Biochemical Analysis of the Mrna Scavenger Decapping Enzymes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mutations in DCPS and EDC3 in Autosomal Recessive Intellectual
Human Molecular Genetics, 2015, Vol. 24, No. 11 3172–3180 doi: 10.1093/hmg/ddv069 Advance Access Publication Date: 20 February 2015 Original Article Downloaded from ORIGINAL ARTICLE Mutations in DCPS and EDC3 in autosomal recessive intellectual disability indicate a crucial role for mRNA http://hmg.oxfordjournals.org/ decapping in neurodevelopment Iltaf Ahmed1,2,†, Rebecca Buchert3,†, Mi Zhou5,†, Xinfu Jiao5,†, Kirti Mittal1, Taimoor I. Sheikh1, Ute Scheller3, Nasim Vasli1, Muhammad Arshad Rafiq1, 6 1 7 2 M. Qasim Brohi , Anna Mikhailov , Muhammad Ayaz , Attya Bhatti , at Universitaet Erlangen-Nuernberg, Wirtschafts- und Sozialwissenschaftliche Z on August 15, 2016 Heinrich Sticht4, Tanveer Nasr8,9, Melissa T. Carter10, Steffen Uebe3, André Reis3, Muhammad Ayub7,11, Peter John2, Megerditch Kiledjian5,*, John B. Vincent1,12,13,* and Rami Abou Jamra3,* 1Molecular Neuropsychiatry and Development Lab, Campbell Family Mental Health Research Institute, Centre for Addiction and Mental Health, 250 College Street, Toronto, Ontario, Canada M5T 1R8, 2Atta-ur-Rehman School of Applied Biosciences (ASAB), National University of Sciences and Technology (NUST), Islamabad 44000, Pakistan, 3Institute of Human Genetics and 4Bioinformatics, Institute of Biochemistry, Friedrich-Alexander-Universität Erlangen-Nürnberg, Erlangen 91054, Germany, 5Department of Cell Biology and Neuroscience, Rutgers University, Piscataway, NJ 08854, USA, 6Sir Cowasji Jehangir Institute of Psychiatry, Hyderabad, Sindh 71000, Pakistan, 7Lahore Institute of Research and Development, -

Dcps Is a Transcript-Specific Modulator of RNA in Mammalian Cells
Downloaded from rnajournal.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press DcpS is a transcript-specific modulator of RNA in mammalian cells MI ZHOU,1 SOPHIE BAIL,1 HEATHER L. PLASTERER,2 JAMES RUSCHE,2 and MEGERDITCH KILEDJIAN1 1Department of Cell Biology and Neuroscience, Rutgers University, Piscataway, New Jersey 08854, USA 2Repligen Corporation, Waltham, Massachusetts 02453, USA ABSTRACT The scavenger decapping enzyme DcpS is a multifunctional protein initially identified by its property to hydrolyze the resulting cap structure following 3′ end mRNA decay. In Saccharomyces cerevisiae, the DcpS homolog Dcs1 is an obligate cofactor for the 5′-3′ exoribonuclease Xrn1 while the Caenorhabditis elegans homolog Dcs-1, facilitates Xrn1 mediated microRNA turnover. In both cases, this function is independent of the decapping activity. Whether DcpS and its decapping activity can affect mRNA steady state or stability in mammalian cells remains unknown. We sought to determine DcpS target genes in mammalian cells using a cell-permeable DcpS inhibitor compound, RG3039 initially developed for therapeutic treatment of spinal muscular atrophy. Global mRNA levels were examined following DcpS decapping inhibition with RG3039. The steady-state levels of 222 RNAs were altered upon RG3039 treatment. Of a subset selected for validation, two transcripts that appear to be long noncoding RNAs HS370762 and BC011766, were dependent on DcpS and its scavenger decapping catalytic activity and referred to as DcpS-responsive noncoding transcripts (DRNT) 1 and 2, respectively. Interestingly, only the increase in DRNT1 transcript was accompanied with an increase of its RNA stability and this increase was dependent on both DcpS and Xrn1. -

Coupling of Spliceosome Complexity to Intron Diversity
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.19.436190; this version posted March 20, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Coupling of spliceosome complexity to intron diversity Jade Sales-Lee1, Daniela S. Perry1, Bradley A. Bowser2, Jolene K. Diedrich3, Beiduo Rao1, Irene Beusch1, John R. Yates III3, Scott W. Roy4,6, and Hiten D. Madhani1,6,7 1Dept. of Biochemistry and Biophysics University of California – San Francisco San Francisco, CA 94158 2Dept. of Molecular and Cellular Biology University of California - Merced Merced, CA 95343 3Department of Molecular Medicine The Scripps Research Institute, La Jolla, CA 92037 4Dept. of Biology San Francisco State University San Francisco, CA 94132 5Chan-Zuckerberg Biohub San Francisco, CA 94158 6Corresponding authors: [email protected], [email protected] 7Lead Contact 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.19.436190; this version posted March 20, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. SUMMARY We determined that over 40 spliceosomal proteins are conserved between many fungal species and humans but were lost during the evolution of S. cerevisiae, an intron-poor yeast with unusually rigid splicing signals. We analyzed null mutations in a subset of these factors, most of which had not been investigated previously, in the intron-rich yeast Cryptococcus neoformans. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Mrna Turnover Philip Mitchell* and David Tollervey†
320 mRNA turnover Philip Mitchell* and David Tollervey† Nuclear RNA-binding proteins can record pre-mRNA are cotransported to the cytoplasm with the mRNP. These processing events in the structure of messenger proteins may preserve a record of the nuclear history of the ribonucleoprotein particles (mRNPs). During initial rounds of pre-mRNA in the cytoplasmic mRNP structure. This infor- translation, the mature mRNP structure is established and is mation can strongly influence the cytoplasmic fate of the monitored by mRNA surveillance systems. Competition for the mRNA and is used by mRNA surveillance systems that act cap structure links translation and subsequent mRNA as a checkpoint of mRNP integrity, particularly in the identi- degradation, which may also involve multiple deadenylases. fication of premature translation termination codons (PTCs). Addresses Cotransport of nuclear mRNA-binding proteins with mRNA Wellcome Trust Centre for Cell Biology, ICMB, University of Edinburgh, from the nucleus to the cytoplasm (nucleocytoplasmic shut- Kings’ Buildings, Edinburgh EH9 3JR, UK tling) was first observed for the heterogeneous nuclear *e-mail: [email protected] ribonucleoprotein (hnRNP) proteins. Some hnRNP proteins †e-mail: [email protected] are stripped from the mRNA at export [1], but hnRNP A1, Current Opinion in Cell Biology 2001, 13:320–325 A2, E, I and K are all exported (see [2]). Although roles for 0955-0674/01/$ — see front matter these hnRNP proteins in transport and translation have been © 2001 Elsevier Science Ltd. All rights reserved. reported [3•,4•], their affects on mRNA stability have been little studied. More is known about hnRNP D/AUF1 and Abbreviations AREs AU-rich sequence elements another nuclear RNA-binding protein, HuR, which act CBC cap-binding complex antagonistically to modulate the stability of a range of DAN deadenylating nuclease mRNAs containing AU-rich sequence elements (AREs) DSEs downstream sequence elements (reviewed in [2]). -

RNA Components of the Spliceosome Regulate Tissue- and Cancer-Specific Alternative Splicing
Downloaded from genome.cshlp.org on September 29, 2021 - Published by Cold Spring Harbor Laboratory Press Research RNA components of the spliceosome regulate tissue- and cancer-specific alternative splicing Heidi Dvinge,1,2,4 Jamie Guenthoer,3 Peggy L. Porter,3 and Robert K. Bradley1,2 1Computational Biology Program, Public Health Sciences Division, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109, USA; 2Basic Sciences Division, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109, USA; 3Human Biology Division, Fred Hutchinson Cancer Research Center, Seattle, Washington 98109, USA Alternative splicing of pre-mRNAs plays a pivotal role during the establishment and maintenance of human cell types. Characterizing the trans-acting regulatory proteins that control alternative splicing has therefore been the focus of much research. Recent work has established that even core protein components of the spliceosome, which are required for splicing to proceed, can nonetheless contribute to splicing regulation by modulating splice site choice. We here show that the RNA components of the spliceosome likewise influence alternative splicing decisions. Although these small nuclear RNAs (snRNAs), termed U1, U2, U4, U5, and U6 snRNA, are present in equal stoichiometry within the spliceosome, we found that their relative levels vary by an order of magnitude during development, across tissues, and across cancer samples. Physiologically relevant perturbation of individual snRNAs drove widespread gene-specific differences in alternative splic- ing but not transcriptome-wide splicing failure. Genes that were particularly sensitive to variations in snRNA abundance in a breast cancer cell line model were likewise preferentially misspliced within a clinically diverse cohort of invasive breast ductal carcinomas. -

Comprehensive Protein Interactome Analysis of a Key RNA Helicase: Detection of Novel Stress Granule Proteins
Biomolecules 2015, 5, 1441-1466; doi:10.3390/biom5031441 OPEN ACCESS biomolecules ISSN 2218-273X www.mdpi.com/journal/biomolecules/ Article Comprehensive Protein Interactome Analysis of a Key RNA Helicase: Detection of Novel Stress Granule Proteins Rebecca Bish 1,†, Nerea Cuevas-Polo 1,†, Zhe Cheng 1, Dolores Hambardzumyan 2, Mathias Munschauer 3, Markus Landthaler 3 and Christine Vogel 1,* 1 Center for Genomics and Systems Biology, Department of Biology, New York University, 12 Waverly Place, New York, NY 10003, USA; E-Mails: [email protected] (R.B.); [email protected] (N.C.-P.); [email protected] (Z.C.) 2 The Cleveland Clinic, Department of Neurosciences, Lerner Research Institute, 9500 Euclid Avenue, Cleveland, OH 44195, USA; E-Mail: [email protected] 3 RNA Biology and Post-Transcriptional Regulation, Max-Delbrück-Center for Molecular Medicine, Berlin-Buch, Robert-Rössle-Str. 10, Berlin 13092, Germany; E-Mails: [email protected] (M.M.); [email protected] (M.L.) † These authors contributed equally to this work. * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +1-212-998-3976; Fax: +1-212-995-4015. Academic Editor: André P. Gerber Received: 10 May 2015 / Accepted: 15 June 2015 / Published: 15 July 2015 Abstract: DDX6 (p54/RCK) is a human RNA helicase with central roles in mRNA decay and translation repression. To help our understanding of how DDX6 performs these multiple functions, we conducted the first unbiased, large-scale study to map the DDX6-centric protein-protein interactome using immunoprecipitation and mass spectrometry. Using DDX6 as bait, we identify a high-confidence and high-quality set of protein interaction partners which are enriched for functions in RNA metabolism and ribosomal proteins. -

Identification of Enhancer of Mrna Decapping 4 As a Novel Fusion Partner of MLL in Acute Myeloid Leukemia
STIMULUS REPORT Identification of enhancer of mRNA decapping 4 as a novel fusion partner of MLL in acute myeloid leukemia Heiko Becker,1-3 Gabriele Greve,1,2 Keisuke Kataoka,4 Jan-Philipp Mallm,5,6 Jesus´ Duque-Afonso,1,2,7 Tobias Ma,1,2 Christoph Niemoller,¨ 1,2 Milena Pantic,1 Justus Duyster,1-3 Michael L. Cleary,7 Julia Schuler,¨ 8 Karsten Rippe,5,6 Seishi Ogawa,3 and Michael Lubbert¨ 1-3 1Department of Medicine I, Medical Center, and 2Faculty of Medicine, University of Freiburg, Freiburg, Germany; 3German Cancer Consortium partner site, Freiburg, Germany; 4Department of Pathology and Tumor Biology, Kyoto University, Kyoto, Japan; 5Division of Chromatin Networks and 6Single-cell Open Laboratory, German Cancer Research Center, Heidelberg, Germany; 7Department of Pathology, Stanford University, Stanford, CA; and 8Charles River Discovery Research Services Germany GmbH, Freiburg, Germany Key Points Introduction • mRNA decapping gene Translocations involving MLL (aka KMT2A) located on chromosome 11q23 occur in acute myeloid EDC4 is a novel fusion leukemia (AML) and lymphoblastic leukemia. In AML, they generally confer an adverse prognosis, unless MLL partner of in AML. the MLLT3 (aka AF9) gene is involved.1 More than 130 different translocation partner genes (TPGs) MLL 2 • Genes functioning in have been identified, forming the recombinome. mRNA decapping may Recently, the scavenger messenger RNA (mRNA) decapping enzyme DCPS has been identified to be compose a distinct required for survival of AML cells, but not normal hematopoietic cells, and a DCPS inhibitor showed antileukemic activity.3,4 DCPS is also 1 of 2 genes (the other being DCP1A) involved in mRNA group of MLL fusion decapping and having been described as TPG of MLL in single leukemia cases.5-7 partners that links MLL MLL function with mRNA Here, we describe a novel fusion with another mRNA decapping component, ie, the enhancer of mRNA decapping 4 gene (EDC4;alsoknownasGE1 or HEDLS), in AML. -

Messenger RNA Reprogramming by Spliceosome-Mediated RNA Trans-Splicing
Messenger RNA reprogramming by spliceosome-mediated RNA trans-splicing Mariano A. Garcia-Blanco J Clin Invest. 2003;112(4):474-480. https://doi.org/10.1172/JCI19462. Perspective Series In the human genome, the majority of protein-encoding genes are interrupted by introns, which are removed from primary transcripts by a macromolecular enzyme known as the spliceosome. Spliceosomes can constitutively remove all the introns in a primary transcript to yield a fully spliced mRNA or alternatively splice primary transcripts leading to the production of many different mRNAs from one gene. This review examines how spliceosomes can recombine two primary transcripts in trans to reprogram messenger RNAs. Find the latest version: https://jci.me/19462/pdf PERSPECTIVE SERIES Genetic repair | Bruce A. Sullenger, Series Editor Messenger RNA reprogramming by spliceosome-mediated RNA trans-splicing Mariano A. Garcia-Blanco Department of Molecular Genetics and Microbiology and Department of Medicine, Duke University Medical Center, Durham, North Carolina, USA In the human genome, the majority of protein-encoding genes are interrupted by introns, which are removed from primary transcripts by a macromolecular enzyme known as the spliceosome. Spliceo- somes can constitutively remove all the introns in a primary transcript to yield a fully spliced mRNA or alternatively splice primary transcripts leading to the production of many different mRNAs from one gene. This review examines how spliceosomes can recombine two primary transcripts in trans to reprogram messenger RNAs. J. Clin. Invest. 112:474–480 (2003). doi:10.1172/JCI200319462. Reprogramming of mRNA mediated RNA trans-splicing (SMaRT) has been used The reprogramming of mRNA is a form of gene ther- to reprogram mRNAs in animal cells in culture, in apy that modifies mRNA without directly changing xenografts, and in animals (9–13). -
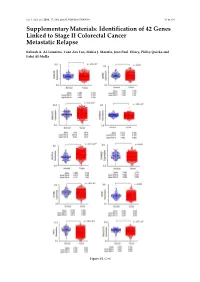
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

The Natural Antisense Transcript NATTD Regulates the Transcription
International Journal of Biochemistry and Cell Biology 110 (2019) 103–110 Contents lists available at ScienceDirect International Journal of Biochemistry and Cell Biology journal homepage: www.elsevier.com/locate/biocel The natural antisense transcript NATTD regulates the transcription of decapping scavenger (DcpS) enzyme T ⁎ Zhu-Zhong Mei , Hongwei Sun, Xiaoli Ou, Lei Li, Junwei Cai, Shuiwang Hu, Juan Wang, ⁎ ⁎ Haihua Luo, Jinghua Liu , Yong Jiang From Guangdong Provincial Key Laboratory of Proteomics, State Key Laboratory of Organ Failure Research, Department of Pathophysiology, Southern Medical University, Guangzhou 510515, China ARTICLE INFO ABSTRACT Keywords: Natural antisense transcripts (NATs) are transcribed from the opposite strand of other genes. Most of them are Natural antisense transcript noncoding RNAs. They have been reported to play important roles in a variety of biological processes. In this NATTD study, we identified a novel NAT, NATTD, which is partially complementary to both the TIRAP/Mal and DcpS DcpS genes. Interestingly, NATTD only positively regulates the expression of DcpS, a decapping scavenger enzyme Epigenetic regulation which is a promising therapeutic target for spinal muscular atrophy. But it has no obvious effects on the ex- pression of TIRAP/Mal gene. The NATTD transcript primarily resides in the nucleus and does not alter the mRNA stability of DcpS. Instead, it is required for the recruitment of RNA polymerase II at the mouse DcpS promoter. Chromatin immunoprecipitation assays revealed that knocking-down -

Role of Dcps in Mammalian RNA Regulation and Human Diseases
ROLE OF DCPS IN MAMMALIAN RNA REGULATION AND HUMAN DISEASES By MI ZHOU A dissertation submitted to the Graduate School-New Brunswick and The Graduate School of Biomedical Sciences Rutgers, The State University of New Jersey In partial fulfillment of the requirements For the degree of Doctor of Philosophy Graduate Program in Cell and Development Biology Written under the direction of Dr. Megerditch Kiledjian And approved by _________________________________ _________________________________ _________________________________ _________________________________ New Brunswick, New Jersey October, 2015 ABSTRACT OF THE DISSERTATION Role of DcpS in Mammalian RNA Regulation and Human Diseases By MI ZHOU Dissertation Director Dr. Megerditch Kiledjian In eukaryotic cells, mRNA degradation plays an important role in the control of gene expression and is therefore highly regulated. The scavenger decapping enzyme DcpS is a multifunctional protein that plays a critical role in mRNA degradation. We first sought to identify DcpS target genes in mammalian cells using a cell permeable DcpS inhibitor compound, RG3039, which was initially developed for therapeutic treatment of Spinal Muscular Atrophy (SMA). Microarray analysis following DcpS decapping inhibition by RG3039 revealed the steady state levels of 222 RNAs were altered. Of a subset selected for validation by qRT-PCR, two non-coding transcripts dependent on DcpS decapping activity, were identified and referred to as DcpS Responsive Noncoding Transcript (DRNT) 1 and 2 respectively. Only the increase in DRNT1 transcript was accompanied with an increase of its RNA stability and this increase was dependent on both DcpS and Xrn1. Our data indicate that DcpS is a transcript-restricted modulator of RNA stability in mammalian cells and the RG3039 ii quinazoline compound is pleotropic, influence gene expression in both an apparent DcpS dependent and independent manner.