Mutations in DCPS and EDC3 in Autosomal Recessive Intellectual
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Dcps Is a Transcript-Specific Modulator of RNA in Mammalian Cells
Downloaded from rnajournal.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press DcpS is a transcript-specific modulator of RNA in mammalian cells MI ZHOU,1 SOPHIE BAIL,1 HEATHER L. PLASTERER,2 JAMES RUSCHE,2 and MEGERDITCH KILEDJIAN1 1Department of Cell Biology and Neuroscience, Rutgers University, Piscataway, New Jersey 08854, USA 2Repligen Corporation, Waltham, Massachusetts 02453, USA ABSTRACT The scavenger decapping enzyme DcpS is a multifunctional protein initially identified by its property to hydrolyze the resulting cap structure following 3′ end mRNA decay. In Saccharomyces cerevisiae, the DcpS homolog Dcs1 is an obligate cofactor for the 5′-3′ exoribonuclease Xrn1 while the Caenorhabditis elegans homolog Dcs-1, facilitates Xrn1 mediated microRNA turnover. In both cases, this function is independent of the decapping activity. Whether DcpS and its decapping activity can affect mRNA steady state or stability in mammalian cells remains unknown. We sought to determine DcpS target genes in mammalian cells using a cell-permeable DcpS inhibitor compound, RG3039 initially developed for therapeutic treatment of spinal muscular atrophy. Global mRNA levels were examined following DcpS decapping inhibition with RG3039. The steady-state levels of 222 RNAs were altered upon RG3039 treatment. Of a subset selected for validation, two transcripts that appear to be long noncoding RNAs HS370762 and BC011766, were dependent on DcpS and its scavenger decapping catalytic activity and referred to as DcpS-responsive noncoding transcripts (DRNT) 1 and 2, respectively. Interestingly, only the increase in DRNT1 transcript was accompanied with an increase of its RNA stability and this increase was dependent on both DcpS and Xrn1. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

1 AGING Supplementary Table 2
SUPPLEMENTARY TABLES Supplementary Table 1. Details of the eight domain chains of KIAA0101. Serial IDENTITY MAX IN COMP- INTERFACE ID POSITION RESOLUTION EXPERIMENT TYPE number START STOP SCORE IDENTITY LEX WITH CAVITY A 4D2G_D 52 - 69 52 69 100 100 2.65 Å PCNA X-RAY DIFFRACTION √ B 4D2G_E 52 - 69 52 69 100 100 2.65 Å PCNA X-RAY DIFFRACTION √ C 6EHT_D 52 - 71 52 71 100 100 3.2Å PCNA X-RAY DIFFRACTION √ D 6EHT_E 52 - 71 52 71 100 100 3.2Å PCNA X-RAY DIFFRACTION √ E 6GWS_D 41-72 41 72 100 100 3.2Å PCNA X-RAY DIFFRACTION √ F 6GWS_E 41-72 41 72 100 100 2.9Å PCNA X-RAY DIFFRACTION √ G 6GWS_F 41-72 41 72 100 100 2.9Å PCNA X-RAY DIFFRACTION √ H 6IIW_B 2-11 2 11 100 100 1.699Å UHRF1 X-RAY DIFFRACTION √ www.aging-us.com 1 AGING Supplementary Table 2. Significantly enriched gene ontology (GO) annotations (cellular components) of KIAA0101 in lung adenocarcinoma (LinkedOmics). Leading Description FDR Leading Edge Gene EdgeNum RAD51, SPC25, CCNB1, BIRC5, NCAPG, ZWINT, MAD2L1, SKA3, NUF2, BUB1B, CENPA, SKA1, AURKB, NEK2, CENPW, HJURP, NDC80, CDCA5, NCAPH, BUB1, ZWILCH, CENPK, KIF2C, AURKA, CENPN, TOP2A, CENPM, PLK1, ERCC6L, CDT1, CHEK1, SPAG5, CENPH, condensed 66 0 SPC24, NUP37, BLM, CENPE, BUB3, CDK2, FANCD2, CENPO, CENPF, BRCA1, DSN1, chromosome MKI67, NCAPG2, H2AFX, HMGB2, SUV39H1, CBX3, TUBG1, KNTC1, PPP1CC, SMC2, BANF1, NCAPD2, SKA2, NUP107, BRCA2, NUP85, ITGB3BP, SYCE2, TOPBP1, DMC1, SMC4, INCENP. RAD51, OIP5, CDK1, SPC25, CCNB1, BIRC5, NCAPG, ZWINT, MAD2L1, SKA3, NUF2, BUB1B, CENPA, SKA1, AURKB, NEK2, ESCO2, CENPW, HJURP, TTK, NDC80, CDCA5, BUB1, ZWILCH, CENPK, KIF2C, AURKA, DSCC1, CENPN, CDCA8, CENPM, PLK1, MCM6, ERCC6L, CDT1, HELLS, CHEK1, SPAG5, CENPH, PCNA, SPC24, CENPI, NUP37, FEN1, chromosomal 94 0 CENPL, BLM, KIF18A, CENPE, MCM4, BUB3, SUV39H2, MCM2, CDK2, PIF1, DNA2, region CENPO, CENPF, CHEK2, DSN1, H2AFX, MCM7, SUV39H1, MTBP, CBX3, RECQL4, KNTC1, PPP1CC, CENPP, CENPQ, PTGES3, NCAPD2, DYNLL1, SKA2, HAT1, NUP107, MCM5, MCM3, MSH2, BRCA2, NUP85, SSB, ITGB3BP, DMC1, INCENP, THOC3, XPO1, APEX1, XRCC5, KIF22, DCLRE1A, SEH1L, XRCC3, NSMCE2, RAD21. -

Lorena Novoa-Aponte
A Strain: fl/fl ∆hep Strain: fl/fl ∆hep AnalysisAAV: ofLuc the ironLuc chaperonePCBP1-WT PCBP1PCBP1-∆Fe interactome:PCBP1-∆RNA PCBP1 variant intersection of DNA repair and iron traffickingAAV: Luc Luc WT ∆Fe ∆RNA PCBP1 −40 Lorena Novoa-Aponte a*, Sarju J. Patel a, Olga Protchenko a, James Wohlschlegel b, Caroline C. Philpott a −40 PCBP1, IHC PCBP1, GAPDH - a Genetics and Metabolism Section,a NIDDK, NIH, Bethesda, MD, USA. b Department of Biological Chemistry, UCLA, Los Angeles, CA, USA C.C. Philpott, et al. *[email protected] 80 B P = 0.0262 60 40 1. Iron is essential but toxic 2. Increased DNA damage in cells lacking PCBP1 ns H&E ? + 20 C.C. Philpott, et al. Iron is used as an essential cofactor by several enzymes involved in DNA ⦿ Increased TUNEL in cells and tissue livers from mice lacking PCBP1. ? replication and repair. However, unchaperoned iron promotes redox ⦿ PCBP1 binds both, iron and single-stranded nucleicTG, nmol/mg protein 0 acids. stress that may affect DNA stability. Strain: Δhep Δhep Δhep ? ⦿ The iron binding activity of PCBP1 controls suppressionAAV: of DNA damage C WT ∆Fe ∆RNA PCBP1 variant ? Iron storage HEK293 cells Mouse Liver ? A 8 siNT+P1var 100 ? A 8 var siPCBP1 2 ? Mammals use the iron siNT+P1 ns var Fe-S cluster assembly 6 siPCBP1+P1siPCBP1 80 ADI1 TUNEL chaperone PCBP1 to var ? 6 siPCBP1+P1 metalate several iron- 60 = 0.0468 = 0.0354 = 0.0465 cell / mm population P P P population + dependent enzymes + 4 + 4 ? 40 Degradation of HIF1α Fig. 3. Iron chaperone-mediated handling of cytosolic labile iron pool. -

Identification of Enhancer of Mrna Decapping 4 As a Novel Fusion Partner of MLL in Acute Myeloid Leukemia
STIMULUS REPORT Identification of enhancer of mRNA decapping 4 as a novel fusion partner of MLL in acute myeloid leukemia Heiko Becker,1-3 Gabriele Greve,1,2 Keisuke Kataoka,4 Jan-Philipp Mallm,5,6 Jesus´ Duque-Afonso,1,2,7 Tobias Ma,1,2 Christoph Niemoller,¨ 1,2 Milena Pantic,1 Justus Duyster,1-3 Michael L. Cleary,7 Julia Schuler,¨ 8 Karsten Rippe,5,6 Seishi Ogawa,3 and Michael Lubbert¨ 1-3 1Department of Medicine I, Medical Center, and 2Faculty of Medicine, University of Freiburg, Freiburg, Germany; 3German Cancer Consortium partner site, Freiburg, Germany; 4Department of Pathology and Tumor Biology, Kyoto University, Kyoto, Japan; 5Division of Chromatin Networks and 6Single-cell Open Laboratory, German Cancer Research Center, Heidelberg, Germany; 7Department of Pathology, Stanford University, Stanford, CA; and 8Charles River Discovery Research Services Germany GmbH, Freiburg, Germany Key Points Introduction • mRNA decapping gene Translocations involving MLL (aka KMT2A) located on chromosome 11q23 occur in acute myeloid EDC4 is a novel fusion leukemia (AML) and lymphoblastic leukemia. In AML, they generally confer an adverse prognosis, unless MLL partner of in AML. the MLLT3 (aka AF9) gene is involved.1 More than 130 different translocation partner genes (TPGs) MLL 2 • Genes functioning in have been identified, forming the recombinome. mRNA decapping may Recently, the scavenger messenger RNA (mRNA) decapping enzyme DCPS has been identified to be compose a distinct required for survival of AML cells, but not normal hematopoietic cells, and a DCPS inhibitor showed antileukemic activity.3,4 DCPS is also 1 of 2 genes (the other being DCP1A) involved in mRNA group of MLL fusion decapping and having been described as TPG of MLL in single leukemia cases.5-7 partners that links MLL MLL function with mRNA Here, we describe a novel fusion with another mRNA decapping component, ie, the enhancer of mRNA decapping 4 gene (EDC4;alsoknownasGE1 or HEDLS), in AML. -
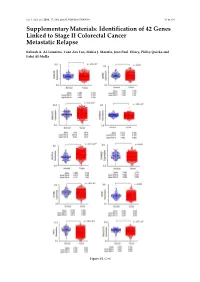
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

A Non-Canonical Role for the EDC4 Decapping Factor in Regulating
RESEARCH ARTICLE A non-canonical role for the EDC4 decapping factor in regulating MARF1- mediated mRNA decay William R Brothers1, Steven Hebert1, Claudia L Kleinman1,2, Marc R Fabian1,3,4* 1Lady Davis Institute for Medical Research, Jewish General Hospital, Montreal, Canada; 2Department of Human Genetics, McGill University, Montreal, Canada; 3Department of Biochemistry, McGill University, Montreal, Canada; 4Department of Oncology, McGill University, Montreal, Canada Abstract EDC4 is a core component of processing (P)-bodies that binds the DCP2 decapping enzyme and stimulates mRNA decay. EDC4 also interacts with mammalian MARF1, a recently identified endoribonuclease that promotes oogenesis and contains a number of RNA binding domains, including two RRMs and multiple LOTUS domains. How EDC4 regulates MARF1 action and the identity of MARF1 target mRNAs is not known. Our transcriptome-wide analysis identifies bona fide MARF1 target mRNAs and indicates that MARF1 predominantly binds their 3’ UTRs via its LOTUS domains to promote their decay. We also show that a MARF1 RRM plays an essential role in enhancing its endonuclease activity. Importantly, we establish that EDC4 impairs MARF1 activity by preventing its LOTUS domains from binding target mRNAs. Thus, EDC4 not only serves as an enhancer of mRNA turnover that binds DCP2, but also as a repressor that binds MARF1 to prevent the decay of MARF1 target mRNAs. Introduction *For correspondence: The regulated decay of eukaryotic mRNA populations plays an important role in the post-transcrip- [email protected] tional control (PTC) of gene expression. These PTC programs are, in turn, critical for regulating a number of biological processes, including during early development, cell proliferation and immune Competing interests: The response. -

Identification of Edc3p As an Enhancer of Mrna Decapping In
Copyright 2004 by the Genetics Society of America Identification of Edc3p as an Enhancer of mRNA Decapping in Saccharomyces cerevisiae Meenakshi Kshirsagar and Roy Parker1 Department of Molecular and Cellular Biology and Howard Hughes Medical Institute, University of Arizona, Tucson, Arizona 85721-0106 Manuscript received July 8, 2003 Accepted for publication October 27, 2003 ABSTRACT The major pathway of mRNA decay in yeast initiates with deadenylation, followed by mRNA decapping and 5Ј–3Ј exonuclease digestion. An in silico approach was used to identify new proteins involved in the mRNA decay pathway. One such protein, Edc3p, was identified as a conserved protein of unknown function having extensive two-hybrid interactions with several proteins involved in mRNA decapping and 5Ј–3Ј degradation including Dcp1p, Dcp2p, Dhh1p, Lsm1p, and the 5Ј–3Ј exonuclease, Xrn1p. We show that Edc3p can stimulate mRNA decapping of both unstable and stable mRNAs in yeast when the decapping enzyme is compromised by temperature-sensitive alleles of either the DCP1 or the DCP2 genes. In these cases, deletion of EDC3 caused a synergistic mRNA-decapping defect at the permissive temperatures. The edc3⌬ had no effect when combined with the lsm1⌬, dhh1⌬,orpat1⌬ mutations, which appear to affect an early step in the decapping pathway. This suggests that Edc3p specifically affects the function of the decapping enzyme per se. Consistent with a functional role in decapping, GFP-tagged Edc3p localizes to cytoplasmic foci involved in mRNA decapping referred to as P-bodies. These results identify Edc3p as a new protein involved in the decapping reaction. N eukaryotic cells, mRNA turnover and its regulation al. -

Phosphorylation of Mrna Decapping Protein Dcp1a by the ERK Signaling Pathway During Early Differentiation of 3T3-L1 Preadipocytes
Phosphorylation of mRNA Decapping Protein Dcp1a by the ERK Signaling Pathway during Early Differentiation of 3T3-L1 Preadipocytes Pei-Yu Chiang1., Yu-Fang Shen1., Yu-Lun Su1, Ching-Han Kao1, Nien-Yi Lin2, Pang-Hung Hsu3, Ming- Daw Tsai1,2,4, Shun-Chang Wang2, Geen-Dong Chang1, Sheng-Chung Lee2,5, Ching-Jin Chang1,2* 1 Graduate Institute of Biochemical Sciences, College of Life Science, National Taiwan University, Taipei, Taiwan, 2 Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan, 3 Department of Life Science, Institute of Bioscience and Biotechnology, National Taiwan Ocean University, Keelung, Taiwan, 4 Genomic Research Center, Academia Sinica, Taipei, Taiwan, 5 Institute of Molecular Medicine, College of Medicine, National Taiwan University, Taipei, Taiwan Abstract Background: Turnover of mRNA is a critical step in the regulation of gene expression, and an important step in mRNA decay is removal of the 59 cap. We previously demonstrated that the expression of some immediate early gene mRNAs is controlled by RNA stability during early differentiation of 3T3-L1 preadipocytes. Methodology/Principal Findings: Here we show that the mouse decapping protein Dcp1a is phosphorylated via the ERK signaling pathway during early differentiation of preadipocytes. Mass spectrometry analysis and site-directed mutagenesis combined with a kinase assay identified ERK pathway–mediated dual phosphorylation at Ser 315 and Ser 319 of Dcp1a. To understand the functional effects of Dcp1a phosphorylation, we examined protein-protein interactions between Dcp1a and other decapping components with co-immunoprecipitation. Dcp1a interacted with Ddx6 and Edc3 through its proline-rich C-terminal extension, whereas the conserved EVH1 (enabled vasodilator-stimulated protein homology 1) domain in the N terminus of Dcp1a showed a stronger interaction with Dcp2. -

The Natural Antisense Transcript NATTD Regulates the Transcription
International Journal of Biochemistry and Cell Biology 110 (2019) 103–110 Contents lists available at ScienceDirect International Journal of Biochemistry and Cell Biology journal homepage: www.elsevier.com/locate/biocel The natural antisense transcript NATTD regulates the transcription of decapping scavenger (DcpS) enzyme T ⁎ Zhu-Zhong Mei , Hongwei Sun, Xiaoli Ou, Lei Li, Junwei Cai, Shuiwang Hu, Juan Wang, ⁎ ⁎ Haihua Luo, Jinghua Liu , Yong Jiang From Guangdong Provincial Key Laboratory of Proteomics, State Key Laboratory of Organ Failure Research, Department of Pathophysiology, Southern Medical University, Guangzhou 510515, China ARTICLE INFO ABSTRACT Keywords: Natural antisense transcripts (NATs) are transcribed from the opposite strand of other genes. Most of them are Natural antisense transcript noncoding RNAs. They have been reported to play important roles in a variety of biological processes. In this NATTD study, we identified a novel NAT, NATTD, which is partially complementary to both the TIRAP/Mal and DcpS DcpS genes. Interestingly, NATTD only positively regulates the expression of DcpS, a decapping scavenger enzyme Epigenetic regulation which is a promising therapeutic target for spinal muscular atrophy. But it has no obvious effects on the ex- pression of TIRAP/Mal gene. The NATTD transcript primarily resides in the nucleus and does not alter the mRNA stability of DcpS. Instead, it is required for the recruitment of RNA polymerase II at the mouse DcpS promoter. Chromatin immunoprecipitation assays revealed that knocking-down -

Role of Dcps in Mammalian RNA Regulation and Human Diseases
ROLE OF DCPS IN MAMMALIAN RNA REGULATION AND HUMAN DISEASES By MI ZHOU A dissertation submitted to the Graduate School-New Brunswick and The Graduate School of Biomedical Sciences Rutgers, The State University of New Jersey In partial fulfillment of the requirements For the degree of Doctor of Philosophy Graduate Program in Cell and Development Biology Written under the direction of Dr. Megerditch Kiledjian And approved by _________________________________ _________________________________ _________________________________ _________________________________ New Brunswick, New Jersey October, 2015 ABSTRACT OF THE DISSERTATION Role of DcpS in Mammalian RNA Regulation and Human Diseases By MI ZHOU Dissertation Director Dr. Megerditch Kiledjian In eukaryotic cells, mRNA degradation plays an important role in the control of gene expression and is therefore highly regulated. The scavenger decapping enzyme DcpS is a multifunctional protein that plays a critical role in mRNA degradation. We first sought to identify DcpS target genes in mammalian cells using a cell permeable DcpS inhibitor compound, RG3039, which was initially developed for therapeutic treatment of Spinal Muscular Atrophy (SMA). Microarray analysis following DcpS decapping inhibition by RG3039 revealed the steady state levels of 222 RNAs were altered. Of a subset selected for validation by qRT-PCR, two non-coding transcripts dependent on DcpS decapping activity, were identified and referred to as DcpS Responsive Noncoding Transcript (DRNT) 1 and 2 respectively. Only the increase in DRNT1 transcript was accompanied with an increase of its RNA stability and this increase was dependent on both DcpS and Xrn1. Our data indicate that DcpS is a transcript-restricted modulator of RNA stability in mammalian cells and the RG3039 ii quinazoline compound is pleotropic, influence gene expression in both an apparent DcpS dependent and independent manner. -

Transcription Start Site Analysis Reveals Widespread Divergent Transcription in D. Melanogaster and Core Promoter-Encoded Enhanc
bioRxiv preprint doi: https://doi.org/10.1101/221952; this version posted November 18, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Transcription start site analysis reveals widespread divergent transcription in D. melanogaster and core promoter-encoded enhancer activities Sarah Rennie1,+, Maria Dalby1,+, Marta Lloret-Llinares2, Stylianos Bakoulis1, Christian Dalager Vaagensø1, Torben Heick Jensen2, and Robin Andersson1,* 1The Bioinformatics Centre, Department of Biology, University of Copenhagen, Copenhagen, Denmark 2Department of Molecular Biology and Genetics, Aarhus University, Aarhus, Denmark +these authors contributed equally to this work *correspondence should be addressed to RA: [email protected] ABSTRACT Mammalian gene promoters and enhancers share many properties. They are composed of a unified promoter architecture of divergent transcripton initiation and gene promoters may exhibit enhancer function. However, it is currently unclear how expression strength of a regulatory element relates to its enhancer strength and if the unifying architecture is conserved across Metazoa. Here we investigate the transcription initiation landscape and its associated RNA decay in D. melanogaster. Surprisingly, we find that the vast majority of active gene-distal enhancers and a large fraction of gene promoters are divergently transcribed from two divergent core promoters. Moreover, we observe quantitative relationships between enhancer potential, expression level and core promoter strength, providing an explanation for indirectly related histone modifications that are reflecting expression levels. Lowly abundant unstable RNAs initiated from weak core promoters are key characteristics of gene-distal developmental enhancers, while the housekeeping enhancer strengths of gene promoters are directly echoing their expression strengths.