Multi-Scale Inference of Genetic Trait Architecture Using
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
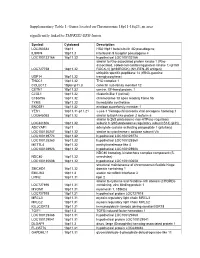
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Glucocorticoid Receptor-Ppara Axis in Fetal Mouse Liver Prepares
RESEARCH ARTICLE Glucocorticoid receptor-PPARa axis in fetal mouse liver prepares neonates for milk lipid catabolism Gianpaolo Rando1†, Chek Kun Tan2†, Nourhe` ne Khaled1, Alexandra Montagner3, Nicolas Leuenberger1, Justine Bertrand-Michel4, Eeswari Paramalingam2, Herve´ Guillou3, Walter Wahli1,2,3* 1Center for Integrative Genomics, University of Lausanne, Lausanne, Switzerland; 2Lee Kong Chian School of Medicine, Nanyang Technological University, Singapore; 3UMR 1331 ToxAlim Research Centre in Food Toxicology, INRA, Universite´ de Toulouse, Toulouse, France; 4IFR 150 Plateforme Metatoul, Institut Fe´de´ratif de Recherche Bio-Me´dicale de Toulouse INSERM U563, Toulouse, France Abstract In mammals, hepatic lipid catabolism is essential for the newborns to efficiently use milk fat as an energy source. However, it is unclear how this critical trait is acquired and regulated. We demonstrate that under the control of PPARa, the genes required for lipid catabolism are transcribed before birth so that the neonatal liver has a prompt capacity to extract energy from milk upon suckling. The mechanism involves a fetal glucocorticoid receptor (GR)-PPARa axis in which GR directly regulates the transcriptional activation of PPARa by binding to its promoter. Certain PPARa target genes such as Fgf21 remain repressed in the fetal liver and become PPARa responsive after birth following an epigenetic switch triggered by b-hydroxybutyrate-mediated inhibition of HDAC3. This study identifies an endocrine developmental axis in which fetal GR primes *For correspondence: walter. the activity of PPARa in anticipation of the sudden shifts in postnatal nutrient source and metabolic [email protected] demands. DOI: 10.7554/eLife.11853.001 †These authors contributed equally to this work Competing interests: The authors declare that no Introduction competing interests exist. -

Homozygosity Mapping of a Dyggve-Melchior-Clausen
714 ORIGINAL ARTICLE J Med Genet: first published as 10.1136/jmg.39.10.714 on 1 October 2002. Downloaded from Homozygosity mapping of a Dyggve-Melchior-Clausen syndrome gene to chromosome 18q21.1 C Thauvin-Robinet, V El Ghouzzi, W Chemaitilly, N Dagoneau, O Boute, G Viot, A Mégarbané, A Sefiani, A Munnich, M Le Merrer, V Cormier-Daire ............................................................................................................................. See end of article for J Med Genet 2002;39:714–717 authors’ affiliations ....................... Correspondence to: Dr V Cormier-Daire, Dyggve-Melchior-Clausen syndrome (DMC) is an autosomal recessive condition characterised by short Département de trunk dwarfism, scoliosis, microcephaly, coarse facies, mental retardation, and characteristic Génétique, Hôpital radiological features. X rays show platyspondyly with double vertebral hump, epiphyseal dysplasia, Necker-Enfants Malades, irregular metaphyses, and a characteristic lacy appearance of the iliac crests. Electron microscopy of 149 rue de Sèvres, 75015 chondrocytes have shown widened cisternae of rough endoplasmic reticulum and biochemical analy- Paris, France; [email protected] ses have shown accumulation of glucosaminoglycan in cartilage, but the pathogenesis of DMC remains unexplained. Here, we report on the homozygosity mapping of a DMC gene to chromosome 18q21.1 Revised version received in seven inbred families (Zmax=9.65 at θ=0 at locus D18S1126) in the genetic interval (1.8 cM) 6 August 2002 Accepted for publication defined by loci D18S455 and D18S363. Despite the various geographical origins of the families 13 August 2002 reported here (Morocco, Tunisia, Portugal, and Lebanon), this condition was genetically homogeneous ....................... in our series. Continuing studies will hopefully lead to the identification of the disease causing gene. -

Discovery Protein Names Uniprot Gene Names Average Log2 L/H
Discovery Ttest IBD vs FDR Average Log2 Average Log2 Ratio Signficant Control p adjusted p Subgroup Protein names Uniprot Gene names L/H ratio Control L/H ratio IBD ibd/control 0.05 FDR value value AUC Specific Immunolgical Aconitate hydratase, mitochondrial A2A274 ACO2 2.570537714 1.404944787 0.311737769 + 1.26481E-09 8.472E-08 0.92417 CD9 antigen A6NNI4 CD9 2.690874857 1.62514501 0.344476348 + 3.02524E-10 2.606E-08 0.95583 Calponin-2 B4DDF4 CNN2 0.388970715 0.995038293 1.833208258 + 0.000445883 0.0044168 0.755 6-phosphogluconate dehydrogenase, decarboxylating B4DQJ8 PGD -0.252090674 0.391282139 1.902888154 + 1.35886E-13 8.373E-11 0.96625 Epithelial cell adhesion molecule B5MCA4 EPCAM 3.405798843 2.468926837 0.39185163 + 0.002975231 0.0215657 0.80625 2,4-dienoyl-CoA reductase, mitochondrial B7Z6B8 DECR1 2.240008 1.557225501 0.50520929 + 0.003411719 0.023728 0.72042 Adenosylhomocysteinase;Putative adenosylhomocysteinase 3 D7UEQ7 AHCYL2 2.14170094 1.336080036 0.446810414 + 0.000222911 0.0024884 0.84583 CD44 antigen E7EPC6 CD44 1.30554758 1.619074953 1.368242916 0.094110279 0.2905348 0.62625 Lactotransferrin;Lactoferricin-H;Kaliocin-1;Lactoferroxin-A;Lactoferroxin- B;Lactoferroxin-C E7ER44 LTF 1.276691743 1.810737039 1.705818912 0.396468431 0.7158 0.59625 + Calpastatin E7ESM9 CAST 1.476781033 1.149459479 0.720851913 0.081528203 0.2635765 0.53083 Mucin-2 E7EUV1 MUC2 2.760394361 2.073574027 0.503173452 0.128786237 0.3581141 0.6325 Acyl-CoA synthetase family member 2, mitochondrial E9PF16 ACSF2 1.026815768 0.304245583 0.485502819 0.210674608 -

Proteomics Analysis and Identi Cation of Critical Proteins and Network
Proteomics Analysis and Identication of Critical Proteins and Network Interactions that Regulate the Specic Deposition of IMF of Jingyuan Chicken Zengwen Huang ( [email protected] ) Ningxia University https://orcid.org/0000-0002-7223-4047 Juan Zhang Ningxia University Yaling Gu Ningxia University Guosheng Xin Ningxia University Zhengyun Cai Ningxia University Xianfeng Yin Neijiang Vocational and Technical College Xiaofang Feng Ningxia University Tong Mu Ningxia University Chaoyun Yang Ningxia University Research article Keywords: Jingyuan chicken, Leg muscle, Breast muscle, Meat quality, Proteomics, IMF Posted Date: December 10th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-54060/v2 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/27 Abstract Background: Improving broiler production eciency and delivering good quality chicken has become an exciting area of research. Many factors affect the quality of chicken, and the IMF content is one of the critical factors determining the quality of chicken. At present, there are many reports on the molecular mechanism of IMF-specic deposition in chicken; however, only a few reports discuss the specic deposition of IMF in different parts of a chicken. Methods: In order to analyze the molecular mechanism of IMF specic deposition in different parts of chickens, the present study has selected 180-days old Jingyuan chicken breast and leg muscles as the research materials, using proteomics technology and screening of PRM protein quantitative detection methods, identication and quantitative verication of proteins that control the IMF-specic deposition in the leg muscles and breast muscles of Jingyuan chickens. The protein was analyzed by advanced bioinformatics using GO, KEGG, R language, Gallus_gallus_UniPort, and other biological software, including tools and related databases for screening and identication. -

Thioesterase Induction by 2,3,7,8-Tetrachlorodibenzo-P-Dioxin
www.nature.com/scientificreports OPEN Thioesterase induction by 2,3, 7,8 ‑te tra chl orodib enz o‑ p ‑dioxin results in a futile cycle that inhibits hepatic β‑oxidation Giovan N. Cholico1,2, Russell R. Fling2,3, Nicholas A. Zacharewski1, Kelly A. Fader1,2, Rance Nault1,2 & Timothy R. Zacharewski1,2* 2,3,7,8‑Tetrachlorodibenzo‑p‑dioxin (TCDD), a persistent environmental contaminant, induces steatosis by increasing hepatic uptake of dietary and mobilized peripheral fats, inhibiting lipoprotein export, and repressing β‑oxidation. In this study, the mechanism of β‑oxidation inhibition was investigated by testing the hypothesis that TCDD dose‑dependently repressed straight‑chain fatty acid oxidation gene expression in mice following oral gavage every 4 days for 28 days. Untargeted metabolomic analysis revealed a dose‑dependent decrease in hepatic acyl‑CoA levels, while octenoyl‑ CoA and dicarboxylic acid levels increased. TCDD also dose‑dependently repressed the hepatic gene expression associated with triacylglycerol and cholesterol ester hydrolysis, fatty acid binding proteins, fatty acid activation, and 3‑ketoacyl‑CoA thiolysis while inducing acyl‑CoA hydrolysis. Moreover, octenoyl‑CoA blocked the hydration of crotonyl‑CoA suggesting short chain enoyl‑CoA hydratase (ECHS1) activity was inhibited. Collectively, the integration of metabolomics and RNA‑seq data suggested TCDD induced a futile cycle of fatty acid activation and acyl‑CoA hydrolysis resulting in incomplete β‑oxidation, and the accumulation octenoyl‑CoA levels that inhibited the activity of short chain enoyl‑CoA hydratase (ECHS1). Although the liver is the largest internal organ, it is second to adipose tissue in regard to lipid storage capacity. Approximately 15–25% (fasted vs fed state, respectively) are derived from chylomicron remnants, 5–30% (fasted vs fed state, respectively) from de novo lipogenesis, and 5–30% from visceral fat tissues 1. -

Comparative Transcriptome Profile Between Iberian Pig Varieties
animals Article Comparative Transcriptome Profile between Iberian Pig Varieties Provides New Insights into Their Distinct Fat Deposition and Fatty Acids Content Ana Villaplana-Velasco 1,2, Jose Luis Noguera 3, Ramona Natacha Pena 4 , Maria Ballester 5, Lourdes Muñoz 6, Elena González 7 , Juan Florencio Tejeda 7 and Noelia Ibáñez-Escriche 8,* 1 Genetics and Genomics, The Roslin Institute, Royal (Dick) School of Veterinary Studies, The University of Edinburgh, Easter Bush Campus, Midlothian, Edinburgh EH25 9RG, UK; [email protected] 2 Centre for Medical Informatics, Usher Institute, The University of Edinburgh, 9 Little France Road, Edinburgh EH16 4UX, UK 3 Animal Breeding and Genetics Program, IRTA, 25198 Lleida, Spain; [email protected] 4 Departament de Ciència Animal, Universitat de Lleida-Agrotecnio Center, 25198 Lleida, Spain; [email protected] 5 Animal Breeding and Genetics Program, IRTA, Torre Marimon, 08140 Caldes de Montbui, Spain; [email protected] 6 INGA FOOD S.A, 06200 Almendralejo, Spain; [email protected] 7 Department of Animal Production and Food Science, Research University Institute of Agricultural Resources (INURA), Escuela de Ingenierías Agrarias, Universidad de Extremadura, 06007 Badajoz, Spain; [email protected] (E.G.); [email protected] (J.F.T.) 8 Department for Animal Science and Tecnology, Universistat Politécnica de València, 46022 Valencia, Spain * Correspondence: [email protected]; Tel.: +34-963-877-438 Citation: Villaplana-Velasco, A.; Noguera, J.L.; Pena, R.N.; Ballester, Simple Summary: Iberian pigs are meat quality models due to their high fat content, high intra- M.; Muñoz, L.; González, E.; Tejeda, J.F.; Ibáñez-Escriche, N. -

Wasin Vol. 9 N. 4 2558.Pmd
Asian Biomedicine Vol. 9 No. 4 August 2015; 455 - 471 DOI: 10.5372/1905-7415.0904.415 Original article Antiaging phenotype in skeletal muscle after endurance exercise is associated with the oxidative phosphorylation pathway Wasin Laohavinija, Apiwat Mutirangurab,c aFaculty of Medicine, Chulalongkorn University, Bangkok 10330, Thailand bDepartment of Anatomy, Faculty of Medicine, Chulalongkorn University, Bangkok 10330, Thailand cCenter of Excellence in Molecular Genetics of Cancer and Human Diseases, Chulalongkorn University, Bangkok 10330, Thailand Background: Performing regular exercise may be beneficial to delay aging. During aging, numerous biochemical and molecular changes occur in cells, including increased DNA instability, epigenetic alterations, cell-signaling disruptions, decreased protein synthesis, reduced adenosine triphosphate (ATP) production capacity, and diminished oxidative phosphorylation. Objectives: To identify the types of exercise and the molecular mechanisms associated with antiaging phenotypes by comparing the profiles of gene expression in skeletal muscle after various types of exercise and aging. Methods: We used bioinformatics data from skeletal muscles reported in the Gene Expression Omnibus repository and used Connection Up- and Down-Regulation Expression Analysis of Microarrays to identify genes significant in antiaging. The significant genes were mapped to molecular pathways and reviewed for their molecular functions, and their associations with molecular and cellular phenotypes using the Database for Annotation, -

Variants in the 3€² Untranslated Region of the Ovine Acetyl
J. Dairy Sci. 100:6285–6297 https://doi.org/10.3168/jds.2016-12326 © 2017, THE AUTHORS. Published by FASS and Elsevier Inc. on behalf of the American Dairy Science Association®. This is an open access article under the CC BY-NC-ND license (http://creativecommons.org/licenses/by-nc-nd/3.0/). Variants in the 3′ untranslated region of the ovine acetyl-coenzyme A acyltransferase 2 gene are associated with dairy traits and exhibit differential allelic expression D. Miltiadou,*1,2 A. L. Hager-Theodorides,†1 S. Symeou,* C. Constantinou,* A. Psifidi,‡ G. Banos,‡§ and O. Tzamaloukas* *Department of Agricultural Sciences, Biotechnology and Food Science, Cyprus University of Technology, 3603 Lemesos, PO Box 50329, Cyprus †Department of Animal Science and Aquaculture, Agricultural University of Athens, 11855 Athens, Greece ‡The Roslin Institute and Royal (Dick) School of Veterinary Studies, University of Edinburgh, EH25 9RG Midlothian, United Kingdom §Animal and Veterinary Sciences, Scotland’s Rural College, EH25 9RG, Midlothian, United Kingdom ABSTRACT compared with the C allele in the udder of randomly selected ewes. We demonstrated for the first time that The acetyl-CoA acyltransferase 2 (ACAA2) gene en- the variants in the 3′ untranslated region of the ovine codes an enzyme of the thiolase family that is involved ACAA2 gene are differentially expressed in homozygous in mitochondrial fatty acid elongation and degradation ewes of each allele and exhibit allelic expression imbal- by catalyzing the last step of the respective β-oxidation ance within heterozygotes in a tissue-specific manner, pathway. The increased energy needs for gluconeogen- supporting the existence of cis-regulatory DNA varia- esis and triglyceride synthesis during lactation are met tion in the ovine ACAA2 gene. -

Mouse Acaa2 Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Acaa2 Knockout Project (CRISPR/Cas9) Objective: To create a Acaa2 knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Acaa2 gene (NCBI Reference Sequence: NM_177470 ; Ensembl: ENSMUSG00000036880 ) is located on Mouse chromosome 18. 10 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 10 (Transcript: ENSMUST00000041053). Exon 2~4 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 2 starts from about 1.43% of the coding region. Exon 2~4 covers 34.68% of the coding region. The size of effective KO region: ~6285 bp. The KO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 2 3 4 10 Legends Exon of mouse Acaa2 Knockout region Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of Exon 2 is aligned with itself to determine if there are tandem repeats. Tandem repeats are found in the dot plot matrix. The gRNA site is selected outside of these tandem repeats. Overview of the Dot Plot (down) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 1727 bp section downstream of Exon 4 is aligned with itself to determine if there are tandem repeats. -

ACAA2 Antibody Rabbit Polyclonal Antibody Catalog # ABV10394
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 ACAA2 Antibody Rabbit Polyclonal Antibody Catalog # ABV10394 Specification ACAA2 Antibody - Product Information Application WB Primary Accession P13437 Other Accession EDL82881.1 Reactivity Human, Mouse, Rat Host Rabbit Clonality Polyclonal Isotype Rabbit IgG Calculated MW 41871 ACAA2 Antibody - Additional Information Gene ID 170465 Positive Control Rat kidney tissue lysate Western blot analysis of ACAA2 using rat Application & Usage Western Blot kidney tissue lysate. analysis (1-4 µg/ml). However, the optimal ACAA2 Antibody - Background concentrations should be ACAA2 (acetylcoenzyme A acyltransferase 2), determined is member of the thiolase family of enzymes individually. and is involved in lipid metabolism. ACAA2 is Blocking peptide localized in the mitochondria. ACCA2 catalyzes is available the last conversion step of acetyl-CoA to separately. 3-oxoacyl-CoA, in the fatty acid oxidation Other Names pathway. Peroxisomal 3-oxoacyl-CoA thiolase A Target/Specificity ACAA2 Antibody Form Liquid Appearance Colorless liquid Formulation 100 µg (0.5 mg/ml) affinity purified rabbit anti-ACAA2 polyclonal antibody in phosphate buffered saline (PBS), pH 7.2, containing 30% glycerol, 0.5% BSA, 5 mM EDTA and 0.01% thimerosal. Page 1/3 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 Handling The antibody solution should be gently mixed before use. Reconstitution & Storage -20 °C Background Descriptions Precautions ACAA2 Antibody is for research use only and not for use in diagnostic or therapeutic procedures. ACAA2 Antibody - Protein Information Name Acaa2 Function In the production of energy from fats, this is one of the enzymes that catalyzes the last step of the mitochondrial beta- oxidation pathway, an aerobic process breaking down fatty acids into acetyl-CoA (Probable). -

Sex Impacts Cardiac Function and the Proteome Response to Thyroid Hormone in Aged Mice Wei Zhong Zhu1, Aaron Olson1,2, Michael Portman1,2 and Dolena Ledee1,2*
Zhu et al. Proteome Science (2020) 18:11 https://doi.org/10.1186/s12953-020-00167-3 RESEARCH Open Access Sex impacts cardiac function and the proteome response to thyroid hormone in aged mice Wei Zhong Zhu1, Aaron Olson1,2, Michael Portman1,2 and Dolena Ledee1,2* Abstract Background: Sex and age have substantial influence on thyroid function. Sex influences the risk and clinical expression of thyroid disorders (TDs), with age a proposed trigger for the development of TDs. Cardiac function is affected by thyroid hormone levels with gender differences. Accordingly, we investigated the proteomic changes involved in sex based cardiac responses to thyroid dysfunction in elderly mice. Methods: Aged (18–20 months) male and female C57BL/6 mice were fed diets to create euthyroid, hypothyroid, or hyperthyroid states. Serial echocardiographs were performed to assess heart function. Proteomic changes in cardiac protein profiles were assessed by 2-D DIGE and LC-MS/MS, and a subset confirmed by immunoblotting. Results: Serial echocardiographs showed ventricular function remained unchanged regardless of treatment. Heart rate and size increased (hyperthyroid) or decreased (hypothyroid) independent of sex. Pairwise comparison between the six groups identified 55 proteins (≥ 1.5-fold difference and p < 0.1). Compared to same-sex controls 26/55 protein changes were in the female hypothyroid heart, whereas 15/55 protein changes were identified in the male hypothyroid, and male and female hyperthyroid heart. The proteins mapped to oxidative phosphorylation, tissue remodeling and inflammatory response pathways. Conclusion: We identified both predicted and novel proteins with gender specific differential expression in response to thyroid hormone status, providing a catalogue of proteins associated with thyroid dysfunction.