Cocaine-Induced Impulsivity Is Differentially Expressed in Male
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Case-Control Study of GRIA1 and GRIA3 Gene Variants in Migraine Jie Fang1, Xingkai An1, Shuai Chen2, Zhenzhen Yu1, Qilin Ma1,2* and Hongli Qu1*
Fang et al. The Journal of Headache and Pain (2016) 17:2 DOI 10.1186/s10194-016-0592-2 RESEARCH ARTICLE Open Access Case-control study of GRIA1 and GRIA3 gene variants in migraine Jie Fang1, Xingkai An1, Shuai Chen2, Zhenzhen Yu1, Qilin Ma1,2* and Hongli Qu1* Abstract Background: As the most abundant excitatory neurotransmitter in the central nervous system, glutamate has been accepted to play a major role in the pathophysiology of migraine. The previous studies have reported the glutamate receptor ionotropic GRIA1 and GRIA3 genes variants associated with migraine. The project aims to investigate the polymorphisms in both genes for their association with migraine in the Chinese Han population. Methods: A Han-Chinese case-control population, including 331 unrelated female migraine patients and 330 matched controls, was studied. Variants in genes (GRIA1 and GRIA3) were genotyped by Multiplex SNaPshot assay. Results: In the group of patients, the frequency of allele C was 84.1 % (557 C alleles) and allele T was 15.9 % (105 T alleles) for the GRIA1 (rs2195450) in migraineurs, this was significantly as compared with the controls (P = .001, OR = 1.786, 95 % CI: 1.28–2.49). And an association was also seen in the migraine with aura (MA) subtype (P = .012, OR = 2.092, 95 % CI: 1.17–3.76) and migraine without aura (MO) subtype (P =.002,OR=1.737, 95 % CI: 1.23–2.45). However, no evidence was found that GRIA1 (rs548294) or GRIA3 (rs3761555) is associated with migraine. Conclusion: Our data of this study confirmed the association of GRIA1 (rs2195450) to female migraine (MA, MO) susceptibility in the Chinese Han population. -

High Resolution Genomic Scans Reveal Genetic Architecture Controlling Alcohol Preference in Bidirectionally Selected Rat Model
Purdue University Purdue e-Pubs Department of Animal Sciences Faculty Department of Animal Sciences Publications 2016 High Resolution Genomic Scans Reveal Genetic Architecture Controlling Alcohol Preference in Bidirectionally Selected Rat Model Chiao-Ling Lo Indiana University School of Medicine Amy C. Lossie Indiana University School of Medicine Tiebing Liang Indiana University School of Medicine Yunlong Liu Indiana University School of Medicine Xiaoling Xuei Indiana University School of Medicine See next page for additional authors Follow this and additional works at: http://docs.lib.purdue.edu/anscpubs Recommended Citation Lo C-L, Lossie AC, Liang T, Liu Y, Xuei X, Lumeng L, et al. (2016) High Resolution Genomic Scans Reveal Genetic Architecture Controlling Alcohol Preference in Bidirectionally Selected Rat Model. PLoS Genet 12(8): e1006178. https://doi.org/10.1371/ journal.pgen.1006178 This document has been made available through Purdue e-Pubs, a service of the Purdue University Libraries. Please contact [email protected] for additional information. Authors Chiao-Ling Lo, Amy C. Lossie, Tiebing Liang, Yunlong Liu, Xiaoling Xuei, Lawrence Lumeng, Feng C. Zhou, and William M. Muir This article is available at Purdue e-Pubs: http://docs.lib.purdue.edu/anscpubs/19 RESEARCH ARTICLE High Resolution Genomic Scans Reveal Genetic Architecture Controlling Alcohol Preference in Bidirectionally Selected Rat Model Chiao-Ling Lo1,2, Amy C. Lossie1,3¤, Tiebing Liang1,4, Yunlong Liu1,5, Xiaoling Xuei1,6, Lawrence Lumeng1,4, Feng C. Zhou1,2,7*, William -

Sex Differences in Glutamate Receptor Gene Expression in Major Depression and Suicide
Molecular Psychiatry (2015) 20, 1057–1068 © 2015 Macmillan Publishers Limited All rights reserved 1359-4184/15 www.nature.com/mp IMMEDIATE COMMUNICATION Sex differences in glutamate receptor gene expression in major depression and suicide AL Gray1, TM Hyde2,3, A Deep-Soboslay2, JE Kleinman2 and MS Sodhi1,4 Accumulating data indicate that the glutamate system is disrupted in major depressive disorder (MDD), and recent clinical research suggests that ketamine, an antagonist of the N-methyl-D-aspartate (NMDA) glutamate receptor (GluR), has rapid antidepressant efficacy. Here we report findings from gene expression studies of a large cohort of postmortem subjects, including subjects with MDD and controls. Our data reveal higher expression levels of the majority of glutamatergic genes tested in the dorsolateral prefrontal cortex (DLPFC) in MDD (F21,59 = 2.32, P = 0.006). Posthoc data indicate that these gene expression differences occurred mostly in the female subjects. Higher expression levels of GRIN1, GRIN2A-D, GRIA2-4, GRIK1-2, GRM1, GRM4, GRM5 and GRM7 were detected in the female patients with MDD. In contrast, GRM5 expression was lower in male MDD patients relative to male controls. When MDD suicides were compared with MDD non-suicides, GRIN2B, GRIK3 and GRM2 were expressed at higher levels in the suicides. Higher expression levels were detected for several additional genes, but these were not statistically significant after correction for multiple comparisons. In summary, our analyses indicate a generalized disruption of the regulation of the GluRs in the DLPFC of females with MDD, with more specific GluR alterations in the suicides and in the male groups. -
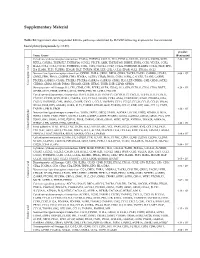
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

Research Article Microarray-Based Comparisons of Ion Channel Expression Patterns: Human Keratinocytes to Reprogrammed Hipscs To
Hindawi Publishing Corporation Stem Cells International Volume 2013, Article ID 784629, 25 pages http://dx.doi.org/10.1155/2013/784629 Research Article Microarray-Based Comparisons of Ion Channel Expression Patterns: Human Keratinocytes to Reprogrammed hiPSCs to Differentiated Neuronal and Cardiac Progeny Leonhard Linta,1 Marianne Stockmann,1 Qiong Lin,2 André Lechel,3 Christian Proepper,1 Tobias M. Boeckers,1 Alexander Kleger,3 and Stefan Liebau1 1 InstituteforAnatomyCellBiology,UlmUniversity,Albert-EinsteinAllee11,89081Ulm,Germany 2 Institute for Biomedical Engineering, Department of Cell Biology, RWTH Aachen, Pauwelstrasse 30, 52074 Aachen, Germany 3 Department of Internal Medicine I, Ulm University, Albert-Einstein Allee 11, 89081 Ulm, Germany Correspondence should be addressed to Alexander Kleger; [email protected] and Stefan Liebau; [email protected] Received 31 January 2013; Accepted 6 March 2013 Academic Editor: Michael Levin Copyright © 2013 Leonhard Linta et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Ion channels are involved in a large variety of cellular processes including stem cell differentiation. Numerous families of ion channels are present in the organism which can be distinguished by means of, for example, ion selectivity, gating mechanism, composition, or cell biological function. To characterize the distinct expression of this group of ion channels we have compared the mRNA expression levels of ion channel genes between human keratinocyte-derived induced pluripotent stem cells (hiPSCs) and their somatic cell source, keratinocytes from plucked human hair. This comparison revealed that 26% of the analyzed probes showed an upregulation of ion channels in hiPSCs while just 6% were downregulated. -

Identification of Key Genes and Pathways Involved in Response To
Deng et al. Biol Res (2018) 51:25 https://doi.org/10.1186/s40659-018-0174-7 Biological Research RESEARCH ARTICLE Open Access Identifcation of key genes and pathways involved in response to pain in goat and sheep by transcriptome sequencing Xiuling Deng1,2†, Dong Wang3†, Shenyuan Wang1, Haisheng Wang2 and Huanmin Zhou1* Abstract Purpose: This aim of this study was to investigate the key genes and pathways involved in the response to pain in goat and sheep by transcriptome sequencing. Methods: Chronic pain was induced with the injection of the complete Freund’s adjuvant (CFA) in sheep and goats. The animals were divided into four groups: CFA-treated sheep, control sheep, CFA-treated goat, and control goat groups (n 3 in each group). The dorsal root ganglions of these animals were isolated and used for the construction of a cDNA= library and transcriptome sequencing. Diferentially expressed genes (DEGs) were identifed in CFA-induced sheep and goats and gene ontology (GO) enrichment analysis was performed. Results: In total, 1748 and 2441 DEGs were identifed in CFA-treated goat and sheep, respectively. The DEGs identi- fed in CFA-treated goats, such as C-C motif chemokine ligand 27 (CCL27), glutamate receptor 2 (GRIA2), and sodium voltage-gated channel alpha subunit 3 (SCN3A), were mainly enriched in GO functions associated with N-methyl- D-aspartate (NMDA) receptor, infammatory response, and immune response. The DEGs identifed in CFA-treated sheep, such as gamma-aminobutyric acid (GABA)-related DEGs (gamma-aminobutyric acid type A receptor gamma 3 subunit [GABRG3], GABRB2, and GABRB1), SCN9A, and transient receptor potential cation channel subfamily V member 1 (TRPV1), were mainly enriched in GO functions related to neuroactive ligand-receptor interaction, NMDA receptor, and defense response. -

RUNNING HEAD: Low Homozygosity in Spanish Sheep
1 RUNNING HEAD: Low homozygosity in Spanish sheep 2 3 Low genome-wide homozygosity in eleven Spanish ovine breeds 4 5 M.G. Luigi 1, T.F. Cardoso 1, 2 , A. Martínez 3, A. Pons 4, L.A. Bermejo 5, J. Jordana 6, J.V. 6 Delgado 3, S. Adán 7, E. Ugarte 8, J. J. Arranz 9, J. Casellas 6 and M. Amills 1, 6 * 7 8 1Department of Animal Genetics, Centre for Research in Agricultural Genomics 9 (CRAG), CSIC-IRTA-UAB-UB, Campus Universitat Autònoma de Barcelona, 10 Bellaterra 08193, Spain; 2CAPES Foundation, Ministry of Education of Brazil, Brasilia 11 D. F., 70.040-020, Brazil; 3Departamento de Genética, Universidad de Córdoba, 12 Córdoba 14071, Spain; 4Unitat de Races Autòctones, Servei de Millora Agrària i 13 Pesquera (SEMILLA), Son Ferriol 07198, Spain; 5Departamento de Ingeniería, 14 Producción y Economía Agrarias, Universidad de La Laguna, 38071 La Laguna, 15 Tenerife, Spain; 6Departament de Ciència Animal i dels Aliments, Facultat de 16 Veterinària, Universitat Autònoma de Barcelona, Bellaterra 08193, Spain; 7Federación 17 de Razas Autóctonas de Galicia (BOAGA), Pazo de Fontefiz, 32152 Coles. Ourense, 18 Spain; 8Neiker-Tecnalia, Campus Agroalimentario de Arkaute, apdo 46 E-01080 19 Vitoria-Gazteiz (Araba), Spain; 9Departamento de Producción Animal, Universidad de 20 León, León 24071, Spain. 21 22 Corresponding author: Marcel Amills. Department of Animal Genetics, Centre for 23 Research in Agricultural Genomics (CRAG), CSIC-IRTA-UAB-UB, Campus 24 Universitat Autònoma de Barcelona, Bellaterra 08193, Spain. Tel. 34 93 5636600. E- 25 mail: [email protected]. 26 Abstract 27 28 The population of Spanish sheep has decreased from 24 to 15 million heads in 29 the last 75 years due to multiple social and economic factors. -

Persistent Pain Maintains Morphine-Seeking Behavior After Morphine Withdrawal Through Reduced Mecp2 Repression of Glua1 in Rat Central Amygdala
The Journal of Neuroscience, February 25, 2015 • 35(8):3689–3700 • 3689 Behavioral/Cognitive Persistent Pain Maintains Morphine-Seeking Behavior after Morphine Withdrawal through Reduced MeCP2 Repression of Glua1 in Rat Central Amygdala Yuan-Yuan Hou, You-Qing Cai, and Zhizhong Z. Pan Department of Anesthesiology and Pain Medicine, The University of Texas MD Anderson Cancer Center, Houston, Texas 77030 As long-term opioids are increasingly used for control of chronic pain, how pain affects the rewarding effect of opioids and hence risk of prescription opioid misuse and abuse remains a healthcare concern and a challenging issue in current pain management. In this study, using a rat model of morphine self-administration, we investigated the molecular mechanisms underlying the impact of pain on operant behavior of morphine intake and morphine seeking before and after morphine withdrawal. We found that rats with persistent pain consumed a similar amount of daily morphine to that in control rats without pain, but maintained their level-pressing behavior of morphine seeking after abstinence of morphine at 0.2 mg/kg, whereas this behavior was gradually diminished in control rats. In the central nucleus of amygdala (CeA), a limbic structure critically involved in the affective dimension of pain, proteins of GluA1 subunits of glutamateAMPAreceptorswereupregulatedduringmorphinewithdrawal,andviralknockdownofCeAGluA1eliminatedthemorphine- seeking behavior in withdrawn rats of the pain group. Chromatin immunoprecipitation analysis revealed that the methyl CpG-binding protein 2 (MeCP2) was enriched in the promoter region of Gria1 encoding GluA1 and this enrichment was significantly attenuated in withdrawn rats of the pain group. Furthermore, viral overexpression of CeA MeCP2 repressed the GluA1 level and eliminated the maintenanceofmorphine-seekingbehavioraftermorphinewithdrawal.TheseresultssuggestdirectMeCp2repressionofGluA1function as a likely mechanism for morphine-seeking behavior maintained by long-lasting affective pain after morphine withdrawal. -

GRIPAP1 / GRASP1 Antibody (N-Terminus) Goat Polyclonal Antibody Catalog # ALS14943
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 GRIPAP1 / GRASP1 Antibody (N-Terminus) Goat Polyclonal Antibody Catalog # ALS14943 Specification GRIPAP1 / GRASP1 Antibody (N-Terminus) - Product Information Application IHC Primary Accession Q4V328 Reactivity Human Host Goat Clonality Polyclonal Calculated MW 96kDa KDa GRIPAP1 / GRASP1 Antibody (N-Terminus) - Additional Information Gene ID 56850 Anti-GRIPAP1 / GRASP1 antibody IHC of Other Names human brain, cortex. GRIP1-associated protein 1, GRASP-1, GRIPAP1, KIAA1167 Target/Specificity Human GRIPAP1 / GRASP1. This antibody is expected to recognize both reported isoforms (as represented by NP_064522.3; NP_997555.1) Reconstitution & Storage Store at -20°C. Minimize freezing and thawing. Precautions GRIPAP1 / GRASP1 Antibody (N-Terminus) is for research use only and not for use in Anti-GRIPAP1 / GRASP1 antibody IHC of diagnostic or therapeutic procedures. human tonsil. GRIPAP1 / GRASP1 Antibody (N-Terminus) GRIPAP1 / GRASP1 Antibody (N-Terminus) - Protein Information - References Bechtel S.,et al.BMC Genomics Name GRIPAP1 (HGNC:18706) 8:399-399(2007). Ross M.T.,et al.Nature 434:325-337(2005). Synonyms KIAA1167 Hirosawa M.,et al.DNA Res. 6:329-336(1999). Function Lubec G.,et al.Submitted (DEC-2008) to Regulates the endosomal recycling back to UniProtKB. the neuronal plasma membrane, possibly Holt L.J.,et al.Nucleic Acids Res. by connecting early and late recycling 28:72-72(2000). endosomal domains and promoting Page 1/2 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 segregation of recycling endosomes from early endosomal membranes. -

Ion Channels
UC Davis UC Davis Previously Published Works Title THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels. Permalink https://escholarship.org/uc/item/1442g5hg Journal British journal of pharmacology, 176 Suppl 1(S1) ISSN 0007-1188 Authors Alexander, Stephen PH Mathie, Alistair Peters, John A et al. Publication Date 2019-12-01 DOI 10.1111/bph.14749 License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: Ion channels. British Journal of Pharmacology (2019) 176, S142–S228 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels Stephen PH Alexander1 , Alistair Mathie2 ,JohnAPeters3 , Emma L Veale2 , Jörg Striessnig4 , Eamonn Kelly5, Jane F Armstrong6 , Elena Faccenda6 ,SimonDHarding6 ,AdamJPawson6 , Joanna L Sharman6 , Christopher Southan6 , Jamie A Davies6 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 3Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 4Pharmacology and Toxicology, Institute of Pharmacy, University of Innsbruck, A-6020 Innsbruck, Austria 5School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 6Centre for Discovery Brain Science, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. -

Gene List of the Targeted NGS MCD and CCA Gene Panel AKT3,ALX1
Gene List of the targeted NGS MCD and CCA gene panel AKT3,ALX1,ALX3,ALX4,AMPD2,ARFGEF2,ARID1B,ARX,ASPM,ATR,ATRX,B3GALTL,BRPF1,c12orf57,C6orf70,CASK,CCND2,CDK5RAP2,CDON,C ENPJ,CEP170,CHMP1A,COL4A1,CREBBP,CYP11A1,DCHS1,DCLK1,DCX,DHCR24,DHCR7,DIS3L2,DISC1,DISP1,DLL1,DMRTA2,DYNC1H1,DYRK1 A,EARS2,EFNB1,EMX1,EOMES,EP300,ERBB4,ERMARD,EXOSC3,FAM36A,FGF8,FGFR1,FGFR2,FLNA,FOXC1,FOXG1,FOXH1,FZD10,GLI2,GLI3,GP R56,GPSM2,HCCS,HESX1,HNRNPU,IGBP1,IGFBP1,ISPD,ITPA,KAL1,KAT6B,KATNB1,KIAA1279,KIF14,KIF1A,KIF1B,KIF21A,KIF2A,KIF5C,KIF7,L1 CAM,LAMB1,LAMC3,LRP2,MCPH1,MED12,MID1,NDE1,NFIB,NPC1,NR2F1,NSD1,NTRK1,NTRK3,OCEL1,OPA1,OTX2,PAFAH1B1,PAX6,PEX1,PHF1 0,PIK3R2,POLR3A,POLR3B,POMT1,POMT2,PTCH1,PTPRS,PYCR1,RAB3GAP1,RARS2,RELN,RFX3,ROBO1,ROBO3,RPS6KA3,RTTN,SATB2,SEPSEC S,SHH,SIX3,SLC12A6,SOX2,SPOCK1,SRPX2,TBCD,TBCE,TCF4,TDGF1,TEAD1,THBS2,TMEM5,TSC1,TSC2,TSEN15,TSEN2,TSEN34,TSEN54,TUBA1 A,TUBA8,TUBB,TUBB2A,TUBB2B,TUBB3,TUBB4A,TUBG1,VAX1,VRK1,WDR47,WDR62,ZBTB18,ZEB2,ZIC2. Gene List of the targeted NGS epilepsy gene panel AARS, ADGRV1, ADRA2B, ADSL, ALDH4A1, ALDH7A1, ALG13, ALPL, ARHGEF15, ARHGEF9, ARX, ASAH1, ATP1A2, ATP1A3, BRD2, CACNA1A, CACNA1H, CACNA2D2, CACNB4, CBL, CDKL5, CERS1, CHD2, CHRNA2, CHRNA4, CHRNB2, CLCN2, CLCN4, CLN8, CLTC, CNKSR2, CNTNAP2, CPA6, CPLX1, CSNK1G1, CSNK2B, CTNND2, DEPDC5, DHDDS, DNM1, DOCK7, DYNC1H1, EEF1A2, EFHC1, EIF2S3, EMC1, EPM2A, FASN, FLNA, FOXG1, GABBR2, GABRA1, GABRA2, GABRA3, GABRB2, GABRB3, GABRD, GABRG2, GAL, GNAO1, GOSR2, GRIA1, GRIN1, GRIN2A, GRIN2B, HCN1, HCN4, HDAC4, HNRNPU, IDH3A, IQSEC2, JRK, KCNA1, KCNA2, KCNB1, -

Glutamate Receptors Within the Mesolimbic Dopamine System Mediate Alcohol Relapse Behavior
The Journal of Neuroscience, November 25, 2015 • 35(47):15523–15538 • 15523 Neurobiology of Disease Glutamate Receptors within the Mesolimbic Dopamine System Mediate Alcohol Relapse Behavior Manuela Eisenhardt,1,2 Sarah Leixner,1,2 Rafael Luja´n,3 Rainer Spanagel,1 and Ainhoa Bilbao1,2 1Institute of Psychopharmacology and 2Behavioral Genetics Research Group, Central Institute of Mental Health, Medical Faculty Mannheim/University of Heidelberg, J5, 68159 Mannheim, Germany, and 3Instituto de Investigacio´n en Discapacidades Neurolo´gicas, Departamento de Ciencias Me´dicas, Facultad de Medicina, Universidad Castilla-La Mancha, Campus Biosanitario, 02006 Albacete, Spain Glutamatergic input within the mesolimbic dopamine (DA) pathway plays a critical role in the development of addictive behavior. Although this is well established for some drugs of abuse, it is not known whether glutamate receptors within the mesolimbic system are involved in mediating the addictive properties of chronic alcohol use. Here we evaluated the contribution of mesolimbic NMDARs and AMPARs in mediating alcohol-seeking responses induced by environmental stimuli and relapse behavior using four inducible mutant mouse lines lacking the glutamate receptor genes Grin1 or Gria1 in either DA transporter (DAT) or D1R-expressing neurons. We first demonstrate the lack of GluN1 or GluA1 in either DAT- or D1R-expressing neurons in our mutant mouse lines by colocalization studies. We then show that GluN1 and GluA1 receptor subunits within these neuronal subpopulations mediate the alcohol deprivation effect, while having no impact on context- plus cue-induced reinstatement of alcohol-seeking behavior. We further validated these results pharmacologically by demonstrating similar reductions in the alcohol deprivation effect after infusion of the NMDAR antagonist me- mantine into the nucleus accumbens and ventral tegmental area of control mice, and a rescue of the mutant phenotype via pharmaco- logical potentiation of AMPAR activity using aniracetam.