Gene Expression Analysis of Zobellia Galactanivorans During The
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Sequence from B4 Sponge with (A) the First BLAST Hit Asbestopluma Lycopodium and (B) the Sequence of M
Supplementary Material Figure S1. Alignments of CO1 (PorCOI2fwd/PorCOI2rev) sequence from B4 sponge with (A) the first BLAST hit Asbestopluma lycopodium and (B) the sequence of M. acerata displaying low query cover. 1 Figure S2. Alignment of CO1 (dgLCO1490/dgHCO2198) sequence from B4 sponge with the first BLAST hit (M. acerata). 2 Figure S3. Alignment of CO1 (dgLCO1490/dgHCO2198) sequence from D4 sponge with the first BLAST hit (H. pilosus). 3 Figure S4. Taxonomy Bar Plot, reporting the relative frequencies (in percentage, %) of the bacteria taxons more representative for each of the four sponges under analysis . Sample code: B4= M. (Oxymycale) acerata; D4= H. pilosus, D6= M. sarai, C6= H. (Rhizoniera) dancoi. Each taxon is highlighted by a different color. 4 Figure S5. Krona plot at the seven increasing complexity levels: (a) Regnum, (b) Phylum, (c) Class, (d) Order, (e) Family, (f) Genus and (g) Species. a) 5 b) 6 c) 7 d) 8 e) 9 f) 10 g) 11 Figure S6. Distribution of ASV’s frequencies. 12 Figure S7. Distribution of ASV’s frequencies for each sample (reported as a blue bar). 13 Table S1. BLAST results from B4 sponge (Mycale (Oxymycale) acerata). The primer names, sequence length in base pairs (bp), first hits (highlighted in bold), hits at low significance displaying the correct species (where present), query cover and identity percentages (%) were reported. Sequence Query Identity Primers BLAST results length (bp) cover (%) (%) Mycale macilenta voucher 0CDN7203‐O small subunit 18S A/B 1700 99 98 ribosomal RNA gene, partial sequence Mycale -

Microbial Community Structure and Function on Sinking Particles in the North Pacific Subtropical Gyre
Microbial community structure and function on sinking particles in the North Pacific Subtropical Gyre The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Fontanez, Kristina M., John M. Eppley, Ty J. Samo, David M. Karl, and Edward F. DeLong. “Microbial Community Structure and Function on Sinking Particles in the North Pacific Subtropical Gyre.” Frontiers in Microbiology 6 (May 19, 2015). As Published http://dx.doi.org/10.3389/fmicb.2015.00469 Publisher Frontiers Research Foundation Version Final published version Citable link http://hdl.handle.net/1721.1/98187 Terms of Use Creative Commons Attribution Detailed Terms http://creativecommons.org/licenses/by/4.0/ ORIGINAL RESEARCH published: 19 May 2015 doi: 10.3389/fmicb.2015.00469 Microbial community structure and function on sinking particles in the North Pacific Subtropical Gyre Kristina M. Fontanez 1, John M. Eppley 1, 2, 3, Ty J. Samo 2, 3, 4, David M. Karl 2, 3 and Edward F. DeLong 1, 2, 3* 1 Department of Civil and Environmental Engineering, Massachusetts Institute of Technology, Cambridge, MA, USA, 2 Department of Oceanography, School of Ocean and Earth Science and Technology, University of Hawaii, Honolulu, HI, USA, 3 Daniel K. Inouye Center for Microbial Oceanography: Research and Education, University of Hawaii, Honolulu, HI, USA, 4 Lawrence Livermore National Laboratory, Nuclear and Chemical Sciences Division, Livermore, CA, USA Sinking particles mediate the transport of carbon and energy to the deep-sea, yet the specific microbes associated with sedimenting particles in the ocean’s interior remain largely uncharacterized. In this study, we used particle interceptor traps (PITs) Edited by: to assess the nature of particle-associated microbial communities collected at a variety Anton F. -

Succession and Cazyme Expression of Marine Bacterial
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.08.416354; this version posted December 8, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Sweet and magnetic: Succession and CAZyme expression of marine bacterial communities encountering a mix of alginate and pectin particles Carina Bunse1,2,*, Hanna Koch3,4,*, Sven Breider3, Meinhard Simon1,3 and Matthias Wietz2,3# 1Helmholtz Institute for Functional Marine Biodiversity at the University of Oldenburg, Germany 2Alfred Wegener Institute Helmholtz Centre for Polar and Marine Research, Germany 3Institute for Chemistry and Biology of the Marine Environment, University of Oldenburg, Germany 4Department of Microbiology, Radboud University Nijmegen, The Netherlands *These authors contributed equally to this work. #Corresponding author HGF-MPG Joint Research Group for Deep-Sea Ecology and Technology Alfred Wegener Institute Helmholtz Centre for Polar and Marine Research Am Handelshafen 12 27570 Bremerhaven Germany Email: [email protected] Telephone +49-471-48311454 bioRxiv preprint doi: https://doi.org/10.1101/2020.12.08.416354; this version posted December 8, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. ABSTRACT Polysaccharide particles are an important nutrient source and microhabitat for marine bacteria. However, substrate-specific bacterial dynamics in a mixture of particle types with different polysaccharide composition, as likely occurring in natural habitats, are undescribed. -

Gene Expression Analysis of Zobellia Galactanivorans During The
ORIGINAL RESEARCH published: 21 September 2017 doi: 10.3389/fmicb.2017.01808 Gene Expression Analysis of Zobellia galactanivorans during the Degradation of Algal Polysaccharides Reveals both Substrate-Specific and Shared Transcriptome-Wide Responses François Thomas 1*, Philippe Bordron 2, 3, 4, Damien Eveillard 5 and Gurvan Michel 1* 1 Sorbonne Universités, UPMC Univ Paris 06, Centre National de la Recherche Scientifique, UMR 8227, Integrative Biology of Marine Models, Station Biologique de Roscoff, Roscoff, France, 2 Sorbonne Universités, UPMC Univ Paris 06, Centre National de la Recherche Scientifique, FR2424, Analysis and Bioinformatics for Marine Science, Station Biologique de 3 4 Edited by: Roscoff, Roscoff, France, Mathomics, Center for Mathematical Modeling, Universidad de Chile, Santiago, Chile, Center for 5 Catherine Ayn Brissette, Genome Regulation (Fondap 15090007), Universidad de Chile, Santiago, Chile, Université de Nantes, Laboratoire des University of North Dakota, Sciences du Numérique de Nantes, Centre National de la Recherche Scientifique, ECN, IMTA, Nantes, France United States Reviewed by: Flavobacteriia are recognized as key players in the marine carbon cycle, due to their Thomas Schweder, ability to efficiently degrade algal polysaccharides both in the open ocean and in coastal University of Greifswald, Germany Matthias Wietz, regions. The chemical complexity of algal polysaccharides, their differences between University of Oldenburg, Germany algal groups and variations through time and space, imply that marine flavobacteria *Correspondence: have evolved dedicated degradation mechanisms and regulation of their metabolism François Thomas [email protected] during interactions with algae. In the present study, we report the first transcriptome-wide Gurvan Michel gene expression analysis for an alga-associated flavobacterium during polysaccharide [email protected] degradation. -

Interactions in Self-Assembled Microbial Communities Saturate with Diversity
bioRxiv preprint doi: https://doi.org/10.1101/347948; this version posted June 16, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Interactions in self-assembled microbial communities saturate with diversity Xiaoqian Yu1, Martin F. Polz2*, Eric J. Alm2,3,4,5* 1. Department of Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA 2. Department of Civil and Environmental Engineering, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA 3. Department of Biological Engineering, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA 4. Center for Microbiome Informatics and Therapeutics, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA 5. Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA *Correspondence should be addressed to: E.J.A. ([email protected]) or M.F.P. ([email protected]) Abstract How the diversity of organisms competing for or sharing resources influences community production is an important question in ecology but has rarely been explored in natural microbial communities. These generally contain large numbers of species making it difficult to disentangle how the effects of different interactions scale with diversity. Here, we show that changing diversity affects measures of community function in relatively simple communities but that increasing richness beyond a threshold has little detectable effect. We generated self-assembled communities with a wide range of diversity by growth of cells from serially diluted seawater on brown algal leachate. We subsequently isolated the most abundant taxa from these communities via dilution-to-extinction in order to compare productivity functions of the entire community to those of individual taxa. -

Genome-Wide Analysis of PL7 Alginate Lyases in the Genus Zobellia
molecules Article Genome-Wide Analysis of PL7 Alginate Lyases in the Genus Zobellia Nadezhda Chernysheva, Evgeniya Bystritskaya, Galina Likhatskaya , Olga Nedashkovskaya and Marina Isaeva * G.B. Elyakov Pacific Institute of Bioorganic Chemistry, Far Eastern Branch, Russian Academy of Sciences, 159, Pr. 100 let Vladivostoku, 690022 Vladivostok, Russia; [email protected] (N.C.); [email protected] (E.B.); [email protected] (G.L.); [email protected] (O.N.) * Correspondence: [email protected]; Tel.: +7-423-231-1168 Abstract: We carried out a detailed investigation of PL7 alginate lyases across the Zobellia genus. The main findings were obtained using the methods of comparative genomics and spatial structure modeling, as well as a phylogenomic approach. Initially, in order to elucidate the alginolytic potential of Zobellia, we calculated the content of polysaccharide lyase (PL) genes in each genome. The genus- specific PLs were PL1, PL6, PL7 (the most abundant), PL14, PL17, and PL40. We revealed that PL7 belongs to subfamilies 3, 5, and 6. They may be involved in local and horizontal gene transfer and gene duplication processes. Most likely, an individual evolution of PL7 genes promotes the genetic variability of the Alginate Utilization System across Zobellia. Apparently, the PL7 alginate lyases may acquire a sub-functionalization due to diversification between in-paralogs. Keywords: Zobellia; genomes; polysaccharide lyase family 7; alginate utilization system; paralogs; orthologs Citation: Chernysheva, N.; Bystritskaya, E.; Likhatskaya, G.; Nedashkovskaya, O.; Isaeva, M. Genome-Wide Analysis of PL7 1. Introduction Alginate Lyases in the Genus Zobellia. Marine algal polysaccharides are an important nutrient source for marine bacteria. To Molecules 2021, 26, 2387. -

The Mannitol Utilization System of the Marine Bacterium Zobellia Galactanivorans Agnès Groisillier, Aurore Labourel, Gurvan Michel, Thierry Tonon
The Mannitol Utilization System of the Marine Bacterium Zobellia galactanivorans Agnès Groisillier, Aurore Labourel, Gurvan Michel, Thierry Tonon To cite this version: Agnès Groisillier, Aurore Labourel, Gurvan Michel, Thierry Tonon. The Mannitol Utilization System of the Marine Bacterium Zobellia galactanivorans. Applied and Environmental Microbiology, Ameri- can Society for Microbiology, 2015, 81, pp.1799 - 1812. 10.1128/AEM.02808-14. hal-01116467 HAL Id: hal-01116467 https://hal.archives-ouvertes.fr/hal-01116467 Submitted on 13 Feb 2015 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. 1 1 The mannitol utilization system of the marine bacterium Zobellia galactanivorans 2 3 4 Agnès Groisillier,a,b, Aurore Labourel,a,b,*, Gurvan Michel,a,b and Thierry Tonona,b,# 5 6 Sorbonne Universités, UPMC Univ Paris 06, UMR 8227, Integrative Biology of Marine 7 Models, Station Biologique de Roscoff, Francea; CNRS, UMR 8227, Integrative Biology of 8 Marine Models, Station Biologique de Roscoff, Franceb 9 10 Running title: Mannitol degradation by Zobellia galactanivorans 11 12 #Address correspondence to Thierry Tonon, [email protected] 13 14 *Present address: University of Newcastle, Cell and Molecular Biosciences, Medical School, 15 Cookson Bidg, Framligton Place, NE2 4HH Newcastle upon Tyne, United Kingdom 16 17 A.G. -
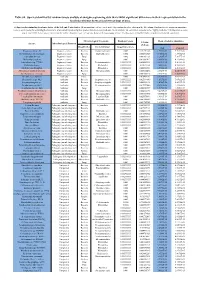
Table S8. Species Identified by Random Forests Analysis of Shotgun Sequencing Data That Exhibit Significant Differences In
Table S8. Species identified by random forests analysis of shotgun sequencing data that exhibit significant differences in their representation in the fecal microbiomes between each two groups of mice. (a) Species discriminating fecal microbiota of the Soil and Control mice. Mean importance of species identified by random forest are shown in the 5th column. Random forests assigns an importance score to each species by estimating the increase in error caused by removing that species from the set of predictors. In our analysis, we considered a species to be “highly predictive” if its importance score was at least 0.001. T-test was performed for the relative abundances of each species between the two groups of mice. P-values were at least 0.05 to be considered statistically significant. Microbiological Taxonomy Random Forests Mean of relative abundance P-Value Species Microbiological Function (T-Test) Classification Bacterial Order Importance Score Soil Control Rhodococcus sp. 2G Engineered strain Bacteria Corynebacteriales 0.002 5.73791E-05 1.9325E-05 9.3737E-06 Herminiimonas arsenitoxidans Engineered strain Bacteria Burkholderiales 0.002 0.005112829 7.1580E-05 1.3995E-05 Aspergillus ibericus Engineered strain Fungi 0.002 0.001061181 9.2368E-05 7.3057E-05 Dichomitus squalens Engineered strain Fungi 0.002 0.018887472 8.0887E-05 4.1254E-05 Acinetobacter sp. TTH0-4 Engineered strain Bacteria Pseudomonadales 0.001333333 0.025523638 2.2311E-05 8.2612E-06 Rhizobium tropici Engineered strain Bacteria Rhizobiales 0.001333333 0.02079554 7.0081E-05 4.2000E-05 Methylocystis bryophila Engineered strain Bacteria Rhizobiales 0.001333333 0.006513543 3.5401E-05 2.2044E-05 Alteromonas naphthalenivorans Engineered strain Bacteria Alteromonadales 0.001 0.000660472 2.0747E-05 4.6463E-05 Saccharomyces cerevisiae Engineered strain Fungi 0.001 0.002980726 3.9901E-05 7.3043E-05 Bacillus phage Belinda Antibiotic Phage 0.002 0.016409765 6.8789E-07 6.0681E-08 Streptomyces sp. -

Transfer of Carbohydrate-Active Enzymes from Marine Bacteria to Japanese Gut Microbiota
Vol 464 | 8 April 2010 | doi:10.1038/nature08937 LETTERS Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota Jan-Hendrik Hehemann1,2{, Gae¨lle Correc1,2, Tristan Barbeyron1,2, William Helbert1,2, Mirjam Czjzek1,2 & Gurvan Michel1,2 Gut microbes supply the human body with energy from dietary not possess the critical residues needed for recognition of agarose or polysaccharides through carbohydrate active enzymes, or k-carrageenan12. CAZymes1, which are absent in the human genome. These enzymes To identify their substrate specificity we cloned and expressed these target polysaccharides from terrestrial plants that dominated diet five GH16 genes in Escherichia coli. However, only Zg1017 and the throughout human evolution2. The array of CAZymes in gut catalytic module of Zg2600 were expressed as soluble proteins and microbes is highly diverse, exemplified by the human gut symbiont could be further analysed (Supplementary Fig. 2). As predicted, these Bacteroides thetaiotaomicron3, which contains 261 glycoside proteins had no activity on commercial agarose (Supplementary hydrolases and polysaccharide lyases, as well as 208 homologues Fig. 3) or k-carrageenan. Consequently, we screened their hydrolytic of susC and susD-genes coding for two outer membrane proteins activity against natural polysaccharides extracted from various marine involved in starch utilization1,4. A fundamental question that, to macrophytes (data not shown). Zg2600 and Zg1017 were found to be our knowledge, has yet to be addressed is how this diversity evolved active only on extracts from the agarophytic red algae Gelidium, by acquiring new genes from microbes living outside the gut. Here Gracilaria and Porphyra, as shown by the release of reducing ends we characterize the first porphyranases from a member of the (Fig. -

Bacteria of the Human Gut Microbiome Catabolize Red Seaweed Glycans with Carbohydrate-Active Enzyme Updates from Extrinsic Microbes
Bacteria of the human gut microbiome catabolize red seaweed glycans with carbohydrate-active enzyme updates from extrinsic microbes Jan-Hendrik Hehemanna, Amelia G. Kellyb, Nicholas A. Pudlob, Eric C. Martensb,1, and Alisdair B. Borastona,1 aDepartment of Biochemistry and Microbiology, University of Victoria, Victoria, BC, Canada V8W 3P6; and bDepartment of Microbiology and Immunology, University of Michigan Medical School, Ann Arbor, MI 48109 Edited by Edward F. DeLong, Massachusetts Institute of Technology, Cambridge, MA, and approved October 7, 2012 (received for review June 28, 2012) Humans host an intestinal population of microbes—collectively re- possible CAZymes encoding genes that appear to have been ac- ferred to as the gut microbiome—which encode the carbohydrate quired by HGT from marine microbes and which may target car- active enzymes, or CAZymes, that are absent from the human ge- bohydrates from red seaweeds (4). nome. These CAZymes help to extract energy from recalcitrant poly- The major matrix polysaccharides in the cell walls of red algae— saccharides. The question then arises as to if and how the microbiome which are the most common dietary red seaweed polysaccharides adapts to new carbohydrate sources when modern humans change consumed by humans and present in many processed foods (11, eating habits. Recent metagenome analysis of microbiomes from 12)—are carrageenans (13), agars, and porphyran (14), and healthy American, Japanese, and Spanish populations identified pu- all contain sulfate esters that are absent in terrestrial plants. tative CAZymes obtained by horizontal gene transfer from marine Furthermore, the sugar backbones can contain unique mono- bacteria, which suggested that human gut bacteria evolved to de- saccharides, such as the 3,6-anhydro-D-galactose present in the grade algal carbohydrates—for example, consumed in form of sushi. -

Gut Microbiota and You
Joint Graduate Presentation Department of Microbiology Gut Microbiota and you Presenter: Henry Chow MPhil Student Supervisor: Prof. Margaret Ip Content 1. Introduction Stomach distress in foreign countries 2. Digestion of Porphyran, a sulphated polysaccharide found in seaweeds Digestion by Z. galactanivorans (marine bacteria) Digestion by B. plebeius (gut bacteria) Digestion of Porphyran between Japanese and Westerners 3. Relation of Food and gut microbiota Gut microbiota Introduction: Stomach distress when eating foreign food? Stomach distress in Foreign places 50% of people travel to foreign places suffer from stomach distress 1 Contaminated food and beverages Allergy from exotic foodstuff Other reasons? 1. Travelers' Diarrhea,Centers for Disease Control and Prevention, http://www.cdc.gov/ncidod/dbmd/diseaseinfo/travelersdiarrhea_g.htm Accessed on 1 December 2010 Diet and digestion 2. Digestion of Porphyran Did you suffer indigestion in Japan? Japanese diet Seaweeds (Nori) • Comprises mainly Sulphated polysaccharides porphyran • Indigestible by human and our gut microbiota • Digestible by Japanese AND… • A marine bacteria which cannot survive in • 2. Cynthia L. Sears “A dynamic partnership: Celebrating our gut flora” Anaerobe, Volume 12, Issue human gut 2, April 2006, Page 114 Zobellia galactanivorans Digestion of porphyran by Z. galactanivorans (marine bacteria) Zobellia galactanivorans • Whole genome sequencing of Z. galactanivorans, a marine Bacteroidetes isolated from Delesseria sanguinea (marine red algae) • Identify -

Morphogenesis of Ulva Mutabilis (Chlorophyta) Induced by Maribacter Species (Bacteroidetes, Flavobacteriaceae)
Botanica Marina 2017; 60(2): 197–206 Short communication Open Access Anne Weiss, Rodrigo Costa and Thomas Wichard* Morphogenesis of Ulva mutabilis (Chlorophyta) induced by Maribacter species (Bacteroidetes, Flavobacteriaceae) DOI 10.1515/bot-2016-0083 related Flavobacteriaceae and a Maribacter strain isolated Received 31 July, 2016; accepted 11 January, 2017; online first from a red alga did not possess any activity. 17 February, 2017 Keywords: bacteroidetes; cell differentiation; green mac- Abstract: Growth and morphogenesis of the sea lettuce Ulva roalga; morphogens; thallusin. (Chlorophyta) depends on the combination of regulative morphogenetic compounds released by specific associated bacteria. Axenic Ulva gametes develop parthenogenetically The green macroalga Ulva mutabilis Føyn (Chlorophyta) is into callus-like colonies consisting of undifferentiated cells not able to develop and differentiate into blade, stem and without normal cell walls. In Ulva mutabilis Føyn, two bac- rhizoid cells under axenic conditions, or when its microbi- terial strains, Maribacter sp. strain MS6 and Roseovarius ome is not appropriate. Instead, the alga forms callus-like, strain MS2, can restore the complete algal morphogenesis slow growing structures with colourless protrusions from forming a tripartite symbiotic community. Morphogenetic the exterior cell wall (Spoerner et al. 2012, Wichard 2015). compounds ( = morphogens) released by the MS6-strain Early experiments of Provasoli (1958) have already pointed induce rhizoid formation and cell wall development in U. out that treatment of Ulva with antibiotics results in abnor- mutabilis, while several bacteria of the Roseobacter clade, mal growth. Further experiments examined the role of including the MS2-strain, promote blade cell division and isolated bacteria in activating developmental and growth thallus elongation.