Rare Coding Variants in ANGPTL6 Are Associated with Familial Forms of Intracranial Aneurysm
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Table S1
Entrez Gene Symbol Gene Name Affymetrix EST Glomchip SAGE Stanford Literature HPA confirmed Gene ID Profiling profiling Profiling Profiling array profiling confirmed 1 2 A2M alpha-2-macroglobulin 0 0 0 1 0 2 10347 ABCA7 ATP-binding cassette, sub-family A (ABC1), member 7 1 0 0 0 0 3 10350 ABCA9 ATP-binding cassette, sub-family A (ABC1), member 9 1 0 0 0 0 4 10057 ABCC5 ATP-binding cassette, sub-family C (CFTR/MRP), member 5 1 0 0 0 0 5 10060 ABCC9 ATP-binding cassette, sub-family C (CFTR/MRP), member 9 1 0 0 0 0 6 79575 ABHD8 abhydrolase domain containing 8 1 0 0 0 0 7 51225 ABI3 ABI gene family, member 3 1 0 1 0 0 8 29 ABR active BCR-related gene 1 0 0 0 0 9 25841 ABTB2 ankyrin repeat and BTB (POZ) domain containing 2 1 0 1 0 0 10 30 ACAA1 acetyl-Coenzyme A acyltransferase 1 (peroxisomal 3-oxoacyl-Coenzyme A thiol 0 1 0 0 0 11 43 ACHE acetylcholinesterase (Yt blood group) 1 0 0 0 0 12 58 ACTA1 actin, alpha 1, skeletal muscle 0 1 0 0 0 13 60 ACTB actin, beta 01000 1 14 71 ACTG1 actin, gamma 1 0 1 0 0 0 15 81 ACTN4 actinin, alpha 4 0 0 1 1 1 10700177 16 10096 ACTR3 ARP3 actin-related protein 3 homolog (yeast) 0 1 0 0 0 17 94 ACVRL1 activin A receptor type II-like 1 1 0 1 0 0 18 8038 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 1 0 0 0 0 19 8751 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 1 0 0 0 0 20 8728 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 1 0 0 0 0 21 81792 ADAMTS12 ADAM metallopeptidase with thrombospondin type 1 motif, 12 1 0 0 0 0 22 9507 ADAMTS4 ADAM metallopeptidase with thrombospondin type 1 -

Noninvasive Sleep Monitoring in Large-Scale Screening of Knock-Out Mice
bioRxiv preprint doi: https://doi.org/10.1101/517680; this version posted January 11, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Noninvasive sleep monitoring in large-scale screening of knock-out mice reveals novel sleep-related genes Shreyas S. Joshi1*, Mansi Sethi1*, Martin Striz1, Neil Cole2, James M. Denegre2, Jennifer Ryan2, Michael E. Lhamon3, Anuj Agarwal3, Steve Murray2, Robert E. Braun2, David W. Fardo4, Vivek Kumar2, Kevin D. Donohue3,5, Sridhar Sunderam6, Elissa J. Chesler2, Karen L. Svenson2, Bruce F. O'Hara1,3 1Dept. of Biology, University of Kentucky, Lexington, KY 40506, USA, 2The Jackson Laboratory, Bar Harbor, ME 04609, USA, 3Signal solutions, LLC, Lexington, KY 40503, USA, 4Dept. of Biostatistics, University of Kentucky, Lexington, KY 40536, USA, 5Dept. of Electrical and Computer Engineering, University of Kentucky, Lexington, KY 40506, USA. 6Dept. of Biomedical Engineering, University of Kentucky, Lexington, KY 40506, USA. *These authors contributed equally Address for correspondence and proofs: Shreyas S. Joshi, Ph.D. Dept. of Biology University of Kentucky 675 Rose Street 101 Morgan Building Lexington, KY 40506 U.S.A. Phone: (859) 257-2805 FAX: (859) 257-1717 Email: [email protected] Running title: Sleep changes in knockout mice bioRxiv preprint doi: https://doi.org/10.1101/517680; this version posted January 11, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Frequent Variations in Cancer-Related Genes May Play Prognostic Role in Treatment of Patients with Chronic Myeloid Leukemia Alexander V
Lavrov et al. BMC Genetics 2016, 17(Suppl 1):14 DOI 10.1186/s12863-015-0308-7 RESEARCH Open Access Frequent variations in cancer-related genes may play prognostic role in treatment of patients with chronic myeloid leukemia Alexander V. Lavrov1,2*, Ekaterina Y. Chelysheva3, Svetlana A. Smirnikhina1, Oleg A. Shukhov3, Anna G. Turkina3, Elmira P. Adilgereeva1 and Sergey I. Kutsev1,2 From The 7th International Young Scientists School “Systems Biology and Bioinformatics” (SBB’2015) Novosibirsk, Russia. 22-25 June 2015 Abstract Background: Genome variability of host genome and cancer cells play critical role in diversity of response to existing therapies and overall success in treating oncological diseases. In chronic myeloid leukemia targeted therapy with tyrosine kinase inhibitors demonstrates high efficacy in most of the patients. However about 15 % of patients demonstrate primary resistance to standard therapy. Whole exome sequencing is a good tool for unbiased search of genetic variations important for prognosis of survival and therapy efficacy in many cancers. We apply this approach to CML patients with optimal response and failure of tyrosine kinase therapy. Results: We analyzed exome variations between optimal responders and failures and found 7 variants in cancer- related genes with different genotypes in two groups of patients. Five of them were found in optimal responders: rs11579366, rs1990236, rs176037, rs10653661, rs3803264 and two in failures: rs3099950, rs9471966. These variants were found in genes associated with cancers (ANKRD35, DNAH9, MAGEC1, TOX3) or participating in cancer-related signaling pathways (THSD1, MORN2, PTCRA). Conclusion: We found gene variants which may become early predictors of the therapy outcome and allow development of new early prognostic tests for estimation of therapy efficacy in CML patients. -

Genomic Abnormalities in Chronic Lymphocytic Leukemia Influence Gene Expression by a Gene Dosage Effect
769-778 24/3/06 12:54 Page 769 INTERNATIONAL JOURNAL OF MOLECULAR MEDICINE 17: 769-778, 2006 769 Genomic abnormalities in chronic lymphocytic leukemia influence gene expression by a gene dosage effect JOHN D. DICKINSON1, AVADHUT JOSHI1, JAVEED IQBAL2, WARREN SANGER3, PHILIP J. BIERMAN4 and SHANTARAM S. JOSHI1 1Department of Genetics, Cell Biology, and Anatomy, 2Department of Pathology and Microbiology, 3Center for Human Genetics, 4Oncology/Hematology Section, Department of Internal Medicine, Center for Research in Leukemia and Lymphoma, University of Nebraska Medical Center, Omaha, NE, USA Received November 16, 2005; Accepted December 21, 2005 Abstract. This work describes the identification and impact lymphocytes and is identified by co-expression of CD5, 19, of somatic genomic abnormalities in human chronic lympho- and 23 cell surface markers (3,4). CLL has been characterized cytic leukemia (CLL). Using molecular cytogenetics (FISH) by broad clinical and biological heterogeneity and not all and G-banding cytogenetic analysis, chromosome abnormalities patients require immediate therapy (3,5,6). Much of the were detected in 37 of 46 (80.4%) CLL patients. 13q14 research in the area of CLL has focused on identifying prog- deletion was the most common finding followed by trisomy nostic factors that can predict which patients will ultimately 12 and 11q22.3 deletion. 17p13 deletion was also detected as undergo tumor progression and require therapy. In addition, were several less frequent chromosome abnormalities. The elucidating the mechanisms of tumor progression in order to presence of these abnormalities significantly influenced the develop new molecular therapies and adjuvant therapies to period of treatment-free survival as well as other clinical induce long-term remissions is an important goal of CLL characteristics. -

Supplementary Table 1 Double Treatment Vs Single Treatment
Supplementary table 1 Double treatment vs single treatment Probe ID Symbol Gene name P value Fold change TC0500007292.hg.1 NIM1K NIM1 serine/threonine protein kinase 1.05E-04 5.02 HTA2-neg-47424007_st NA NA 3.44E-03 4.11 HTA2-pos-3475282_st NA NA 3.30E-03 3.24 TC0X00007013.hg.1 MPC1L mitochondrial pyruvate carrier 1-like 5.22E-03 3.21 TC0200010447.hg.1 CASP8 caspase 8, apoptosis-related cysteine peptidase 3.54E-03 2.46 TC0400008390.hg.1 LRIT3 leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 1.86E-03 2.41 TC1700011905.hg.1 DNAH17 dynein, axonemal, heavy chain 17 1.81E-04 2.40 TC0600012064.hg.1 GCM1 glial cells missing homolog 1 (Drosophila) 2.81E-03 2.39 TC0100015789.hg.1 POGZ Transcript Identified by AceView, Entrez Gene ID(s) 23126 3.64E-04 2.38 TC1300010039.hg.1 NEK5 NIMA-related kinase 5 3.39E-03 2.36 TC0900008222.hg.1 STX17 syntaxin 17 1.08E-03 2.29 TC1700012355.hg.1 KRBA2 KRAB-A domain containing 2 5.98E-03 2.28 HTA2-neg-47424044_st NA NA 5.94E-03 2.24 HTA2-neg-47424360_st NA NA 2.12E-03 2.22 TC0800010802.hg.1 C8orf89 chromosome 8 open reading frame 89 6.51E-04 2.20 TC1500010745.hg.1 POLR2M polymerase (RNA) II (DNA directed) polypeptide M 5.19E-03 2.20 TC1500007409.hg.1 GCNT3 glucosaminyl (N-acetyl) transferase 3, mucin type 6.48E-03 2.17 TC2200007132.hg.1 RFPL3 ret finger protein-like 3 5.91E-05 2.17 HTA2-neg-47424024_st NA NA 2.45E-03 2.16 TC0200010474.hg.1 KIAA2012 KIAA2012 5.20E-03 2.16 TC1100007216.hg.1 PRRG4 proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) 7.43E-03 2.15 TC0400012977.hg.1 SH3D19 -

High-Density Array Comparative Genomic Hybridization Detects Novel Copy Number Alterations in Gastric Adenocarcinoma
ANTICANCER RESEARCH 34: 6405-6416 (2014) High-density Array Comparative Genomic Hybridization Detects Novel Copy Number Alterations in Gastric Adenocarcinoma ALINE DAMASCENO SEABRA1,2*, TAÍSSA MAÍRA THOMAZ ARAÚJO1,2*, FERNANDO AUGUSTO RODRIGUES MELLO JUNIOR1,2, DIEGO DI FELIPE ÁVILA ALCÂNTARA1,2, AMANDA PAIVA DE BARROS1,2, PAULO PIMENTEL DE ASSUMPÇÃO2, RAQUEL CARVALHO MONTENEGRO1,2, ADRIANA COSTA GUIMARÃES1,2, SAMIA DEMACHKI2, ROMMEL MARIO RODRÍGUEZ BURBANO1,2 and ANDRÉ SALIM KHAYAT1,2 1Human Cytogenetics Laboratory and 2Oncology Research Center, Federal University of Pará, Belém Pará, Brazil Abstract. Aim: To investigate frequent quantitative alterations gastric cancer is the second most frequent cancer in men and of intestinal-type gastric adenocarcinoma. Materials and the third in women (4). The state of Pará has a high Methods: We analyzed genome-wide DNA copy numbers of 22 incidence of gastric adenocarcinoma and this disease is a samples and using CytoScan® HD Array. Results: We identified public health problem, since mortality rates are above the 22 gene alterations that to the best of our knowledge have not Brazilian average (5). been described for gastric cancer, including of v-erb-b2 avian This tumor can be classified into two histological types, erythroblastic leukemia viral oncogene homolog 4 (ERBB4), intestinal and diffuse, according to Laurén (4, 6, 7). The SRY (sex determining region Y)-box 6 (SOX6), regulator of intestinal type predominates in high-risk areas, such as telomere elongation helicase 1 (RTEL1) and UDP- Brazil, and arises from precursor lesions, whereas the diffuse Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 5 type has a similar distribution in high- and low-risk areas and (B4GALT5). -
Downloaded from the Mouse Lysosome Gene Database, Mlgdb
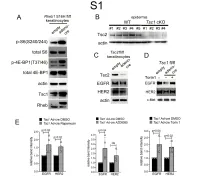
1 Supplemental Figure Legends 2 3 Supplemental Figure S1: Epidermal-specific mTORC1 gain-of-function models show 4 increased mTORC1 activation and down-regulate EGFR and HER2 protein expression in a 5 mTORC1-sensitive manner. (A) Immunoblotting of Rheb1 S16H flox/flox keratinocyte cultures 6 infected with empty or adenoviral cre recombinase for markers of mTORC1 (p-S6, p-4E-BP1) 7 activity. (B) Tsc1 cKO epidermal lysates also show decreased expression of TSC2 by 8 immunoblotting of the same experiment as in Figure 2A. (C) Immunoblotting of Tsc2 flox/flox 9 keratinocyte cultures infected with empty or adenoviral cre recombinase showing decreased EGFR 10 and HER2 protein expression. (D) Expression of EGFR and HER2 was decreased in Tsc1 cre 11 keratinocytes compared to empty controls, and up-regulated in response to Torin1 (1µM, 24 hrs), 12 by immunoblot analyses. Immunoblots are contemporaneous and parallel from the same biological 13 replicate and represent the same experiment as depicted in Figure 7B. (E) Densitometry 14 quantification of representative immunoblot experiments shown in Figures 2E and S1D (r≥3; error 15 bars represent STDEV; p-values by Student’s T-test). 16 17 18 19 20 21 22 23 Supplemental Figure S2: EGFR and HER2 transcription are unchanged with epidermal/ 24 keratinocyte Tsc1 or Rptor loss. Egfr and Her2 mRNA levels in (A) Tsc1 cKO epidermal lysates, 25 (B) Tsc1 cKO keratinocyte lysates and(C) Tsc1 cre keratinocyte lysates are minimally altered 26 compared to their respective controls. (r≥3; error bars represent STDEV; p-values by Student’s T- 27 test). -

Microrna-15A and -16-1 Act Via MYB to Elevate Fetal Hemoglobin Expression in Human Trisomy 13
MicroRNA-15a and -16-1 act via MYB to elevate fetal hemoglobin expression in human trisomy 13 Vijay G. Sankarana,b,c,d, Tobias F. Mennee,f, Danilo Šcepanovicg, Jo-Anne Vergilioh, Peng Jia, Jinkuk Kima,g,i, Prathapan Thirua, Stuart H. Orkind,e,f,j, Eric S. Landera,b,k,l, and Harvey F. Lodisha,b,l,1 aWhitehead Institute for Biomedical Research, Cambridge, MA 02142; bBroad Institute of Massachusetts Institute of Technology and Harvard University, Cambridge, MA 02142; Departments of cMedicine and hPathology and eDivision of Hematology/Oncology, Children’s Hospital Boston, Boston, MA 02115; Departments of dPediatrics and kSystems Biology, Harvard Medical School, Boston, MA 02115; lDepartment of Biology and iHoward Hughes Medical Institute, Massachusetts Institute of Technology, Cambridge, MA 02142; gHarvard–Massachusetts Institute of Technology Division of Health Sciences and Technology, Cambridge, MA 02142; fDepartment of Pediatric Oncology, Dana-Farber Cancer Institute, Boston, MA 02115; and jThe Howard Hughes Medical Institute, Boston, MA 02115 Contributed by Harvey F. Lodish, December 13, 2010 (sent for review November 8, 2010) Many human aneuploidy syndromes have unique phenotypic other hematopoietic cells shifts to the bone marrow, the pre- consequences, but in most instances it is unclear whether these dominant postnatal site for hematopoiesis, another switch phenotypes are attributable to alterations in the dosage of specific occurs, resulting in down-regulation of γ-globin and concomitant genes. In human trisomy 13, there is delayed switching and up-regulation of the adult β-globin gene (6, 8, 10). There is persistence of fetal hemoglobin (HbF) and elevation of embryonic a limited understanding of the molecular control of these globin hemoglobin in newborns. -

Associations Between Genetically Predicted Blood Protein Biomarkers and Pancreatic 2 Cancer Risk 3
Author Manuscript Published OnlineFirst on May 21, 2020; DOI: 10.1158/1055-9965.EPI-20-0091 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Zhu et al. – Page 1 1 Associations between genetically predicted blood protein biomarkers and pancreatic 2 cancer risk 3 4 Jingjing Zhu 1, Xiang Shu 2, Xingyi Guo 2, Duo Liu 1,3, Jiandong Bao 2, Roger L Milne 4, 5, 6, 5 Graham G Giles 4, 5, 6, Chong Wu 7, Mengmeng Du 8, Emily White 9,10, Harvey A Risch11, Nuria 6 Malats 12, Eric J. Duell 13, Phyllis J. Goodman 14, Donghui Li 15, Paige Bracci 16, Verena Katzke 7 17, Rachel E Neale 18, Steven Gallinger 19, Stephen K Van Den Eeden 20, Alan A Arslan 21, 8 Federico Canzian 22, Charles Kooperberg 9, Laura E Beane-Freeman 23, Ghislaine Scelo 24, Kala 9 Visvanathan25, Christopher A. Haiman 26, Loïc Le Marchand 1, Herbert Yu 1, Gloria M Petersen 10 27, Rachael Stolzenberg-Solomon 23, Alison P Klein 25,28, Qiuyin Cai 2, Jirong Long 2, Xiao-Ou 11 Shu 2, Wei Zheng 2, Lang Wu 1 12 13 1. Cancer Epidemiology Division, Population Sciences in the Pacific Program, University of 14 Hawaii Cancer Center, University of Hawaii at Manoa, Honolulu, HI, USA 15 2. Division of Epidemiology, Department of Medicine, Vanderbilt Epidemiology Center, 16 Vanderbilt-Ingram Cancer Center, Vanderbilt University Medical Center, Nashville, TN, USA 17 3. Department of Pharmacy, Harbin Medical University Cancer Hospital, Harbin, China 18 4. Cancer Epidemiology Division, Cancer Council Victoria, Melbourne, VIC, Australia 19 5. -

Multi-Transcriptomic Analysis Points to Early Organelle Dysfunction in Human Astrocytes in Alzheimer’S Disease
Multi-Transcriptomic Analysis Points to Early Organelle Dysfunction in Human Astrocytes in Alzheimer’s Disease Elena Galea ( [email protected] ) Universitat Autonoma de Barcelona https://orcid.org/0000-0003-4537-9897 Laura D. Weinstock Georgia Tech: Georgia Institute of Technology Raquel Larramona-Arcas Universitat Autonoma de Barcelona Alyssa F. Pybus Georgia Tech: Georgia Institute of Technology Lydia Giménez-Llort Universitat Autonoma de Barcelona Carole Escartin Universite Paris-Saclay Levi B. Wood Georgia Tech: Georgia Institute of Technology Research Article Keywords: Alzheimer’s disease, astrocytes, hierarchical clustering, MCI, mitochondria, perisynaptic astrocyte processes, RNA seq, gene enrichment scores, transcriptomics. Posted Date: June 25th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-426597/v2 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/34 Abstract Background Recent gene proling of human AD astrocytes using single-nucleus RNA sequencing is limited by the low number of differentially expressed genes detected, and the small size of cohorts. We improved on prior studies with a novel systems-biology-based approach that can be used with available data from large cohorts. Methods Brain-cell specic gene clusters were generated from RNA sequencing data from isolated healthy human brain cells using a cell-type enrichment score and clustering. The cell-specic gene clusters were localized in whole-brain transcriptomes from 766 subjects diagnosed with AD or mild cognitive impairment (MCI) by the Mount Sinai Hospital, the Mayo Clinic, and the Religious Order Study/Memory and Aging Project (ROSMAP). Gene clusters were organized into functional categories, and changes among subject groups determined by gene set variation analysis (GSVA) and principal component analysis (PCA). -

Stringdb Package Vignette
STRINGdb Package Vignette Andrea Franceschini 15 March 2015 1 INTRODUCTION STRING (https://www.string-db.org) is a database of known and predicted protein-protein interac- tions. The interactions include direct (physical) and indirect (functional) associations. The database contains information from numerous sources, including experimental repositories, computational pre- diction methods and public text collections. Each interaction is associated with a combined confidence score that integrates the various evidences. We currently cover over 24 milions proteins from 5090 organisms. As you will learn in this guide, the STRING database can be usefull to add meaning to list of genes (e.g. the best hits coming out from a screen or the most differentially expressed genes coming out from a Microarray/RNAseq experiment.) We provide the STRINGdb R package in order to facilitate our users in accessing the STRING database from R. In this guide we explain, with examples, most of the package's features and function- alities. In the STRINGdb R package we use the new ReferenceClasses of R (search for "ReferenceClasses" in the R documentation.). Besides we make use of the iGraph package (http://igraph.sourceforge.net) as a data structure to represent our protein-protein interaction network. To begin, you should first know the NCBI taxonomy identifiers of the organism on which you have performed the experiment (e.g. 9606 for Human, 10090 for mouse). If you don't know that, you can search the NCBI Taxonomy (http://www.ncbi.nlm.nih.gov/taxonomy) or start looking at our species table (that you can also use to verify that your organism is represented in the STRING database). -

The Glomerular Transcriptome and a Predicted Protein–Protein Interaction Network
BASIC RESEARCH www.jasn.org The Glomerular Transcriptome and a Predicted Protein–Protein Interaction Network Liqun He,* Ying Sun,* Minoru Takemoto,* Jenny Norlin,* Karl Tryggvason,* Tore Samuelsson,† and Christer Betsholtz*‡ *Division of Matrix Biology, Department of Medical Biochemistry and Biophysics, and ‡Department of Medicine, Karolinska Institutet, Stockholm, and †Department of Medical Biochemistry, Go¨teborg University, Go¨teborg, Sweden ABSTRACT To increase our understanding of the molecular composition of the kidney glomerulus, we performed a meta-analysis of available glomerular transcriptional profiles made from mouse and man using five different methodologies. We generated a combined catalogue of glomerulus-enriched genes that emerged from these different sources and then used this to construct a predicted protein–protein interaction network in the glomerulus (GlomNet). The combined glomerulus-enriched gene catalogue provides the most comprehensive picture of the molecular composition of the glomerulus currently available, and GlomNet contributes an integrative systems biology approach to the understanding of glomerular signaling networks that operate during development, function, and disease. J Am Soc Nephrol 19: 260–268, 2008. doi: 10.1681/ASN.2007050588 Many kidney diseases and, importantly, approxi- nins have been shown to be mutated in Alport syn- mately two thirds of all cases of ESRD originate with drome and Pierson congenital nephrotic syndromes, glomerular disease. Most cases of glomerular disease respectively.11,12 Genetic studies in mice have further are caused by systemic disorders (e.g., diabetes, hyper- revealed genes and proteins of importance for glomer- tension, lupus, obesity) for which the molecular ulus development and function, such as podoca- pathogeneses of the glomerular complications are un- lyxin,13 CD2AP,14 NEPH1,15 FAT1,16 forkhead box known.