Processivity of DNA Polymerases: Two Mechanisms, One Goal Zvi Kelman1*, Jerard Hurwitz1 and Mike O’Donnell2
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Klenow Fragment, #EP0054
Description Klenow Fragment is the Large Fragment of DNA Polymerase I, E.coli . It exhibits 5' →3' polymerase activity and 3' →5' exonuclease (proofreading) activity, but lacks 5' →3' exonuclease activity of DNA Polymerase I. PRODUCT INFORMATION Applications Klenow Fragment • DNA blunting by fill-in of 5’-overhangs or removal of 3‘-overhangs. (1), see protocols on back page. • Random-primed DNA labeling (2-4). #EP0054 300 U • Labeling by fill-in 5 ’-overhangs of dsDNA. Lot: _ Expiry Date: _ • DNA sequencing by the Sanger method (5). • Site-specific mutagenesis of DNA with synthetic oligonucleotides (6). Concentration: 2 U/µL • Second strand synthesis of cDNA (7). Source Supplied with: 1 mL of 10X Reaction Buffer E.coli cells with a cloned fragment of the polA gene. Molecular Weight 68 kDa monomer. Store at -20 °C Definition of Activity Unit One unit of the enzyme catalyzes the incorporation of 10 nmol of deoxyribonucleotides into a polynucleotide fraction (adsorbed on DE-81) in 30 min at 37°C. Enzyme activity is assayed in the following mixture: 50 mM Tris-HCl (pH 8.0 at 25°C), 5 mM MgCl 2, 1 mM DTT, In total 2 vials. 0.033 mM dNTP, 0.4 M Bq/mL [3H]-dTTP and 62.5 µg/mL activated salmon milt DNA. www.thermoscientific.com/onebio Rev.9 V Storage Buffer CERTIFICATE OF ANALYSIS The enzyme is supplied in: 25 mM Tris-HCl (pH 7.5), Endodeoxyribonuclease Assay 0.1 mM EDTA, 1 mM DTT and 50% (v/v) glycerol. 10X Reaction Buffer No conversion of covalently closed circular DNA to nicked DNA was detected after incubation of 20 units of Klenow 500 mM Tris-HCl (pH 8.0 at 25°C), 50 mM MgCl 2, 10 mM DTT. -

A Mutation in DNA Polymerase Α Rescues WEE1KO Sensitivity to HU
International Journal of Molecular Sciences Article A Mutation in DNA Polymerase α Rescues WEE1KO Sensitivity to HU Thomas Eekhout 1,2 , José Antonio Pedroza-Garcia 1,2 , Pooneh Kalhorzadeh 1,2, Geert De Jaeger 1,2 and Lieven De Veylder 1,2,* 1 Department of Plant Biotechnology and Bioinformatics, Ghent University, 9052 Gent, Belgium; [email protected] (T.E.); [email protected] (J.A.P.-G.); [email protected] (P.K.); [email protected] (G.D.J.) 2 Center for Plant Systems Biology, VIB, 9052 Gent, Belgium * Correspondence: [email protected] Abstract: During DNA replication, the WEE1 kinase is responsible for safeguarding genomic integrity by phosphorylating and thus inhibiting cyclin-dependent kinases (CDKs), which are the driving force of the cell cycle. Consequentially, wee1 mutant plants fail to respond properly to problems arising during DNA replication and are hypersensitive to replication stress. Here, we report the identification of the pola-2 mutant, mutated in the catalytic subunit of DNA polymerase α, as a suppressor mutant of wee1. The mutated protein appears to be less stable, causing a loss of interaction with its subunits and resulting in a prolonged S-phase. Keywords: replication stress; DNA damage; cell cycle checkpoint Citation: Eekhout, T.; Pedroza- 1. Introduction Garcia, J.A.; Kalhorzadeh, P.; De Jaeger, G.; De Veylder, L. A Mutation DNA replication is a highly complex process that ensures the chromosomes are in DNA Polymerase α Rescues correctly replicated to be passed onto the daughter cells during mitosis. Replication starts WEE1KO Sensitivity to HU. Int. -

Klenow Fragment Is a Mesophilic DNA Polymerase Derived from the E.Coli Polymerase I DNA- Klenow Fragment Dependent Repair Enzyme
Product Specifications P7060L Rev C Product Information Product Description: Klenow Fragment is a mesophilic DNA polymerase derived from the E.coli Polymerase I DNA- Klenow Fragment dependent repair enzyme. The enzyme exhibits DNA synthesis and proofreading (3′→5′) nuclease activities, and, Part Number P7060L in the absence of the holoenzyme’s (5′→3′) nuclease domain, displays a moderate strand displacement activity Concentration 5,000 U/mL during DNA synthesis. The protein is expressed as a Unit Size 2,500 U truncated product of the E.coli PolA gene. Storage Temperature -25⁰C to -15⁰C Product Specifications P7060 Specific SS DS E. coli DNA Assay SDS Purity DS Exonuclease Activity Exonuclease Endonuclease Contamination Units Tested n/a n/a 50 50 50 50 Specification >99% 5,000 U/mg Functional Functional No Conversion <10 copies Source of Protein: A recombinant E. coli strain carrying the Klenow Fragment gene. Unit Definition: 1 unit is defined as the amount of polymerase required to convert 10 nmol of dNTPs into acid insoluble material in 30 minutes at 37°C. Molecular weight: 68,202 Daltons Quality Control Analysis: Unit Activity is measured using a 2-fold serial dilution method. Dilutions of enzyme were made in a 50% glycerol Klenow (3’-5’ exo-) storage solution and added to 50 µL reactions containing Calf Thymus DNA, 1X Klenow Reaction Buffer, 3H-dTTP and 100 µM dNTPs. Reactions were incubated 10 minutes at 37°C, plunged on ice, and analyzed using the method of Sambrook and Russell (Molecular Cloning, v3, 2001, pp. A8.25-A8.26). Protein Concentration (OD280) is determined by OD280 absorbance. -

DNA Polymerase V Activity Is Autoregulated by a Novel Intrinsic DNA-Dependent
1 2 DNA polymerase V activity is autoregulated by a novel intrinsic DNA-dependent 3 ATPase 4 Aysen L. Erdem1, Malgorzata Jaszczur1, Jeffrey G. Bertram1, Roger Woodgate2, Michael M. Cox3 & 5 Myron F. Goodman1 6 1Departments of Biological Sciences and Chemistry, University of Southern California, University 7 Park, Los Angeles, California 90089-2910, USA. 2Laboratory of Genomic Integrity, National 8 Institute of Child Health and Human Development, National Institutes of Health, Bethesda, 9 Maryland 20892-3371, USA. 3Department of Biochemistry, University of Wisconsin-Madison, 10 Madison, Wisconsin 53706, USA. 11 12 Escherichia coli DNA polymerase V (pol V), a heterotrimeric complex composed of UmuD′2C, 13 is marginally active. ATP and RecA play essential roles in the activation of pol V for DNA 14 synthesis including translesion synthesis (TLS). We have established three features of the roles 15 of ATP and RecA. 1) RecA-activated DNA polymerase V (pol V Mut), is a DNA-dependent 16 ATPase; 2) bound ATP is required for DNA synthesis; 3) pol V Mut function is regulated by 17 ATP, with ATP required to bind primer/template (p/t) DNA and ATP hydrolysis triggering 18 dissociation from the DNA. Pol V Mut formed with an ATPase-deficient RecA E38K/K72R 19 mutant hydrolyzes ATP rapidly, establishing the DNA-dependent ATPase as an intrinsic 20 property of pol V Mut distinct from the ATP hydrolytic activity of RecA when bound to 21 single-stranded (ss)DNA as a nucleoprotein filament (RecA*). No similar ATPase activity or 22 autoregulatory mechanism has previously been found for a DNA polymerase. -

Human Glucokinase Gene
Proc. Nati. Acad. Sci. USA Vol. 89, pp. 7698-7702, August 1992 Genetics Human glucokinase gene: Isolation, characterization, and identification of two missense mutations linked to early-onset non-insulin-dependent (type 2) diabetes mellitus (glucose/metabolism/phosphorylation/structure4unctlon/chromosome 7) M. STOFFEL*, PH. FROGUELt, J. TAKEDA*, H. ZOUALItt, N. VIONNET*, S. NISHI*§, I. T. WEBER¶, R. W. HARRISON¶, S. J. PILKISII, S. LESAGEtt, M. VAXILLAIREtt, G. VELHOtt, F. SUNtt, F. lIRSt, PH. PASSAt, D. COHENt, AND G. I. BELL*"** *Howard Hughes Medical Institute, and Departments of Biochemistry and Molecular Biology, and of Medicine, The University of Chicago, 5841 South Maryland Avenue, MC1028, Chicago, IL 60637; §Second Division of Internal Medicine, Hamamatsu University School of Medicine, Hamamatsu, Shizuoka 431-32, Japan; IDepartment of Pharmacology, Jefferson Cancer Institute, Thomas Jefferson University, Philadelphia, PA 19107; IlDepartment of Physiology and Biophysics, State University of New York, Stony Brook, NY 11794; tCentre d'Etude du Polymorphisme Humain, 27 rue Juliette Dodu, and Service d'Endocrinologie, H6pital Saint-Louis, 75010 Paris, France; and tG6ndthon, 1 rue de l'Internationale, 91000 Evry, France Communicated by Jean Dausset, May 28, 1992 ABSTRACT DNA polymorphisms in the glucokinase gene by maintaining a gradient for glucose transport into these cells have recently been shown to be tightly linked to early-onset thereby regulating hepatic glucose disposal. In (3 cells, glu- non-insulin-dependent diabetes mellitus in "80% of French cokinase is believed to be part of the glucose-sensing mech- families with this form of diabetes. We previously identified a anism and to be involved in the regulation ofinsulin secretion. -

DNA Bound by the Oxytricha Telomere Protein Is Accessible to Telomerase and Other DNA Polymerases DOROTHY E
Proc. Natl. Acad. Sci. USA Vol. 91, pp. 405-409, January 1994 Biochemistry DNA bound by the Oxytricha telomere protein is accessible to telomerase and other DNA polymerases DOROTHY E. SHIPPEN*, ELIZABETH H. BLACKBURNt, AND CAROLYN M. PRICE0§ tDepartment of Microbiology and Immunology, University of California, San Francisco, CA 94143; and tDepartment of Chemistry, University of Nebraska, Lincoln, NB 68588 Contributed by Elizabeth H. Blackburn, August 25, 1993 ABSTRACT Macronuclear telomeres in Oxytricha exist as oftelomere protein in these two populations is not altered by DNA-protein complexes in which the termini of the G-rich additional nuclease treatment. strands are bound by a 97-kDa telomere protein. During The fragment of DNA bound by the majority of telomere telome'ic DNA replication, the replication machinery must protein molecules corresponds to the most terminal 13 or 14 have access to the G-rich strand. However, given the stability nucleotides of the T4G4T4G4 overhang (4). Dimethyl sulfate of telomere protein binding, it has been unclear how this is footprinting demonstrated that the complex formed between accomplished. In this study we investigated the ability of the telomere protein and the residual DNA fragment retains several different DNA polymerases to access telomeric DNA in the same DNA-protein contacts present at native telomeres Oxytricha telomere protein-DNA complexes. Although DNA (4). Thus, these telomeric DNA-protein complexes are useful bound by the telomere protein is not degraded by micrococcal substrates for in vitro investigations of telomere structure nuclease or labeled by terminal deoxynucleotidyltrnsferase, (10). In this study we have employed the DNA-protein this DNA serves as an efficient primer for the addition of complexes to analyze the interaction of protein-bound telo- telomeric repeats by telomerase, a specialized RNA-dependent meric DNA with components of the DNA replication ma- DNA polymerase (ribonucleoprotein reverse tanscriptase), chinery. -

Arthur Kornberg Discovered (The First) DNA Polymerase Four
Arthur Kornberg discovered (the first) DNA polymerase Using an “in vitro” system for DNA polymerase activity: 1. Grow E. coli 2. Break open cells 3. Prepare soluble extract 4. Fractionate extract to resolve different proteins from each other; repeat; repeat 5. Search for DNA polymerase activity using an biochemical assay: incorporate radioactive building blocks into DNA chains Four requirements of DNA-templated (DNA-dependent) DNA polymerases • single-stranded template • deoxyribonucleotides with 5’ triphosphate (dNTPs) • magnesium ions • annealed primer with 3’ OH Synthesis ONLY occurs in the 5’-3’ direction Fig 4-1 E. coli DNA polymerase I 5’-3’ polymerase activity Primer has a 3’-OH Incoming dNTP has a 5’ triphosphate Pyrophosphate (PP) is lost when dNMP adds to the chain E. coli DNA polymerase I: 3 separable enzyme activities in 3 protein domains 5’-3’ polymerase + 3’-5’ exonuclease = Klenow fragment N C 5’-3’ exonuclease Fig 4-3 E. coli DNA polymerase I 3’-5’ exonuclease Opposite polarity compared to polymerase: polymerase activity must stop to allow 3’-5’ exonuclease activity No dNTP can be re-made in reversed 3’-5’ direction: dNMP released by hydrolysis of phosphodiester backboneFig 4-4 Proof-reading (editing) of misincorporated 3’ dNMP by the 3’-5’ exonuclease Fidelity is accuracy of template-cognate dNTP selection. It depends on the polymerase active site structure and the balance of competing polymerase and exonuclease activities. A mismatch disfavors extension and favors the exonuclease.Fig 4-5 Superimposed structure of the Klenow fragment of DNA pol I with two different DNAs “Fingers” “Thumb” “Palm” red/orange helix: 3’ in red is elongating blue/cyan helix: 3’ in blue is getting edited Fig 4-6 E. -

Teacher Notes Science Nspired
DNA Replication TEACHER NOTES SCIENCE NSPIRED Science Objectives In this lesson, students will • Explore how the structure of DNA supports its semi-conservative replication • Identify the name and function of several key enzymes in DNA replication • Recognize the function of Okazaki fragments in DNA replication Vocabulary TI-Nspire™ Technology Skills: • Double helix • Leading strand • Download a TI-Nspire • Lagging strand • Polymerase I • Polymerase III • Helicase document • Base pairs • Okazaki Fragments • Open a document • Primase • Move between pages • Open Directions Box About the Lesson • Explore ‘Hot Spots’ • Using three simulations, students will interact with DNA replication to explore semi-conservative replication and identify Tech Tips: specific enzymes and their roles in replication. Assessments are Make sure that students know embedded in the activity to engage discussion and gauge how to move between pages by learning. pressing /¡ (left arrow) and • As a result, students will: /¢ (right arrow). • Learn the basic functions of the following DNA replication enzymes: helicase, primase, ligase, polymerase I and III. Lesson Materials: • Learn how the double helix reproduces in a semi- Student Activity • DNA_Replication_Student.do conservative fashion c • Learn the role and function of Okazaki fragments in • DNA_Replication_Student.pd replication of the lagging strand. f TI-Nspire document TI-Nspire™ Navigator™ • DNA_Replication.tns Not required. If using Navigator: • Send out the DNA_Replication.tns file. • Monitor student progress -
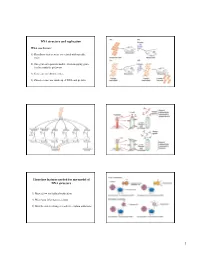
DNA Structure and Replication Three Key Features Needed for Any Model
DNA structure and replication What was known? 1) Hereditary factors were associated with specific traits 2) One-gene-one-protein model - from mapping genes for biosynthetic pathways 3) Genes are on chromosomes 4) Chromosomes are made up of DNA and protein Three key features needed for any model of DNA structure 1) Must allow for faithful replication 2) Must have information content 3) Must be able to change in order to explain mutations 1 Rosalind Franklin James Watson Francis Crick X-ray diffraction Maurice Wilkins Photograph of B-DNA Linus Pauling Nature 171, 737-738 (1953) © Macmillan Publishers Ltd. Molecular structure of Nucleic Acids WATSON, J. D. & CRICK, F. H. C. Features of the double helix A Structure for Deoxyribose Nucleic Acid 1) Two parallel strands We wish to suggest a structure for the salt of deoxyribose nucleic acid (D.N.A.). This structure has novel features which are of considerable biological interest………… 2) Bases held together by H-bonds 3) Phosphodiester backbone 4) The base is attached to the position 1 on the sugar 5) Base pair stack - provides Figure 1. This figure is purely diagrammatic. The two ribbons symbolize the two phophate-sugar chains, and the horizonal rods stability the pairs of bases holding the chains together. The vertical line marks the fibre axis. 6) Contains a major and …………….It has not escaped our notice that the specific pairing we minor groove have postulated immediately suggests a possible copying mechanism for the genetic material. Three key features needed for any model of DNA structure -

Enhancing Terminal Deoxynucleotidyl Transferase Activity on Substrates
G C A T T A C G G C A T genes Communication Enhancing Terminal Deoxynucleotidyl Transferase 0 Activity on Substrates with 3 Terminal Structures for Enzymatic De Novo DNA Synthesis 1,2,3, 1,2,3, 1,2,4 Sebastian Barthel y , Sebastian Palluk y, Nathan J. Hillson , 1,2,5,6,7,8,9 1,2,5,10, , Jay D. Keasling and Daniel H. Arlow * y 1 Joint BioEnergy Institute, Emeryville, CA 94608, USA; [email protected] (S.B.); [email protected] (S.P.); [email protected] (N.J.H.); [email protected] (J.D.K.) 2 Biological Systems and Engineering Division, Lawrence Berkeley National Lab, Berkeley, CA 94720, USA 3 Department of Biology, Technische Universität Darmstadt, 64287 Darmstadt, Germany 4 DOE Joint Genome Institute, Walnut Creek, CA 94598, USA 5 Institute for Quantitative Biosciences, UC Berkeley, Berkeley, CA 94720, USA 6 Department of Chemical and Biomolecular Engineering, UC Berkeley, Berkeley, CA 94720, USA 7 Department of Bioengineering UC Berkeley, Berkeley, CA 94720, USA 8 Novo Nordisk Foundation Center for Biosustainability, Technical University of Denmark, 2970 Hørsholm, Denmark 9 Center for Synthetic Biochemistry, Institute for Synthetic Biology, Shenzhen Institutes for Advanced Technologies, Shenzhen 518055, China 10 Biophysics Graduate Group, UC Berkeley, Berkeley, CA 94720, USA * Correspondence: [email protected]; Tel.: +1-248-227-5556 Current address: Ansa Biotechnologies, Berkeley, CA 94170, USA. y Received: 8 December 2019; Accepted: 7 January 2020; Published: 16 January 2020 Abstract: Enzymatic oligonucleotide synthesis methods based on the template-independent polymerase terminal deoxynucleotidyl transferase (TdT) promise to enable the de novo synthesis of long oligonucleotides under mild, aqueous conditions. -

T5 DNA Polymerase: Structural-Functional Relationships to Other DNA Polymerases (DNA Polymerase I/Proofreading/Processivity/Evolution) MARK C
Proc. Nati. Acad. Sci. USA Vol. 86, pp. 4465-4469, June 1989 Biochemistry T5 DNA polymerase: Structural-functional relationships to other DNA polymerases (DNA polymerase I/proofreading/processivity/evolution) MARK C. LEAVITT AND JUNETSU ITO Department of Microbiology and Immunology, University of Arizona Health Sciences Center, Tucson, AZ 85724 Communicated by Lester 0. Krampitz, April 10, 1989 (receivedfor review February 1, 1989) ABSTRACT T5 DNA polymerase, a highly processive sin- proceed through double-stranded regions in template sec- gle-polypeptide enzyme, has been analyzed for its primary ondary structures or supercoiled plasmid templates. structural features. The amino acid sequence of T5 DNA We present here the DNA sequence of the T5 DNA polymerase has a high degree of homology with that of DNA polymerase gene* and the deduced amino acid sequence ofits polymerase I from Escherichia coli and retains many of the product. Comparisons of the primary structure of this en- amino acid residues that have been implicated in the 3' -* 5' zyme with other DNA polymerases suggest differences that exonuclease and DNA polymerase activities of that enzyme. may account for the high processivity of this enzyme. We Alignment with sequences of polymerase I and T7 DNA poly- also demonstrate the conservation of residues thought to be merase was used to identify regions possibly involved in the intimately involved in 3' -* 5' exonuclease and polymerase high processivity of this enzyme. Further, amino acid sequence activities. Finally, two amino acid sequence segments, which comparisons ofT5 DNA polymerase with a large group ofDNA may be involved in the 3' -*5' exonuclease function of these polymerases previously shown to exhibit little similarity to enzymes, appear to be highly conserved among a wide polymerase I indicate certain sequence segments are shared variety of DNA polymerases. -

Primer Release Is the Rate-Limiting Event in Lagging-Strand Synthesis
Primer release is the rate-limiting event in lagging- strand synthesis mediated by the T7 replisome Alfredo J. Hernandeza, Seung-Joo Leea, and Charles C. Richardsona,1 aDepartment of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115 Contributed by Charles C. Richardson, April 18, 2016 (sent for review December 30, 2015; reviewed by Nicholas E. Dixon and I. Robert Lehman) DNA replication occurs semidiscontinuously due to the antiparallel but also for enabling the use of short oligoribonucleotides by T7 DNA strands and polarity of enzymatic DNA synthesis. Although DNA polymerase. Critically, the primase domain also fulfills two the leading strand is synthesized continuously, the lagging strand additional roles apart from primer synthesis: it prevents disso- is synthesized in small segments designated Okazaki fragments. ciation of the extremely short tetramer, stabilizing it with the Lagging-strand synthesis is a complex event requiring repeated template, and it secures it in the polymerase active site (10, 12). cycles of RNA primer synthesis, transfer to the lagging-strand Here we show that the rate-limiting step in initiation of Okazaki polymerase, and extension effected by cooperation between DNA fragments by the T7 replisome is primer release from the primase primase and the lagging-strand polymerase. We examined events domain of gp4. In the absence of gp2.5, an additional step, distinct controlling Okazaki fragment initiation using the bacteriophage from primer release, also limits primer extension. The presence of T7 replication system. Primer utilization by T7 DNA polymerase is gp2.5 promotes efficient primer formation and primer utilization. slower than primer formation.