BLM Bloom syndrome VA1-10010 NM_005269 Human GLI glioma-associated oncogene homolog (zinc finger protein) VA1-10011 NM_002614 Human PDZK1 PDZ domain containing 1 VA1-10015 NM_003225 Human TFF1 Trefoil factor 1 (breast cancer, estrogen-inducible sequence expressed in) VA1-10016 NM_002276 Human KRT19 keratin 19 VA1-10022 NM_002659 Human PLAUR plasminogen activator, urokinase receptor VA1-10025 NM_017669 Human ERCC6L excision repair cross-complementing rodent repair deficiency, complementation group 6-like VA1-10029 NM_017699 Human SIDT1 SID1 transmembrane family, member 1 VA1-10032 NM_000077 Human CDKN2A cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) VA1-10040 NM_003150 Human STAT3 signal transducer and activator of transcripton 3 (acute-phase response factor) VA1-10046 NM_004707 Human ATG12 ATG12 autophagy related 12 homolog (S. cerevisiae) VA1-10047 NM_000737 Human CGB chorionic gonadotropin, beta polypeptide VA1-10048 NM_001017420 Human ESCO2 establishment of cohesion 1 homolog 2 (S. cerevisiae) VA1-10050 NM_197978 Human HEMGN hemogen VA1-10051 NM_001738 Human CA1 Carbonic anhydrase I VA1-10052 NM_000184 Human HBG2 Hemoglobin, gamma G VA1-10053 NM_005330 Human HBE1 Hemoglobin, epsilon 1 VA1-10054 NR_003367 Human PVT1 Pvt1 oncogene homolog (mouse) VA1-10061 NM_000454 Human SOD1 Superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) -

(12) Patent Application Publication (10) Pub. No.: US 2003/0082511 A1 Brown Et Al
US 20030082511A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2003/0082511 A1 Brown et al. (43) Pub. Date: May 1, 2003 (54) IDENTIFICATION OF MODULATORY Publication Classification MOLECULES USING INDUCIBLE PROMOTERS (51) Int. Cl." ............................... C12O 1/00; C12O 1/68 (52) U.S. Cl. ..................................................... 435/4; 435/6 (76) Inventors: Steven J. Brown, San Diego, CA (US); Damien J. Dunnington, San Diego, CA (US); Imran Clark, San Diego, CA (57) ABSTRACT (US) Correspondence Address: Methods for identifying an ion channel modulator, a target David B. Waller & Associates membrane receptor modulator molecule, and other modula 5677 Oberlin Drive tory molecules are disclosed, as well as cells and vectors for Suit 214 use in those methods. A polynucleotide encoding target is San Diego, CA 92121 (US) provided in a cell under control of an inducible promoter, and candidate modulatory molecules are contacted with the (21) Appl. No.: 09/965,201 cell after induction of the promoter to ascertain whether a change in a measurable physiological parameter occurs as a (22) Filed: Sep. 25, 2001 result of the candidate modulatory molecule. Patent Application Publication May 1, 2003 Sheet 1 of 8 US 2003/0082511 A1 KCNC1 cDNA F.G. 1 Patent Application Publication May 1, 2003 Sheet 2 of 8 US 2003/0082511 A1 49 - -9 G C EH H EH N t R M h so as se W M M MP N FIG.2 Patent Application Publication May 1, 2003 Sheet 3 of 8 US 2003/0082511 A1 FG. 3 Patent Application Publication May 1, 2003 Sheet 4 of 8 US 2003/0082511 A1 KCNC1 ITREXCHO KC 150 mM KC 2000000 so 100 mM induced Uninduced Steady state O 100 200 300 400 500 600 700 Time (seconds) FIG. -

An Update on the Metabolic Roles of Carbonic Anhydrases in the Model Alga Chlamydomonas Reinhardtii
H OH metabolites OH Review An Update on the Metabolic Roles of Carbonic Anhydrases in the Model Alga Chlamydomonas reinhardtii Ashok Aspatwar 1,* ID , Susanna Haapanen 1 and Seppo Parkkila 1,2 1 Faculty of Medicine and Life Sciences, University of Tampere, FI-33014 Tampere, Finland; [email protected].fi (S.H.); [email protected].fi (S.P.) 2 Fimlab, Ltd., and Tampere University Hospital, FI-33520 Tampere, Finland * Correspondence: [email protected].fi; Tel.: +358-46-596-2117 Received: 11 January 2018; Accepted: 10 March 2018; Published: 13 March 2018 Abstract: Carbonic anhydrases (CAs) are metalloenzymes that are omnipresent in nature. − + CAs catalyze the basic reaction of the reversible hydration of CO2 to HCO3 and H in all living organisms. Photosynthetic organisms contain six evolutionarily different classes of CAs, which are namely: α-CAs, β-CAs, γ-CAs, δ-CAs, ζ-CAs, and θ-CAs. Many of the photosynthetic organisms contain multiple isoforms of each CA family. The model alga Chlamydomonas reinhardtii contains 15 CAs belonging to three different CA gene families. Of these 15 CAs, three belong to the α-CA gene family; nine belong to the β-CA gene family; and three belong to the γ-CA gene family. The multiple copies of the CAs in each gene family may be due to gene duplications within the particular CA gene family. The CAs of Chlamydomonas reinhardtii are localized in different subcellular compartments of this unicellular alga. The presence of a large number of CAs and their diverse subcellular localization within a single cell suggests the importance of these enzymes in the metabolic and biochemical roles they perform in this unicellular alga. -

The Effects of Betaine Treatment on Rats Fed an Acute Bolus of Ethanol at 3 and 12 H Post Bolus: a Microarray Analysis
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by PubMed Central Genes Nutr (2010) 5:321–329 DOI 10.1007/s12263-010-0173-y RESEARCH PAPER The effects of betaine treatment on rats fed an acute bolus of ethanol at 3 and 12 h post bolus: a microarray analysis J. Li • F. Bardag-Gorce • J. Oliva • B. A. French • J. Dedes • S. W. French Received: 24 November 2009 / Accepted: 19 February 2010 / Published online: 19 March 2010 Ó The Author(s) 2010. This article is published with open access at Springerlink.com Abstract Betaine, a methyl donor active in methionine Introduction metabolism, is effective in preventing and reversing experimental alcohol liver disease. The metabolic and Ethanol, given as a bolus of 6 g/kg body weight, increases molecular biologic mechanisms involved in this prevention the blood and urinary alcohol to a high level at 3 h post are only partially known. To further investigate how betaine bolus. At 12 h, the blood and urinary alcohol levels return modifies the effects of ethanol on the liver, rats were given to low a level [2]. At these 2 time points, there are major an acute ethanol bolus with or without betaine and the changes in gene expression in the liver but only minimal results were compared to isocaloric dextrose-fed controls. evidence of epigenetic changes [2]. Only at 12 h is there a Livers were subjected to microarray analysis, and functional decrease in global DNA methylation [2]. pathways and individual gene expression changes were Previously, in using this model, S-adenosylmethionine analyzed. -

And Primate Evolution Gene Copy Number Variation Spanning 60
Downloaded from genome.cshlp.org on January 27, 2009 - Published by Cold Spring Harbor Laboratory Press Gene copy number variation spanning 60 million years of human and primate evolution Laura Dumas, Young H. Kim, Anis Karimpour-Fard, et al. Genome Res. 2007 17: 1266-1277 originally published online July 31, 2007 Access the most recent version at doi:10.1101/gr.6557307 Supplemental http://genome.cshlp.org/content/suppl/2007/07/26/gr.6557307.DC1.html Material References This article cites 51 articles, 20 of which can be accessed free at: http://genome.cshlp.org/content/17/9/1266.full.html#ref-list-1 Article cited in: http://genome.cshlp.org/content/17/9/1266.full.html#related-urls Email alerting Receive free email alerts when new articles cite this article - sign up in the box at the service top right corner of the article or click here To subscribe to Genome Research go to: http://genome.cshlp.org/subscriptions Copyright © 2007, Cold Spring Harbor Laboratory Press Downloaded from genome.cshlp.org on January 27, 2009 - Published by Cold Spring Harbor Laboratory Press Article Gene copy number variation spanning 60 million years of human and primate evolution Laura Dumas,1 Young H. Kim,2 Anis Karimpour-Fard,3 Michael Cox,1,4,5 Janet Hopkins,1,4,5 Jonathan R. Pollack,2 and James M. Sikela1,4,5,6 1Human Medical Genetics Program, University of Colorado at Denver and Health Sciences Center, Aurora, Colorado 80045, USA; 2Department of Pathology, Stanford University, Stanford, California 94305, USA; 3Department of Preventative Medicine and Biometrics,