Isolation and Physiological Characterization of Microorganisms
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Marsarchaeota Are an Aerobic Archaeal Lineage Abundant in Geothermal Iron Oxide Microbial Mats
Marsarchaeota are an aerobic archaeal lineage abundant in geothermal iron oxide microbial mats Authors: Zackary J. Jay, Jacob P. Beam, Mansur Dlakic, Douglas B. Rusch, Mark A. Kozubal, and William P. Inskeep This is a postprint of an article that originally appeared in Nature Microbiology on May 14, 2018. The final version can be found at https://dx.doi.org/10.1038/s41564-018-0163-1. Jay, Zackary J. , Jacob P. Beam, Mensur Dlakic, Douglas B. Rusch, Mark A. Kozubal, and William P. Inskeep. "Marsarchaeota are an aerobic archaeal lineage abundant in geothermal iron oxide microbial mats." Nature Microbiology 3, no. 6 (May 2018): 732-740. DOI: 10.1038/ s41564-018-0163-1. Made available through Montana State University’s ScholarWorks scholarworks.montana.edu Marsarchaeota are an aerobic archaeal lineage abundant in geothermal iron oxide microbial mats Zackary J. Jay1,4,7, Jacob P. Beam1,5,7, Mensur Dlakić2, Douglas B. Rusch3, Mark A. Kozubal1,6 and William P. Inskeep 1* The discovery of archaeal lineages is critical to our understanding of the universal tree of life and evolutionary history of the Earth. Geochemically diverse thermal environments in Yellowstone National Park provide unprecedented opportunities for studying archaea in habitats that may represent analogues of early Earth. Here, we report the discovery and character- ization of a phylum-level archaeal lineage proposed and herein referred to as the ‘Marsarchaeota’, after the red planet. The Marsarchaeota contains at least two major subgroups prevalent in acidic, microaerobic geothermal Fe(III) oxide microbial mats across a temperature range from ~50–80 °C. Metagenomics, single-cell sequencing, enrichment culturing and in situ transcrip- tional analyses reveal their biogeochemical role as facultative aerobic chemoorganotrophs that may also mediate the reduction of Fe(III). -
A Deep Learning Approach to Pattern Recognition for Short DNA Sequences”
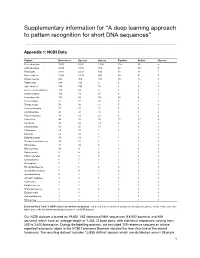
Supplementary information for “A deep learning approach to pattern recognition for short DNA sequences” Appendix 1: NCBI Data Phylum References Species Genera Families Orders Classes Proteobacteria 7,053 5,061 1,106 158 55 9 Actinobacteria 4,768 3,313 383 68 29 6 Firmicutes 3,814 2,531 499 56 13 7 Bacteroidetes 1,934 1,525 360 39 8 7 Euryarchaeota 834 450 100 28 13 8 Tenericutes 266 195 8 5 4 1 Spirochaetes 146 100 16 6 4 1 Deinococcus-Thermus 118 99 9 3 2 1 Crenarchaeota 113 61 27 8 5 1 Cyanobacteria 113 88 59 30 8 2 Fusobacteria 75 37 10 2 1 1 Thermotogae 70 48 13 5 4 1 Verrucomicrobia 57 53 22 7 4 3 Acidobacteria 45 41 19 5 5 4 Planctomycetes 44 31 23 5 3 2 Chloroflexi 44 35 26 15 12 8 Aquificae 43 32 14 4 2 1 Synergistetes 31 25 15 1 1 1 Chlamydiae 28 18 7 5 2 1 Chlorobi 21 16 5 1 1 1 Deferribacteres 15 11 7 1 1 1 Thermodesulfobacteria 14 12 5 1 1 1 Nitrospirae 11 10 3 1 1 1 Fibrobacteres 10 4 3 3 3 3 Balneolaeota 9 9 4 1 1 1 Chrysiogenetes 6 4 3 1 1 1 Lentisphaerae 6 5 3 3 3 2 Dictyoglomi 5 2 1 1 1 1 Rhodothermaeota 5 5 3 2 1 1 Gemmatimonadetes 5 4 3 2 2 2 Ignavibacteriae 4 2 2 2 1 1 Armatimonadetes 4 3 3 3 3 3 Caldiserica 3 1 1 1 1 1 Calditrichaeota 2 2 1 1 1 1 Thaumarchaeota 2 2 2 2 2 2 Elusimicrobia 2 1 1 1 1 1 Kiritimatiellaeota 1 1 1 1 1 1 Nitrospinae 1 1 1 1 1 1 Extended Data Table 1: NCBI dataset breakdown by phylum. -

Supplementary Materials
SUPPLEMENTARY MATERIALS Table S1. Chemical characteristics of the two digestate forms (SD and WD). Values quoted are expressed as % of air-dry digestate (means followed by standard error in brackets). References for the employed methods used for determination of each chemical characteristic is also reported. SD WD Reference Org C % 44.4 (0.33) 1.1 (0.01) [81] Tot N % 1.4 (0.01) 0.4 (0.01) [82] C/N 31.4 (0.17) 3.1 (0.04) NH4-N % n.d 0.2 (0.00) [83] K % 1.7 (0.00) n.d. [84] P % 0.9 (0.01) n.d. [84] S % 0.23 (0.02) n.d. [85] SD = solid digestate; WD = whole digestate. Table S2. Soil physical and chemical characteristics at the beginning of trial (t0) (means from 9 observations followed by standard errors in brackets). Clay (%) 41.9 (1.22) Silt (%) 47.8 (2.13) Moisture (%) 24.46 (1.24) Bulk density (g cm-3) 1.39 (0.04) pH 8.3 (0) CaCO3 (%) 11.4 (0.7) TOC (g kg-1) 12.8 (0.3) TN (g kg-1) 1.4 (0) C/N 9.4 (0.3) CEC (cmol(+) kg-1) 21.0 (0.7) Exchangeable Bases (mg kg-1) K 278.7 (8.0) Na 22.0 (3.0) Mg 201.7 (25.6) Ca 3718.6 (176.3) Available Microelements (mg kg-1) Cu 28.0 (5.0) Zn 1.7 (0.2) Fe 15.4 (0.5) Mn 16.3 (0.5) TOC = total organic C; TN = total N; CEC = cation exchange capacity Table S3. -

Phylogenetics of Archaeal Lipids Amy Kelly 9/27/2006 Outline
Phylogenetics of Archaeal Lipids Amy Kelly 9/27/2006 Outline • Phlogenetics of Archaea • Phlogenetics of archaeal lipids • Papers Phyla • Two? main phyla – Euryarchaeota • Methanogens • Extreme halophiles • Extreme thermophiles • Sulfate-reducing – Crenarchaeota • Extreme thermophiles – Korarchaeota? • Hyperthermophiles • indicated only by environmental DNA sequences – Nanoarchaeum? • N. equitans a fast evolving euryarchaeal lineage, not novel, early diverging archaeal phylum – Ancient archael group? • In deepest brances of Crenarchaea? Euryarchaea? Archaeal Lipids • Methanogens – Di- and tetra-ethers of glycerol and isoprenoid alcohols – Core mostly archaeol or caldarchaeol – Core sometimes sn-2- or Images removed due to sn-3-hydroxyarchaeol or copyright considerations. macrocyclic archaeol –PMI • Halophiles – Similar to methanogens – Exclusively synthesize bacterioruberin • Marine Crenarchaea Depositional Archaeal Lipids Biological Origin Environment Crocetane methanotrophs? methane seeps? methanogens, PMI (2,6,10,15,19-pentamethylicosane) methanotrophs hypersaline, anoxic Squalane hypersaline? C31-C40 head-to-head isoprenoids Smit & Mushegian • “Lost” enzymes of MVA pathway must exist – Phosphomevalonate kinase (PMK) – Diphosphomevalonate decarboxylase – Isopentenyl diphosphate isomerase (IPPI) Kaneda et al. 2001 Rohdich et al. 2001 Boucher et al. • Isoprenoid biosynthesis of archaea evolved through a combination of processes – Co-option of ancestral enzymes – Modification of enzymatic specificity – Orthologous and non-orthologous gene -

Heat Resistant Thermophilic Endospores in Cold Estuarine Sediments
Heat resistant thermophilic endospores in cold estuarine sediments Emma Bell Thesis submitted for the degree of Doctor of Philosophy School of Civil Engineering and Geosciences Faculty of Science, Agriculture and Engineering February 2016 Abstract Microbial biogeography explores the spatial and temporal distribution of microorganisms at multiple scales and is influenced by environmental selection and passive dispersal. Understanding the relative contribution of these factors can be challenging as their effects can be difficult to differentiate. Dormant thermophilic endospores in cold sediments offer a natural model for studies focusing on passive dispersal. Understanding distributions of these endospores is not confounded by the influence of environmental selection; rather their occurrence is due exclusively to passive transport. Sediment heating experiments were designed to investigate the dispersal histories of various thermophilic spore-forming Firmicutes in the River Tyne, a tidal estuary in North East England linking inland tributaries with the North Sea. Microcosm incubations at 50-80°C were monitored for sulfate reduction and enriched bacterial populations were characterised using denaturing gradient gel electrophoresis, functional gene clone libraries and high-throughput sequencing. The distribution of thermophilic endospores among different locations along the estuary was spatially variable, indicating that dispersal vectors originating in both warm terrestrial and marine habitats contribute to microbial diversity in estuarine and marine environments. In addition to their persistence in cold sediments, some endospores displayed a remarkable heat-resistance surviving multiple rounds of autoclaving. These extremely heat-resistant endospores are genetically similar to those detected in deep subsurface environments, including geothermal groundwater investigated from a nearby terrestrial borehole drilled to >1800 m depth with bottom temperatures in excess of 70°C. -

Bacterial Profiles and Antibiogrants of the Bacteria Isolated of the Exposed Pulps of Dog and Cheetah Canine Teeth
Bacterial profiles and antibiogrants of the bacteria isolated of the exposed pulps of dog and cheetah canine teeth A dissertation submitted to the Faculty of Veterinary Science, University of Pretoria. In partial fulfillment of the requirements for the degree Master of Science (Veterinary Science) Promoter: Dr. Gerhard Steenkamp Co-promoter: Ms. Anna-Mari Bosman Department of Companion Animal Clinical Studies Faculty of Veterinary Science University of Pretoria Pretoria (January 2012) J.C. Almansa Ruiz © University of Pretoria Declaration I declare that the dissertation that I hereby submit for the Masters of Science degree in Veterinary Science at the University of Pretoria has not previously been submitted by me for degree purposes at any other university. J.C. Almansa Ruiz ii Dedications To one of the most amazing hunters of the African bush, the Cheetah, that has made me dream since I was a child, and the closest I had been to one, before starting this project was in National Geographic documentaries. I wish all of them a better future in which their habitat will be more respected. To all conservationists, especially to Carla Conradie and Dave Houghton, for spending their lives saving these animals which are suffering from the consequences of the encroachment of human beings into their territory. Some, such as George Adamson, the lion conservationist, even lost their lives in this mission. To the conservationist, Lawrence Anthony, for risking his life in a suicide mission to save the animals in the Baghdad Zoo, when the conflict in Iraq exploded. To my mother and father Jose Maria and Rosa. -

7. References
University of Akureyri Department of Natural Resource Science 7. References Ammann, E. C., Reed, L. L., & Durichek, J. J. (1968). Gas consumptions and Growth Rate of Hydrogenomonas eutropha in Continuous Culture. Applied Microbiology , 16, (6), 822-826. Aguiar, P., Beveridge, T. J., & Reysenbach, A.-L. (2004). Sulfurihydrogenibium azorense, sp. nov., a thermophilic hydrogen oxidizing microaerophile from terrestrial hot springs in the Azores. International Journal of Systematic and Evolutionary Microbiology , 54, 33-39. Altschul, S., Gish, W., Miller, W., Myers, E., & Lipman, D. (1990). "Basic local alignment search tool.". J. Mol. Biol. , 215:403-410. Amend, J., & Shock, E. (2001). Energetics of overall metabolic reactions of thermophilic and hyperthermophilic Archaea and Bacteria. FEMS Microbiology Reviews , 25, 175-243. Aragno, M. (1978). Enrichment, isolation and preliminary characterization of a thermophilic, endospore-forming hydrogen bacterium. FEMS Micobiol. Lett. , 3: 13-15. Aragno, M. (1992). The Thermophilic, Aerobic, Hydrogen-Oxidizing (Knallgas) Bacteria. In A. Balows, H. Trüper, M. Dworkin, W. Harder, & K. Schleifer, The Prokaryotes, a handbook on biology of bacteria. 2nd ed. vol. 4 (pp. 3917-3933.). New York: Springer Verlag. Aragno, M., & Schlegel, H. G. (1992). The mesophilic Hydrogen-Oxidizing (Knallgas) Bacteria. In A. Balows, H. Truper, M. Dworkin, W. Harder, & K.-H. Schleifer, The Prokaryotes 2nd. ed. (pp. 344-384). New York: Springer. Ármannson, H. (2002, May 30-31). Erindi á ráðstefnu um málefni veitufyrirtækja . Grænt bókhald í jarðhita- samanburður á útblæstri við aðra orkugjafa . Akureyri, Iceland: Samorka. Bae, S., Kwak, K., Kim, S., Chung, S., & Igarashi, Y. (2001). Isolation and Characterization of CO2-Fixing Hydrogen -Oxidizing Marine 109 University of Akureyri Department of Natural Resource Science Bacteria. -

A Korarchaeal Genome Reveals Insights Into the Evolution of the Archaea
A korarchaeal genome reveals insights into the evolution of the Archaea James G. Elkinsa,b, Mircea Podarc, David E. Grahamd, Kira S. Makarovae, Yuri Wolfe, Lennart Randauf, Brian P. Hedlundg, Ce´ line Brochier-Armaneth, Victor Kunini, Iain Andersoni, Alla Lapidusi, Eugene Goltsmani, Kerrie Barryi, Eugene V. Koonine, Phil Hugenholtzi, Nikos Kyrpidesi, Gerhard Wannerj, Paul Richardsoni, Martin Kellerc, and Karl O. Stettera,k,l aLehrstuhl fu¨r Mikrobiologie und Archaeenzentrum, Universita¨t Regensburg, D-93053 Regensburg, Germany; cBiosciences Division, Oak Ridge National Laboratory, Oak Ridge, TN 37831; dDepartment of Chemistry and Biochemistry, University of Texas, Austin, TX 78712; eNational Center for Biotechnology Information, National Library of Medicine, National Institutes of Health, Bethesda, MD 20894; fDepartment of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520; gSchool of Life Sciences, University of Nevada, Las Vegas, NV 89154; hLaboratoire de Chimie Bacte´rienne, Unite´ Propre de Recherche 9043, Centre National de la Recherche Scientifique, Universite´de Provence Aix-Marseille I, 13331 Marseille Cedex 3, France; iU.S. Department of Energy Joint Genome Institute, Walnut Creek, CA 94598; jInstitute of Botany, Ludwig Maximilians University of Munich, D-80638 Munich, Germany; and kInstitute of Geophysics and Planetary Physics, University of California, Los Angeles, CA 90095 Communicated by Carl R. Woese, University of Illinois at Urbana–Champaign, Urbana, IL, April 2, 2008 (received for review January 7, 2008) The candidate division Korarchaeota comprises a group of uncul- and sediment samples from Obsidian Pool as an inoculum. The tivated microorganisms that, by their small subunit rRNA phylog- cultivation system supported the stable growth of a mixed commu- eny, may have diverged early from the major archaeal phyla nity of hyperthermophilic bacteria and archaea including an or- Crenarchaeota and Euryarchaeota. -

Review Article Diversity of the DNA Replication System in the Archaea Domain
Hindawi Publishing Corporation Archaea Volume 2014, Article ID 675946, 15 pages http://dx.doi.org/10.1155/2014/675946 Review Article Diversity of the DNA Replication System in the Archaea Domain Felipe Sarmiento,1 Feng Long,1 Isaac Cann,2 and William B. Whitman1 1 Department of Microbiology, University of Georgia, 541 Biological Science Building, Athens, GA 30602-2605, USA 2 3408 Institute for Genomic Biology, University of Illinois, 1206 W Gregory Drive, Urbana, IL 61801, USA Correspondence should be addressed to William B. Whitman; [email protected] Received 21 October 2013; Accepted 16 February 2014; Published 26 March 2014 Academic Editor: Yoshizumi Ishino Copyright © 2014 Felipe Sarmiento et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. The precise and timely duplication of the genome is essential for cellular life. It is achieved by DNA replication, a complex process that is conserved among the three domains of life. Even though the cellular structure of archaea closely resembles that of bacteria, the information processing machinery of archaea is evolutionarily more closely related to the eukaryotic system, especially for the proteins involved in the DNA replication process. While the general DNA replication mechanism is conserved among the different domains of life, modifications in functionality and in some of the specialized replication proteins are observed. Indeed, Archaea possess specific features unique to this domain. Moreover, even though the general pattern of the replicative system is the samein all archaea, a great deal of variation exists between specific groups. -

And Thermo-Adaptation in Hyperthermophilic Archaea: Identification of Compatible Solutes, Accumulation Profiles, and Biosynthetic Routes in Archaeoglobus Spp
Universidade Nova de Lisboa Osmo- andInstituto thermo de Tecnologia-adaptation Química e Biológica in hyperthermophilic Archaea: Subtitle Subtitle Luís Pedro Gafeira Gonçalves Osmo- and thermo-adaptation in hyperthermophilic Archaea: identification of compatible solutes, accumulation profiles, and biosynthetic routes in Archaeoglobus spp. OH OH OH CDP c c c - CMP O O - PPi O3P P CTP O O O OH OH OH OH OH OH O- C C C O P O O P i Dissertation presented to obtain the Ph.D degree in BiochemistryO O- Instituto de Tecnologia Química e Biológica | Universidade Nova de LisboaP OH O O OH OH OH Oeiras, Luís Pedro Gafeira Gonçalves January, 2008 2008 Universidade Nova de Lisboa Instituto de Tecnologia Química e Biológica Osmo- and thermo-adaptation in hyperthermophilic Archaea: identification of compatible solutes, accumulation profiles, and biosynthetic routes in Archaeoglobus spp. This dissertation was presented to obtain a Ph. D. degree in Biochemistry at the Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa. By Luís Pedro Gafeira Gonçalves Supervised by Prof. Dr. Helena Santos Oeiras, January, 2008 Apoio financeiro da Fundação para a Ciência e Tecnologia (POCI 2010 – Formação Avançada para a Ciência – Medida IV.3) e FSE no âmbito do Quadro Comunitário de apoio, Bolsa de Doutoramento com a referência SFRH / BD / 5076 / 2001. ii ACKNOWNLEDGMENTS The work presented in this thesis, would not have been possible without the help, in terms of time and knowledge, of many people, to whom I am extremely grateful. Firstly and mostly, I need to thank my supervisor, Prof. Helena Santos, for her way of thinking science, her knowledge, her rigorous criticism, and her commitment to science. -

Textile Waste and Microplastic Induce Activity and Development of Unique
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.08.939876; this version posted February 10, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Textile waste and microplastic induce activity and 2 development of unique hydrocarbon-degrading marine 3 bacterial communities 4 5 Elsa B. Girard1, Melanie Kaliwoda2, Wolfgang W. Schmahl1,2,3, Gert Wörheide1,3,4 and 6 William D. Orsi1,3* 7 8 1 Department of Earth and Environmental Sciences, Ludwig-Maximilians-Universität München, 9 80333 Munich, Germany 10 2 SNSB - Mineralogische Staatssammlung München, 80333 München, Germany 11 3 GeoBio-CenterLMU, Ludwig-Maximilians-Universität München, 80333 Munich, Germany 12 4 SNSB - Bayerische Staatssammlung für Paläontologie und Geologie, 80333 Munich, Germany 13 *Corresponding author (e-mail: [email protected]) 14 15 16 17 18 KEYWORDS 19 Microplastic, Fiber, Hydrocarbon-degrading bacteria, Microbial community, Pollution 20 21 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.02.08.939876; this version posted February 10, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 22 ABSTRACT 23 Biofilm-forming microbial communities on plastics and textile fibers are of growing interest since 24 they have potential to contribute to disease outbreaks and material biodegradability in the 25 environment. -

Corrosion of Carbon Steel by Bacteria from North
Corrosion of carbon steel by bacteria from North Sea offshore seawater injection systems: Laboratory investigation Marko Stipaničev, Florin Turcu, Loïc Esnault, Omar Rosas, Régine Basséguy, Magdalena Sztyler, Iwona B. Beech To cite this version: Marko Stipaničev, Florin Turcu, Loïc Esnault, Omar Rosas, Régine Basséguy, et al.. Corrosion of carbon steel by bacteria from North Sea offshore seawater injection systems: Laboratory investigation. Bioelectrochemistry, Elsevier, 2014, 97, pp.76-88. 10.1016/j.bioelechem.2013.09.006. hal-01913355 HAL Id: hal-01913355 https://hal.archives-ouvertes.fr/hal-01913355 Submitted on 6 Nov 2018 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. OATAO is an open access repository that collects the work of Toulouse researchers and makes it freely available over the web where possible This is an author’s version published in: http://oatao.univ-toulouse.fr/20292 Official URL: https://doi.org/10.1016/j.bioelechem.2013.09.006 To cite this version: Stipaničev, Marko and Turcu, Florin and Esnault, Loïc and Rosas, Omar and Basséguy, Régine and Sztyler, Magdalena and Beech, Iwona B. Corrosion of carbon steel by bacteria from North Sea offshore seawater injection systems: Laboratory investigation.