The Effect of the Human Peptide GHK on Gene Expression Relevant to Nervous System Function and Cognitive Decline
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 Evidence for Gliadin Antibodies As Causative Agents in Schizophrenia
1 Evidence for gliadin antibodies as causative agents in schizophrenia. C.J.Carter PolygenicPathways, 20 Upper Maze Hill, Saint-Leonard’s on Sea, East Sussex, TN37 0LG [email protected] Tel: 0044 (0)1424 422201 I have no fax Abstract Antibodies to gliadin, a component of gluten, have frequently been reported in schizophrenia patients, and in some cases remission has been noted following the instigation of a gluten free diet. Gliadin is a highly immunogenic protein, and B cell epitopes along its entire immunogenic length are homologous to the products of numerous proteins relevant to schizophrenia (p = 0.012 to 3e-25). These include members of the DISC1 interactome, of glutamate, dopamine and neuregulin signalling networks, and of pathways involved in plasticity, dendritic growth or myelination. Antibodies to gliadin are likely to cross react with these key proteins, as has already been observed with synapsin 1 and calreticulin. Gliadin may thus be a causative agent in schizophrenia, under certain genetic and immunological conditions, producing its effects via antibody mediated knockdown of multiple proteins relevant to the disease process. Because of such homology, an autoimmune response may be sustained by the human antigens that resemble gliadin itself, a scenario supported by many reports of immune activation both in the brain and in lymphocytes in schizophrenia. Gluten free diets and removal of such antibodies may be of therapeutic benefit in certain cases of schizophrenia. 2 Introduction A number of studies from China, Norway, and the USA have reported the presence of gliadin antibodies in schizophrenia 1-5. Gliadin is a component of gluten, intolerance to which is implicated in coeliac disease 6. -

Downloaded from the National Database for Autism Research (NDAR)
International Journal of Molecular Sciences Article Phenotypic Subtyping and Re-Analysis of Existing Methylation Data from Autistic Probands in Simplex Families Reveal ASD Subtype-Associated Differentially Methylated Genes and Biological Functions Elizabeth C. Lee y and Valerie W. Hu * Department of Biochemistry and Molecular Medicine, The George Washington University, School of Medicine and Health Sciences, Washington, DC 20037, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-202-994-8431 Current address: W. Harry Feinstone Department of Molecular Microbiology and Immunology, y Johns Hopkins Bloomberg School of Public Health, Baltimore, MD 21205, USA. Received: 25 August 2020; Accepted: 17 September 2020; Published: 19 September 2020 Abstract: Autism spectrum disorder (ASD) describes a group of neurodevelopmental disorders with core deficits in social communication and manifestation of restricted, repetitive, and stereotyped behaviors. Despite the core symptomatology, ASD is extremely heterogeneous with respect to the severity of symptoms and behaviors. This heterogeneity presents an inherent challenge to all large-scale genome-wide omics analyses. In the present study, we address this heterogeneity by stratifying ASD probands from simplex families according to the severity of behavioral scores on the Autism Diagnostic Interview-Revised diagnostic instrument, followed by re-analysis of existing DNA methylation data from individuals in three ASD subphenotypes in comparison to that of their respective unaffected siblings. We demonstrate that subphenotyping of cases enables the identification of over 1.6 times the number of statistically significant differentially methylated regions (DMR) and DMR-associated genes (DAGs) between cases and controls, compared to that identified when all cases are combined. Our analyses also reveal ASD-related neurological functions and comorbidities that are enriched among DAGs in each phenotypic subgroup but not in the combined case group. -

Id4, a New Candidate Gene for Senile Osteoporosis, Acts As a Molecular Switch Promoting Osteoblast Differentiation
Id4, a New Candidate Gene for Senile Osteoporosis, Acts as a Molecular Switch Promoting Osteoblast Differentiation Yoshimi Tokuzawa1., Ken Yagi1., Yzumi Yamashita1, Yutaka Nakachi1, Itoshi Nikaido1, Hidemasa Bono1, Yuichi Ninomiya1, Yukiko Kanesaki-Yatsuka1, Masumi Akita2, Hiromi Motegi3, Shigeharu Wakana3, Tetsuo Noda3,4, Fred Sablitzky5, Shigeki Arai6, Riki Kurokawa6, Toru Fukuda7, Takenobu Katagiri7, Christian Scho¨ nbach8,9, Tatsuo Suda1, Yosuke Mizuno1, Yasushi Okazaki1* 1 Division of Functional Genomics and Systems Medicine, Research Center for Genomic Medicine, Saitama Medical University, Hidaka, Saitama, Japan, 2 Division of Morphological Science, Biomedical Research Center, Saitama Medical University, Iruma-gun, Saitama, Japan, 3 RIKEN BioResource Center, Tsukuba, Ibaraki, Japan, 4 The Cancer Institute of the Japanese Foundation for Cancer Research, Koto-ward, Tokyo, Japan, 5 Developmental Genetics and Gene Control, Institute of Genetics, University of Nottingham, Queen’s Medical Center, Nottingham, United Kingdom, 6 Division of Gene Structure and Function, Research Center for Genomic Medicine, Saitama Medical University, Hidaka, Saitama, Japan, 7 Division of Pathophysiology, Research Center for Genomic Medicine, Saitama Medical University, Hidaka, Saitama, Japan, 8 Division of Genomics and Genetics, Nanyang Technological University School of Biological Sciences, Singapore, Singapore, 9 Department of Bioscience and Bioinformatics, Kyushu Institute of Technology, Iizuka, Fukuoka, Japan Abstract Excessive accumulation of bone marrow adipocytes observed in senile osteoporosis or age-related osteopenia is caused by the unbalanced differentiation of MSCs into bone marrow adipocytes or osteoblasts. Several transcription factors are known to regulate the balance between adipocyte and osteoblast differentiation. However, the molecular mechanisms that regulate the balance between adipocyte and osteoblast differentiation in the bone marrow have yet to be elucidated. -

A Core Transcriptional Network Composed of Pax2/8, Gata3 and Lim1 Regulates Key Players of Pro/Mesonephros Morphogenesis
Developmental Biology 382 (2013) 555–566 Contents lists available at ScienceDirect Developmental Biology journal homepage: www.elsevier.com/locate/developmentalbiology Genomes and Developmental Control A core transcriptional network composed of Pax2/8, Gata3 and Lim1 regulates key players of pro/mesonephros morphogenesis Sami Kamel Boualia a, Yaned Gaitan a, Mathieu Tremblay a, Richa Sharma a, Julie Cardin b, Artur Kania b, Maxime Bouchard a,n a Goodman Cancer Research Centre and Department of Biochemistry, McGill University, 1160 Pine Ave. W., Montreal, Quebec, Canada H3A 1A3 b Institut de Recherches Cliniques de Montréal, Montréal, Québec, Canada H2W 1R7, Department of Anatomy and Cell Biology, Division of Experimental Medicine, McGill University, Montréal, Quebec, Canada, H3A 2B2 and Faculté de médecine, Université de Montréal, Montréal, Quebec, Canada, H3C 3J7. article info abstract Article history: Translating the developmental program encoded in the genome into cellular and morphogenetic Received 23 January 2013 functions requires the deployment of elaborate gene regulatory networks (GRNs). GRNs are especially Received in revised form crucial at the onset of organ development where a few regulatory signals establish the different 27 July 2013 programs required for tissue organization. In the renal system primordium (the pro/mesonephros), Accepted 30 July 2013 important regulators have been identified but their hierarchical and regulatory organization is still Available online 3 August 2013 elusive. Here, we have performed a detailed analysis of the GRN underlying mouse pro/mesonephros Keywords: development. We find that a core regulatory subcircuit composed of Pax2/8, Gata3 and Lim1 turns on a Kidney development deeper layer of transcriptional regulators while activating effector genes responsible for cell signaling Transcription and tissue organization. -

Ali Shokoohmand Thesis (PDF 8MB)
IDENTIFYING THE MOLECULAR MEDIATORS OF VN AND THE IGF:VN COMPLEX-STIMULATED BREAST CANCER CELL SURVIVAL Ali Shokoohmand School of Biomedical Sciences, Faculty of Health Queensland University of Technology, Australia A thesis submitted for the degree of Doctor of Philosophy of the Queensland University of Technology 2015 QUT Verified Signature Acknowledgements Commencing, pursuing and completing this dissertation like any other project, required abundant resources as well as strong motivation, which wouldn’t have been possible without the people who provided me with the much needed encouragement, support, scientific advice and help with experiments. Therefore, I would like to express my gratitude to the people below. I would like to thank my principal supervisor ‘Dr Mr’ Hollier whose advice, guidance and encouragement has been always available for me through my PhD journey. Thanks for encouraging me to work hard at all times and keeping me motivated during my PhD. Your work ethic and your scientific expertise will always inspire me and I truly learned a lot from you. Abhi, I have always been appreciating to have you beside me. Your encouragement and support was a very great thing to me. I have learnt many deals from you. Your encouragement and support always helped me to work harder. Above all, I always enjoyed talking with you about our cultures, people and countries. I am sure we still have many things to talk about! Zee, thanks for the support you have given throughout my PhD. Without your support, this journey could have been harder for me. Derek, I would like to thank you for listening to me sometimes and being here for me. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplementary Table 3 Complete List of RNA-Sequencing Analysis of Gene Expression Changed by ≥ Tenfold Between Xenograft and Cells Cultured in 10%O2
Supplementary Table 3 Complete list of RNA-Sequencing analysis of gene expression changed by ≥ tenfold between xenograft and cells cultured in 10%O2 Expr Log2 Ratio Symbol Entrez Gene Name (culture/xenograft) -7.182 PGM5 phosphoglucomutase 5 -6.883 GPBAR1 G protein-coupled bile acid receptor 1 -6.683 CPVL carboxypeptidase, vitellogenic like -6.398 MTMR9LP myotubularin related protein 9-like, pseudogene -6.131 SCN7A sodium voltage-gated channel alpha subunit 7 -6.115 POPDC2 popeye domain containing 2 -6.014 LGI1 leucine rich glioma inactivated 1 -5.86 SCN1A sodium voltage-gated channel alpha subunit 1 -5.713 C6 complement C6 -5.365 ANGPTL1 angiopoietin like 1 -5.327 TNN tenascin N -5.228 DHRS2 dehydrogenase/reductase 2 leucine rich repeat and fibronectin type III domain -5.115 LRFN2 containing 2 -5.076 FOXO6 forkhead box O6 -5.035 ETNPPL ethanolamine-phosphate phospho-lyase -4.993 MYO15A myosin XVA -4.972 IGF1 insulin like growth factor 1 -4.956 DLG2 discs large MAGUK scaffold protein 2 -4.86 SCML4 sex comb on midleg like 4 (Drosophila) Src homology 2 domain containing transforming -4.816 SHD protein D -4.764 PLP1 proteolipid protein 1 -4.764 TSPAN32 tetraspanin 32 -4.713 N4BP3 NEDD4 binding protein 3 -4.705 MYOC myocilin -4.646 CLEC3B C-type lectin domain family 3 member B -4.646 C7 complement C7 -4.62 TGM2 transglutaminase 2 -4.562 COL9A1 collagen type IX alpha 1 chain -4.55 SOSTDC1 sclerostin domain containing 1 -4.55 OGN osteoglycin -4.505 DAPL1 death associated protein like 1 -4.491 C10orf105 chromosome 10 open reading frame 105 -4.491 -

Primate Specific Retrotransposons, Svas, in the Evolution of Networks That Alter Brain Function
Title: Primate specific retrotransposons, SVAs, in the evolution of networks that alter brain function. Olga Vasieva1*, Sultan Cetiner1, Abigail Savage2, Gerald G. Schumann3, Vivien J Bubb2, John P Quinn2*, 1 Institute of Integrative Biology, University of Liverpool, Liverpool, L69 7ZB, U.K 2 Department of Molecular and Clinical Pharmacology, Institute of Translational Medicine, The University of Liverpool, Liverpool L69 3BX, UK 3 Division of Medical Biotechnology, Paul-Ehrlich-Institut, Langen, D-63225 Germany *. Corresponding author Olga Vasieva: Institute of Integrative Biology, Department of Comparative genomics, University of Liverpool, Liverpool, L69 7ZB, [email protected] ; Tel: (+44) 151 795 4456; FAX:(+44) 151 795 4406 John Quinn: Department of Molecular and Clinical Pharmacology, Institute of Translational Medicine, The University of Liverpool, Liverpool L69 3BX, UK, [email protected]; Tel: (+44) 151 794 5498. Key words: SVA, trans-mobilisation, behaviour, brain, evolution, psychiatric disorders 1 Abstract The hominid-specific non-LTR retrotransposon termed SINE–VNTR–Alu (SVA) is the youngest of the transposable elements in the human genome. The propagation of the most ancient SVA type A took place about 13.5 Myrs ago, and the youngest SVA types appeared in the human genome after the chimpanzee divergence. Functional enrichment analysis of genes associated with SVA insertions demonstrated their strong link to multiple ontological categories attributed to brain function and the disorders. SVA types that expanded their presence in the human genome at different stages of hominoid life history were also associated with progressively evolving behavioural features that indicated a potential impact of SVA propagation on a cognitive ability of a modern human. -

The Prospective Value of Dopamine Receptors on Bio-Behavior of Tumor
Journal of Cancer 2019, Vol. 10 1622 Ivyspring International Publisher Journal of Cancer 2019; 10(7): 1622-1632. doi: 10.7150/jca.27780 Review The Prospective Value of Dopamine Receptors on Bio-Behavior of Tumor Xu Wang1,2, Zhi-Bin Wang1,2, Chao Luo1,2,4, Xiao-Yuan Mao1,2, Xi Li1,2, Ji-Ye Yin1,2, Wei Zhang1,2,3, Hong-Hao Zhou1,2,3, Zhao-Qian Liu1,2,3 1. Department of Clinical Pharmacology, Xiangya Hospital, Central South University, Changsha 410008, P. R. China; 2. Institute of Clinical Pharmacology, Central South University, Hunan Key Laboratory of Pharmacogenetics, Changsha 410078, P. R. China; 3. National Clinical Research Center for Geriatric Disorders, Xiangya Hospital, Central South University, Changsha 410008, P. R. China; 4. School of Life Sciences, Central South University, Changsha, Hunan 410078. Corresponding author: Professor Zhao-Qian Liu: Department of Clinical Pharmacology, Xiangya Hospital, Central South University, Changsha 410008, P. R. China; Institute of Clinical Pharmacology, Central South University; Hunan Key Laboratory of Pharmacogenetics, Changsha 410078, P. R. China. Tel: +86 731 89753845, Fax: +86 731 82354476, E-mail: [email protected]. © Ivyspring International Publisher. This is an open access article distributed under the terms of the Creative Commons Attribution (CC BY-NC) license (https://creativecommons.org/licenses/by-nc/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2018.06.10; Accepted: 2019.02.07; Published: 2019.03.03 Abstract Dopamine receptors are belong to the family of G protein-coupled receptor. There are five types of dopamine receptor (DR), including DRD1, DRD2, DRD3, DRD4, and DRD5, which are divided into two major groups: the D1-like receptors (DRD1 and DRD5), and the D2-like receptors (DRD2, DRD3, and DRD4). -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Binding Mode Exploration of B1 Receptor Antagonists' by the Use of Molecular Dynamics and Docking Simulation—How Different T
International Journal of Molecular Sciences Article Binding Mode Exploration of B1 Receptor Antagonists’ by the Use of Molecular Dynamics and Docking Simulation—How Different Target Engagement Can Determine Different Biological Effects Marica Gemei 1,*, Carmine Talarico 1 , Laura Brandolini 1, Candida Manelfi 1, Lorena Za 2, Silvia Bovolenta 2, Chiara Liberati 2, Luigi Del Vecchio 3, Roberto Russo 4 , Carmen Cerchia 4, Marcello Allegretti 1 and Andrea Rosario Beccari 1 1 Dompé Farmaceutici SpA, via Campo di Pile, 67100 L’Aquila, Italy; [email protected] (C.T.); [email protected] (L.B.); candida.manelfi@dompe.com (C.M.); [email protected] (M.A.); [email protected] (A.R.B.) 2 Axxam, Via Meucci 3, Bresso, 20091 Milano, Italy; [email protected] (L.Z.); [email protected] (S.B.); [email protected] (C.L.) 3 Ceinge Biotecnologie Avanzate, via G. Salvatore 486, 80145 Napoli, Italy; [email protected] 4 Department of Pharmacy, University of Naples “Federico II”, via D. Montesano, 49, 80131 Napoli, Italy; [email protected] (R.R.); [email protected] (C.C.) * Correspondence: [email protected]; Tel.: +34-06-465916 Received: 26 August 2020; Accepted: 12 October 2020; Published: 16 October 2020 Abstract: The kinin B1 receptor plays a critical role in the chronic phase of pain and inflammation. The development of B1 antagonists peaked in recent years but almost all promising molecules failed in clinical trials. Little is known about these molecules’ mechanisms of action and additional information will be necessary to exploit the potential of the B1 receptor. -
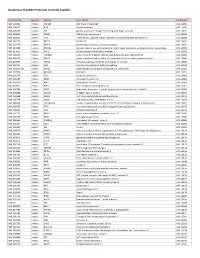
Quantigene Flowrna Probe Sets Currently Available
QuantiGene FlowRNA Probe Sets Currently Available Accession No. Species Symbol Gene Name Catalog No. NM_003452 Human ZNF189 zinc finger protein 189 VA1-10009 NM_000057 Human BLM Bloom syndrome VA1-10010 NM_005269 Human GLI glioma-associated oncogene homolog (zinc finger protein) VA1-10011 NM_002614 Human PDZK1 PDZ domain containing 1 VA1-10015 NM_003225 Human TFF1 Trefoil factor 1 (breast cancer, estrogen-inducible sequence expressed in) VA1-10016 NM_002276 Human KRT19 keratin 19 VA1-10022 NM_002659 Human PLAUR plasminogen activator, urokinase receptor VA1-10025 NM_017669 Human ERCC6L excision repair cross-complementing rodent repair deficiency, complementation group 6-like VA1-10029 NM_017699 Human SIDT1 SID1 transmembrane family, member 1 VA1-10032 NM_000077 Human CDKN2A cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) VA1-10040 NM_003150 Human STAT3 signal transducer and activator of transcripton 3 (acute-phase response factor) VA1-10046 NM_004707 Human ATG12 ATG12 autophagy related 12 homolog (S. cerevisiae) VA1-10047 NM_000737 Human CGB chorionic gonadotropin, beta polypeptide VA1-10048 NM_001017420 Human ESCO2 establishment of cohesion 1 homolog 2 (S. cerevisiae) VA1-10050 NM_197978 Human HEMGN hemogen VA1-10051 NM_001738 Human CA1 Carbonic anhydrase I VA1-10052 NM_000184 Human HBG2 Hemoglobin, gamma G VA1-10053 NM_005330 Human HBE1 Hemoglobin, epsilon 1 VA1-10054 NR_003367 Human PVT1 Pvt1 oncogene homolog (mouse) VA1-10061 NM_000454 Human SOD1 Superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult))