Glucocorticoid Receptor-Ppara Axis in Fetal Mouse Liver Prepares
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gastronauts Symposium Booklet
DUKE / DUKE-NUSDUKE / DUKE-NUS GASTRONAUTSGASTRONAUTS SINGAPORESINGAPORE A GLOBAL SYMPOSIUMA GLOBAL ON GUT-BRAIN SYMPOSIUM MATTERS ON GUT-BRAIN MATTERS 3 - 4 MAY 20183 - 4 MAY 2018 CONTENTS 03 Message from Co-Chairs 04 Symposium Highlights 08 SESSION ONE Session Chair: Diego Bohórquez Speaker 1.1 Raphael H. Valdivia Speaker 1.2 Jonathan Kotula Speaker 1.3 Xiuqin Zhang Speaker 1.4 Matthew Chang 14 SESSION TWO Session Chair: Irene Miguel-Aliaga Speaker 2.1 Feifan Guo Speaker 2.2 Walter Wahli Speaker 2.3 Paul M Yen Speaker 2.4 Greg S. B. Suh 20 DISCUSSION Chair: Pae Wu Iain Dickson Jermont Chen Justin Gallivan Robert Kokoska 21 SESSION THREE Session Chair: Xiling Shen Speaker 3.1 John Furness Speaker 3.2 Arthur Beyder Speaker 3.3 Nicholas Spencer Speaker 3.4 Sven Pettersson CONTENTS 27 SESSION FOUR Session Chair: Raphael H. Valdivia Speaker 4.1 Nick Barker Speaker 4.2 Xiling Shen Speaker 4.3 Hyunsoo Shawn Je Speaker 4.4 Maxime M. Mahe 33 SESSION FIVE Session Chair: Nicholas Spencer Speaker 5.1 Lawrence Carin Speaker 5.2 John Doyle Speaker 5.3 Warren Grill 38 SESSION SIX Session Chair: Arthur Beyder Speaker 6.1 Karl Herrup Speaker 6.2 David L. Silver Speaker 6.3 Rodger A. Liddle 43 SESSION SEVEN Session Chair: John Furness Speaker 7.1 Diego Bohórquez Speaker 7.2 Luis R. Saraiva Speaker 7.3 Irene Miguel-Aliaga Speaker 7.4 Ivan de Araujo MESSAGE FROM CO-CHAIRS Dear Gastronauts, On behalf of the organising committee, we are thrilled to welcome you to Singapore: A place known for its cultural tapestry Diego Bohórquez Assistant Professor of Medicine and where people from around the globe and Neurobiology Duke University come to imagine new possibilities. -
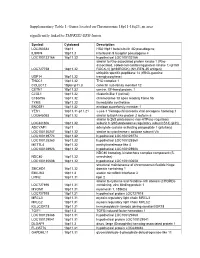
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Roles of Estrogens in the Healthy and Diseased Oviparous Vertebrate Liver
H OH metabolites OH Review Roles of Estrogens in the Healthy and Diseased Oviparous Vertebrate Liver Blandine Tramunt 1,2, Alexandra Montagner 1, Nguan Soon Tan 3 , Pierre Gourdy 1,2 , Hervé Rémignon 4,5 and Walter Wahli 3,5,6,* 1 Institut des Maladies Métaboliques et Cardiovasculaires (I2MC-UMR1297), INSERM/UPS, Université de Toulouse, F-31432 Toulouse, France; [email protected] (B.T.); [email protected] (A.M.); [email protected] (P.G.) 2 Service de Diabétologie, Maladies Métaboliques et Nutrition, CHU de Toulouse, F-31059 Toulouse, France 3 Lee Kong Chian School of Medicine, Nanyang Technological University Singapore, Clinical Sciences Building, Singapore 308232, Singapore; [email protected] 4 INP-ENSAT, Université de Toulouse, F-31320 Castanet-Tolosan, France; [email protected] 5 Toxalim Research Center in Food Toxicology (UMR 1331), INRAE, National Veterinary College of Toulouse (ENVT), Purpan College of Engineers of the Institut National Polytechnique de Toulouse (INP-PURPAN), Université Toulouse III—Paul Sabatier (UPS), Université de Toulouse, F-31300 Toulouse, France 6 Center for Integrative Genomics, Université de Lausanne, Le Génopode, CH-1015 Lausanne, Switzerland * Correspondence: [email protected] Abstract: The liver is a vital organ that sustains multiple functions beneficial for the whole organism. It is sexually dimorphic, presenting sex-biased gene expression with implications for the phenotypic differences between males and females. Estrogens are involved in this sex dimorphism and their actions in the liver of several reptiles, fishes, amphibians, and birds are discussed. The liver partici- pates in reproduction by producing vitellogenins (yolk proteins) and eggshell proteins under the Citation: Tramunt, B.; Montagner, A.; control of estrogens that act via two types of receptors active either mainly in the cell nucleus (ESR) Tan, N.S.; Gourdy, P.; Rémignon, H.; or the cell membrane (GPER1). -

Ppars and Microbiota in Skeletal Muscle Health and Wasting
International Journal of Molecular Sciences Review PPARs and Microbiota in Skeletal Muscle Health and Wasting Ravikumar Manickam 1 , Kalina Duszka 2 and Walter Wahli 3,4,5,* 1 Department of Pharmaceutical Sciences, University of South Florida, 12901 Bruce B. Downs Blvd., Tampa, FL 33612, USA; [email protected] 2 Department of Nutritional Sciences, University of Vienna, Althanstrasse 14, 1090 Vienna, Austria; [email protected] 3 Center for Integrative Genomics, University of Lausanne, Le Génopode, CH-1015 Lausanne, Switzerland 4 Toxalim, INRAE, Chemin de Tournefeuille 180, F-31027 Toulouse, France 5 Lee Kong Chian School of Medicine, Nanyang Technological University Singapore, Clinical Sciences Building, 11 Mandalay Road, Singapore 308232, Singapore * Correspondence: [email protected] Received: 18 October 2020; Accepted: 26 October 2020; Published: 29 October 2020 Abstract: Skeletal muscle is a major metabolic organ that uses mostly glucose and lipids for energy production and has the capacity to remodel itself in response to exercise and fasting. Skeletal muscle wasting occurs in many diseases and during aging. Muscle wasting is often accompanied by chronic low-grade inflammation associated to inter- and intra-muscular fat deposition. During aging, muscle wasting is advanced due to increased movement disorders, as a result of restricted physical exercise, frailty, and the pain associated with arthritis. Muscle atrophy is characterized by increased protein degradation, where the ubiquitin-proteasomal and autophagy-lysosomal pathways, atrogenes, and growth factor signaling all play an important role. Peroxisome proliferator-activated receptors (PPARs) are members of the nuclear receptor family of transcription factors, which are activated by fatty acids and their derivatives. -

Glucocorticoid Receptor-PPAR Alpha Axis in Fetal Mouse Liver Prepares
Glucocorticoid receptor-PPAR alpha axis in fetal mouse liver prepares neonates for milk lipid catabolism Gianpaolo Rando, Chek Kun Tan, Nourhène Khaled, Alexandra Montagner, Nicolas Leuenberger, Justine Bertrand-Michel, Eeswari Paramalingam, Hervé Guillou, Walter Wahli To cite this version: Gianpaolo Rando, Chek Kun Tan, Nourhène Khaled, Alexandra Montagner, Nicolas Leuenberger, et al.. Glucocorticoid receptor-PPAR alpha axis in fetal mouse liver prepares neonates for milk lipid catabolism. eLife, eLife Sciences Publication, 2016, 5, 31 p. 10.7554/eLife.11853. hal-01602433 HAL Id: hal-01602433 https://hal.archives-ouvertes.fr/hal-01602433 Submitted on 27 May 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License RESEARCH ARTICLE Glucocorticoid receptor-PPARa axis in fetal mouse liver prepares neonates for milk lipid catabolism Gianpaolo Rando1†, Chek Kun Tan2†, Nourhe` ne Khaled1, Alexandra Montagner3, Nicolas Leuenberger1, Justine Bertrand-Michel4, Eeswari Paramalingam2, Herve´ -

Homozygosity Mapping of a Dyggve-Melchior-Clausen
714 ORIGINAL ARTICLE J Med Genet: first published as 10.1136/jmg.39.10.714 on 1 October 2002. Downloaded from Homozygosity mapping of a Dyggve-Melchior-Clausen syndrome gene to chromosome 18q21.1 C Thauvin-Robinet, V El Ghouzzi, W Chemaitilly, N Dagoneau, O Boute, G Viot, A Mégarbané, A Sefiani, A Munnich, M Le Merrer, V Cormier-Daire ............................................................................................................................. See end of article for J Med Genet 2002;39:714–717 authors’ affiliations ....................... Correspondence to: Dr V Cormier-Daire, Dyggve-Melchior-Clausen syndrome (DMC) is an autosomal recessive condition characterised by short Département de trunk dwarfism, scoliosis, microcephaly, coarse facies, mental retardation, and characteristic Génétique, Hôpital radiological features. X rays show platyspondyly with double vertebral hump, epiphyseal dysplasia, Necker-Enfants Malades, irregular metaphyses, and a characteristic lacy appearance of the iliac crests. Electron microscopy of 149 rue de Sèvres, 75015 chondrocytes have shown widened cisternae of rough endoplasmic reticulum and biochemical analy- Paris, France; [email protected] ses have shown accumulation of glucosaminoglycan in cartilage, but the pathogenesis of DMC remains unexplained. Here, we report on the homozygosity mapping of a DMC gene to chromosome 18q21.1 Revised version received in seven inbred families (Zmax=9.65 at θ=0 at locus D18S1126) in the genetic interval (1.8 cM) 6 August 2002 Accepted for publication defined by loci D18S455 and D18S363. Despite the various geographical origins of the families 13 August 2002 reported here (Morocco, Tunisia, Portugal, and Lebanon), this condition was genetically homogeneous ....................... in our series. Continuing studies will hopefully lead to the identification of the disease causing gene. -

Walter Wahli, Metabolomics 2017, 7:2(Suppl) Conferenceseries.Com
Walter Wahli, Metabolomics 2017, 7:2(Suppl) conferenceseries.com http://dx.doi.org/10.4172/2153-0769-C1-036 8th International Conference and Exhibition on Metabolomics & Systems Biology May 08-10, 2017 Singapore Walter Wahli Nanyang Technological University, Singapore PPARα in hepatocytes integrates microbiome-derived signals and is protective against fatty liver in both neonate and adult mice he liver is a key organ of metabolic homeostasis with functions that oscillate in response to food intake. Germ-free mice Tdisplay altered daily oscillation of clock gene expression with a change in the expression of clock output regulators. These alterations in microbiome-sensitive gene expression are associated with daily alterations in lipid, glucose and xenobiotic metabolism as revealed by hepatic metabolome analyses. Hepatic lipid catabolism is essential for the newborns to use milk fat as an energy source. PPARα in hepatocytes is critical for this function. PPARα expression is stimulated a few days before birth, which prepares the receptor for its physiological role in harnessing milk lipids after birth. This mechanism involves a fetal glucocorticoid receptor (GR)-PPARα axis in which GR directly binds to the Pparα promoter to stimulate its activity. In turn, under the control of PPARα, the expression of genes required for lipid catabolism is enhanced before birth so that the neonatal liver has a prompt capacity to extract energy from milk upon suckling. Interestingly, the PPARα target gene Fgf21 is not stimulated in the fetal liver and responds to PPARα only after birth following an epigenetic switch triggered by β-hydroxybutyrate-mediated inhibition of histone deacetylase 3. -

Discovery Protein Names Uniprot Gene Names Average Log2 L/H
Discovery Ttest IBD vs FDR Average Log2 Average Log2 Ratio Signficant Control p adjusted p Subgroup Protein names Uniprot Gene names L/H ratio Control L/H ratio IBD ibd/control 0.05 FDR value value AUC Specific Immunolgical Aconitate hydratase, mitochondrial A2A274 ACO2 2.570537714 1.404944787 0.311737769 + 1.26481E-09 8.472E-08 0.92417 CD9 antigen A6NNI4 CD9 2.690874857 1.62514501 0.344476348 + 3.02524E-10 2.606E-08 0.95583 Calponin-2 B4DDF4 CNN2 0.388970715 0.995038293 1.833208258 + 0.000445883 0.0044168 0.755 6-phosphogluconate dehydrogenase, decarboxylating B4DQJ8 PGD -0.252090674 0.391282139 1.902888154 + 1.35886E-13 8.373E-11 0.96625 Epithelial cell adhesion molecule B5MCA4 EPCAM 3.405798843 2.468926837 0.39185163 + 0.002975231 0.0215657 0.80625 2,4-dienoyl-CoA reductase, mitochondrial B7Z6B8 DECR1 2.240008 1.557225501 0.50520929 + 0.003411719 0.023728 0.72042 Adenosylhomocysteinase;Putative adenosylhomocysteinase 3 D7UEQ7 AHCYL2 2.14170094 1.336080036 0.446810414 + 0.000222911 0.0024884 0.84583 CD44 antigen E7EPC6 CD44 1.30554758 1.619074953 1.368242916 0.094110279 0.2905348 0.62625 Lactotransferrin;Lactoferricin-H;Kaliocin-1;Lactoferroxin-A;Lactoferroxin- B;Lactoferroxin-C E7ER44 LTF 1.276691743 1.810737039 1.705818912 0.396468431 0.7158 0.59625 + Calpastatin E7ESM9 CAST 1.476781033 1.149459479 0.720851913 0.081528203 0.2635765 0.53083 Mucin-2 E7EUV1 MUC2 2.760394361 2.073574027 0.503173452 0.128786237 0.3581141 0.6325 Acyl-CoA synthetase family member 2, mitochondrial E9PF16 ACSF2 1.026815768 0.304245583 0.485502819 0.210674608 -

Proteomics Analysis and Identi Cation of Critical Proteins and Network
Proteomics Analysis and Identication of Critical Proteins and Network Interactions that Regulate the Specic Deposition of IMF of Jingyuan Chicken Zengwen Huang ( [email protected] ) Ningxia University https://orcid.org/0000-0002-7223-4047 Juan Zhang Ningxia University Yaling Gu Ningxia University Guosheng Xin Ningxia University Zhengyun Cai Ningxia University Xianfeng Yin Neijiang Vocational and Technical College Xiaofang Feng Ningxia University Tong Mu Ningxia University Chaoyun Yang Ningxia University Research article Keywords: Jingyuan chicken, Leg muscle, Breast muscle, Meat quality, Proteomics, IMF Posted Date: December 10th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-54060/v2 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/27 Abstract Background: Improving broiler production eciency and delivering good quality chicken has become an exciting area of research. Many factors affect the quality of chicken, and the IMF content is one of the critical factors determining the quality of chicken. At present, there are many reports on the molecular mechanism of IMF-specic deposition in chicken; however, only a few reports discuss the specic deposition of IMF in different parts of a chicken. Methods: In order to analyze the molecular mechanism of IMF specic deposition in different parts of chickens, the present study has selected 180-days old Jingyuan chicken breast and leg muscles as the research materials, using proteomics technology and screening of PRM protein quantitative detection methods, identication and quantitative verication of proteins that control the IMF-specic deposition in the leg muscles and breast muscles of Jingyuan chickens. The protein was analyzed by advanced bioinformatics using GO, KEGG, R language, Gallus_gallus_UniPort, and other biological software, including tools and related databases for screening and identication. -

Thioesterase Induction by 2,3,7,8-Tetrachlorodibenzo-P-Dioxin
www.nature.com/scientificreports OPEN Thioesterase induction by 2,3, 7,8 ‑te tra chl orodib enz o‑ p ‑dioxin results in a futile cycle that inhibits hepatic β‑oxidation Giovan N. Cholico1,2, Russell R. Fling2,3, Nicholas A. Zacharewski1, Kelly A. Fader1,2, Rance Nault1,2 & Timothy R. Zacharewski1,2* 2,3,7,8‑Tetrachlorodibenzo‑p‑dioxin (TCDD), a persistent environmental contaminant, induces steatosis by increasing hepatic uptake of dietary and mobilized peripheral fats, inhibiting lipoprotein export, and repressing β‑oxidation. In this study, the mechanism of β‑oxidation inhibition was investigated by testing the hypothesis that TCDD dose‑dependently repressed straight‑chain fatty acid oxidation gene expression in mice following oral gavage every 4 days for 28 days. Untargeted metabolomic analysis revealed a dose‑dependent decrease in hepatic acyl‑CoA levels, while octenoyl‑ CoA and dicarboxylic acid levels increased. TCDD also dose‑dependently repressed the hepatic gene expression associated with triacylglycerol and cholesterol ester hydrolysis, fatty acid binding proteins, fatty acid activation, and 3‑ketoacyl‑CoA thiolysis while inducing acyl‑CoA hydrolysis. Moreover, octenoyl‑CoA blocked the hydration of crotonyl‑CoA suggesting short chain enoyl‑CoA hydratase (ECHS1) activity was inhibited. Collectively, the integration of metabolomics and RNA‑seq data suggested TCDD induced a futile cycle of fatty acid activation and acyl‑CoA hydrolysis resulting in incomplete β‑oxidation, and the accumulation octenoyl‑CoA levels that inhibited the activity of short chain enoyl‑CoA hydratase (ECHS1). Although the liver is the largest internal organ, it is second to adipose tissue in regard to lipid storage capacity. Approximately 15–25% (fasted vs fed state, respectively) are derived from chylomicron remnants, 5–30% (fasted vs fed state, respectively) from de novo lipogenesis, and 5–30% from visceral fat tissues 1. -

The Center for Integrative Genomics Report 2005–2006
The Center for integrative genomics Report 2005–2006 $FOUSFJOUÏHSBUJG EFHÏOPNJRVF Index PRESENTATION Director’s message 4 Scientific advisory committee 6 Organigram of the CIG 7 RESEARCH The structure and function of genomes and their evolution Alexandre reymond – Genome structure and expression 10 Henrik Kaessmann – Evolutionary genomics 12 Victor Jongeneel – Cancer genomics 14 The regulation of gene expression Nouria hernandez – Mechanisms of transcription regulation 16 Winship herr – Regulation of cell proliferation 18 Christian Fankhauser – Light–regulated development in plants 20 The genomics of complex functions Mehdi tafti – Genetics of sleep and the sleep EEG 22 Paul Franken – Genetics and energetics of sleep homeostasis and circadian rythms 24 Walter Wahli – Peroxisome Proliferator–Activated Receptors (PPARs) as regulators of metabolic and tissue repair processes 26 Béatrice Desvergne – PPARbeta and fine tuning of cell fate decision 30 Liliane Michalik – Roles of PPARs in skin biology and angiogenesis 34 Bernard thorens – Molecular and physiological analysis of energy homeostasis in health and disease 36 CORE fACILITIES Core facilities of the CIG 39 DNA array facility – DAF 40 Protein analysis facility – PAF 42 Core facilities associated with the CIG 44 EdUCATION Lectures and courses 48 Seminars at the CIG 51 CIG retreat 56 The CIG & the public 56 FUNDING (ACKNOwlEDGEMENTS) 58 PEOplE 59 CIG Timeline CIG Timeline Inauguration of the Pro- tein Analysis Facility (PAF – December 2002) Inauguration of the DNA Array Facility (DAF – March 2003) Arrival of Prof. Kaessmann (September 2003) Installation Arrival of of Vital–IT Prof. Thorens in the Génopode (October 2003) Arrival of Inauguration of the Prof. Hernandez The construction of Prof. Hernandez, Cellular Imaging Facility becomes the second the big animal facility Herr and Tafti at the Génopode CIG director is rejected by popular (September 2004) referendum Arrival of Arrival of Arrival Arrival Inauguration First CIG retreat Prof. -

Comparative Transcriptome Profile Between Iberian Pig Varieties
animals Article Comparative Transcriptome Profile between Iberian Pig Varieties Provides New Insights into Their Distinct Fat Deposition and Fatty Acids Content Ana Villaplana-Velasco 1,2, Jose Luis Noguera 3, Ramona Natacha Pena 4 , Maria Ballester 5, Lourdes Muñoz 6, Elena González 7 , Juan Florencio Tejeda 7 and Noelia Ibáñez-Escriche 8,* 1 Genetics and Genomics, The Roslin Institute, Royal (Dick) School of Veterinary Studies, The University of Edinburgh, Easter Bush Campus, Midlothian, Edinburgh EH25 9RG, UK; [email protected] 2 Centre for Medical Informatics, Usher Institute, The University of Edinburgh, 9 Little France Road, Edinburgh EH16 4UX, UK 3 Animal Breeding and Genetics Program, IRTA, 25198 Lleida, Spain; [email protected] 4 Departament de Ciència Animal, Universitat de Lleida-Agrotecnio Center, 25198 Lleida, Spain; [email protected] 5 Animal Breeding and Genetics Program, IRTA, Torre Marimon, 08140 Caldes de Montbui, Spain; [email protected] 6 INGA FOOD S.A, 06200 Almendralejo, Spain; [email protected] 7 Department of Animal Production and Food Science, Research University Institute of Agricultural Resources (INURA), Escuela de Ingenierías Agrarias, Universidad de Extremadura, 06007 Badajoz, Spain; [email protected] (E.G.); [email protected] (J.F.T.) 8 Department for Animal Science and Tecnology, Universistat Politécnica de València, 46022 Valencia, Spain * Correspondence: [email protected]; Tel.: +34-963-877-438 Citation: Villaplana-Velasco, A.; Noguera, J.L.; Pena, R.N.; Ballester, Simple Summary: Iberian pigs are meat quality models due to their high fat content, high intra- M.; Muñoz, L.; González, E.; Tejeda, J.F.; Ibáñez-Escriche, N.