Persistent Pain Maintains Morphine-Seeking Behavior After Morphine Withdrawal Through Reduced Mecp2 Repression of Glua1 in Rat Central Amygdala
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Case-Control Study of GRIA1 and GRIA3 Gene Variants in Migraine Jie Fang1, Xingkai An1, Shuai Chen2, Zhenzhen Yu1, Qilin Ma1,2* and Hongli Qu1*
Fang et al. The Journal of Headache and Pain (2016) 17:2 DOI 10.1186/s10194-016-0592-2 RESEARCH ARTICLE Open Access Case-control study of GRIA1 and GRIA3 gene variants in migraine Jie Fang1, Xingkai An1, Shuai Chen2, Zhenzhen Yu1, Qilin Ma1,2* and Hongli Qu1* Abstract Background: As the most abundant excitatory neurotransmitter in the central nervous system, glutamate has been accepted to play a major role in the pathophysiology of migraine. The previous studies have reported the glutamate receptor ionotropic GRIA1 and GRIA3 genes variants associated with migraine. The project aims to investigate the polymorphisms in both genes for their association with migraine in the Chinese Han population. Methods: A Han-Chinese case-control population, including 331 unrelated female migraine patients and 330 matched controls, was studied. Variants in genes (GRIA1 and GRIA3) were genotyped by Multiplex SNaPshot assay. Results: In the group of patients, the frequency of allele C was 84.1 % (557 C alleles) and allele T was 15.9 % (105 T alleles) for the GRIA1 (rs2195450) in migraineurs, this was significantly as compared with the controls (P = .001, OR = 1.786, 95 % CI: 1.28–2.49). And an association was also seen in the migraine with aura (MA) subtype (P = .012, OR = 2.092, 95 % CI: 1.17–3.76) and migraine without aura (MO) subtype (P =.002,OR=1.737, 95 % CI: 1.23–2.45). However, no evidence was found that GRIA1 (rs548294) or GRIA3 (rs3761555) is associated with migraine. Conclusion: Our data of this study confirmed the association of GRIA1 (rs2195450) to female migraine (MA, MO) susceptibility in the Chinese Han population. -

Sex Differences in Glutamate Receptor Gene Expression in Major Depression and Suicide
Molecular Psychiatry (2015) 20, 1057–1068 © 2015 Macmillan Publishers Limited All rights reserved 1359-4184/15 www.nature.com/mp IMMEDIATE COMMUNICATION Sex differences in glutamate receptor gene expression in major depression and suicide AL Gray1, TM Hyde2,3, A Deep-Soboslay2, JE Kleinman2 and MS Sodhi1,4 Accumulating data indicate that the glutamate system is disrupted in major depressive disorder (MDD), and recent clinical research suggests that ketamine, an antagonist of the N-methyl-D-aspartate (NMDA) glutamate receptor (GluR), has rapid antidepressant efficacy. Here we report findings from gene expression studies of a large cohort of postmortem subjects, including subjects with MDD and controls. Our data reveal higher expression levels of the majority of glutamatergic genes tested in the dorsolateral prefrontal cortex (DLPFC) in MDD (F21,59 = 2.32, P = 0.006). Posthoc data indicate that these gene expression differences occurred mostly in the female subjects. Higher expression levels of GRIN1, GRIN2A-D, GRIA2-4, GRIK1-2, GRM1, GRM4, GRM5 and GRM7 were detected in the female patients with MDD. In contrast, GRM5 expression was lower in male MDD patients relative to male controls. When MDD suicides were compared with MDD non-suicides, GRIN2B, GRIK3 and GRM2 were expressed at higher levels in the suicides. Higher expression levels were detected for several additional genes, but these were not statistically significant after correction for multiple comparisons. In summary, our analyses indicate a generalized disruption of the regulation of the GluRs in the DLPFC of females with MDD, with more specific GluR alterations in the suicides and in the male groups. -
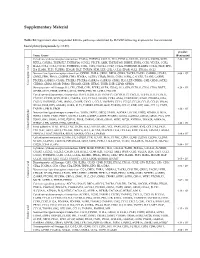
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

Ion Channels
UC Davis UC Davis Previously Published Works Title THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels. Permalink https://escholarship.org/uc/item/1442g5hg Journal British journal of pharmacology, 176 Suppl 1(S1) ISSN 0007-1188 Authors Alexander, Stephen PH Mathie, Alistair Peters, John A et al. Publication Date 2019-12-01 DOI 10.1111/bph.14749 License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: Ion channels. British Journal of Pharmacology (2019) 176, S142–S228 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels Stephen PH Alexander1 , Alistair Mathie2 ,JohnAPeters3 , Emma L Veale2 , Jörg Striessnig4 , Eamonn Kelly5, Jane F Armstrong6 , Elena Faccenda6 ,SimonDHarding6 ,AdamJPawson6 , Joanna L Sharman6 , Christopher Southan6 , Jamie A Davies6 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 3Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 4Pharmacology and Toxicology, Institute of Pharmacy, University of Innsbruck, A-6020 Innsbruck, Austria 5School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 6Centre for Discovery Brain Science, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. -

Gene List of the Targeted NGS MCD and CCA Gene Panel AKT3,ALX1
Gene List of the targeted NGS MCD and CCA gene panel AKT3,ALX1,ALX3,ALX4,AMPD2,ARFGEF2,ARID1B,ARX,ASPM,ATR,ATRX,B3GALTL,BRPF1,c12orf57,C6orf70,CASK,CCND2,CDK5RAP2,CDON,C ENPJ,CEP170,CHMP1A,COL4A1,CREBBP,CYP11A1,DCHS1,DCLK1,DCX,DHCR24,DHCR7,DIS3L2,DISC1,DISP1,DLL1,DMRTA2,DYNC1H1,DYRK1 A,EARS2,EFNB1,EMX1,EOMES,EP300,ERBB4,ERMARD,EXOSC3,FAM36A,FGF8,FGFR1,FGFR2,FLNA,FOXC1,FOXG1,FOXH1,FZD10,GLI2,GLI3,GP R56,GPSM2,HCCS,HESX1,HNRNPU,IGBP1,IGFBP1,ISPD,ITPA,KAL1,KAT6B,KATNB1,KIAA1279,KIF14,KIF1A,KIF1B,KIF21A,KIF2A,KIF5C,KIF7,L1 CAM,LAMB1,LAMC3,LRP2,MCPH1,MED12,MID1,NDE1,NFIB,NPC1,NR2F1,NSD1,NTRK1,NTRK3,OCEL1,OPA1,OTX2,PAFAH1B1,PAX6,PEX1,PHF1 0,PIK3R2,POLR3A,POLR3B,POMT1,POMT2,PTCH1,PTPRS,PYCR1,RAB3GAP1,RARS2,RELN,RFX3,ROBO1,ROBO3,RPS6KA3,RTTN,SATB2,SEPSEC S,SHH,SIX3,SLC12A6,SOX2,SPOCK1,SRPX2,TBCD,TBCE,TCF4,TDGF1,TEAD1,THBS2,TMEM5,TSC1,TSC2,TSEN15,TSEN2,TSEN34,TSEN54,TUBA1 A,TUBA8,TUBB,TUBB2A,TUBB2B,TUBB3,TUBB4A,TUBG1,VAX1,VRK1,WDR47,WDR62,ZBTB18,ZEB2,ZIC2. Gene List of the targeted NGS epilepsy gene panel AARS, ADGRV1, ADRA2B, ADSL, ALDH4A1, ALDH7A1, ALG13, ALPL, ARHGEF15, ARHGEF9, ARX, ASAH1, ATP1A2, ATP1A3, BRD2, CACNA1A, CACNA1H, CACNA2D2, CACNB4, CBL, CDKL5, CERS1, CHD2, CHRNA2, CHRNA4, CHRNB2, CLCN2, CLCN4, CLN8, CLTC, CNKSR2, CNTNAP2, CPA6, CPLX1, CSNK1G1, CSNK2B, CTNND2, DEPDC5, DHDDS, DNM1, DOCK7, DYNC1H1, EEF1A2, EFHC1, EIF2S3, EMC1, EPM2A, FASN, FLNA, FOXG1, GABBR2, GABRA1, GABRA2, GABRA3, GABRB2, GABRB3, GABRD, GABRG2, GAL, GNAO1, GOSR2, GRIA1, GRIN1, GRIN2A, GRIN2B, HCN1, HCN4, HDAC4, HNRNPU, IDH3A, IQSEC2, JRK, KCNA1, KCNA2, KCNB1, -

Glutamate Receptors Within the Mesolimbic Dopamine System Mediate Alcohol Relapse Behavior
The Journal of Neuroscience, November 25, 2015 • 35(47):15523–15538 • 15523 Neurobiology of Disease Glutamate Receptors within the Mesolimbic Dopamine System Mediate Alcohol Relapse Behavior Manuela Eisenhardt,1,2 Sarah Leixner,1,2 Rafael Luja´n,3 Rainer Spanagel,1 and Ainhoa Bilbao1,2 1Institute of Psychopharmacology and 2Behavioral Genetics Research Group, Central Institute of Mental Health, Medical Faculty Mannheim/University of Heidelberg, J5, 68159 Mannheim, Germany, and 3Instituto de Investigacio´n en Discapacidades Neurolo´gicas, Departamento de Ciencias Me´dicas, Facultad de Medicina, Universidad Castilla-La Mancha, Campus Biosanitario, 02006 Albacete, Spain Glutamatergic input within the mesolimbic dopamine (DA) pathway plays a critical role in the development of addictive behavior. Although this is well established for some drugs of abuse, it is not known whether glutamate receptors within the mesolimbic system are involved in mediating the addictive properties of chronic alcohol use. Here we evaluated the contribution of mesolimbic NMDARs and AMPARs in mediating alcohol-seeking responses induced by environmental stimuli and relapse behavior using four inducible mutant mouse lines lacking the glutamate receptor genes Grin1 or Gria1 in either DA transporter (DAT) or D1R-expressing neurons. We first demonstrate the lack of GluN1 or GluA1 in either DAT- or D1R-expressing neurons in our mutant mouse lines by colocalization studies. We then show that GluN1 and GluA1 receptor subunits within these neuronal subpopulations mediate the alcohol deprivation effect, while having no impact on context- plus cue-induced reinstatement of alcohol-seeking behavior. We further validated these results pharmacologically by demonstrating similar reductions in the alcohol deprivation effect after infusion of the NMDAR antagonist me- mantine into the nucleus accumbens and ventral tegmental area of control mice, and a rescue of the mutant phenotype via pharmaco- logical potentiation of AMPAR activity using aniracetam. -

The Glutamate Receptor Ion Channels
0031-6997/99/5101-0007$03.00/0 PHARMACOLOGICAL REVIEWS Vol. 51, No. 1 Copyright © 1999 by The American Society for Pharmacology and Experimental Therapeutics Printed in U.S.A. The Glutamate Receptor Ion Channels RAYMOND DINGLEDINE,1 KARIN BORGES, DEREK BOWIE, AND STEPHEN F. TRAYNELIS Department of Pharmacology, Emory University School of Medicine, Atlanta, Georgia This paper is available online at http://www.pharmrev.org I. Introduction ............................................................................. 8 II. Gene families ............................................................................ 9 III. Receptor structure ...................................................................... 10 A. Transmembrane topology ............................................................. 10 B. Subunit stoichiometry ................................................................ 10 C. Ligand-binding sites located in a hinged clamshell-like gorge............................. 13 IV. RNA modifications that promote molecular diversity ....................................... 15 A. Alternative splicing .................................................................. 15 B. Editing of AMPA and kainate receptors ................................................ 17 V. Post-translational modifications .......................................................... 18 A. Phosphorylation of AMPA and kainate receptors ........................................ 18 B. Serine/threonine phosphorylation of NMDA receptors .................................. -

Ligand-Gated Ion Channels
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2015/16: Ligand-gated ion channels. British Journal of Pharmacology (2015) 172, 5870–5903 THE CONCISE GUIDE TO PHARMACOLOGY 2015/16: Ligand-gated ion channels Stephen PH Alexander1, John A Peters2, Eamonn Kelly3, Neil Marrion3, Helen E Benson4, Elena Faccenda4, Adam J Pawson4, Joanna L Sharman4, Christopher Southan4, Jamie A Davies4 and CGTP Collaborators L 1 School of Biomedical Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK, N 2Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK, 3School of Physiology and Pharmacology, University of Bristol, Bristol, BS8 1TD, UK, 4Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2015/16 provides concise overviews of the key properties of over 1750 human drug targets with their pharmacology, plus links to an open access knowledgebase of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. The full contents can be found at http://onlinelibrary.wiley.com/ doi/10.1111/bph.13350/full. Ligand-gated ion channels are one of the eight major pharmacological targets into which the Guide is divided, with the others being: ligand-gated ion channels, voltage- gated ion channels, other ion channels, nuclear hormone receptors, catalytic receptors, enzymes and transporters. These are presented with nomenclature guidance and summary information on the best available pharmacological tools, alongside key references and suggestions for further reading. The Concise Guide is published in landscape format in order to facilitate comparison of related targets. -

Long-Term Potentiation Is Independent of the C-Tail of the Glua1 AMPA
RESEARCH ARTICLE Long-term potentiation is independent of the C-tail of the GluA1 AMPA receptor subunit Javier Dı´az-Alonso1†*, Wade Morishita2, Salvatore Incontro1, Jeffrey Simms3, Julia Holtzman3, Michael Gill3, Lennart Mucke3,4, Robert C Malenka2, Roger A Nicoll1* 1Department of Cellular and Molecular Pharmacology, University of California, San Francisco, San Francisco, United States; 2Nancy Pritzker Laboratory, Department of Psychiatry and Behavioral Sciences, Stanford University School of Medicine, Stanford, United States; 3Gladstone Institute of Neurological Disease, San Francisco, United States; 4Department of Neurology, University of California, San Francisco, San Francisco, United States Abstract We tested the proposal that the C-terminal domain (CTD) of the AMPAR subunit GluA1 is required for LTP. We found that a knock-in mouse lacking the CTD of GluA1 expresses normal LTP and spatial memory, assayed by the Morris water maze. Our results support a model in which LTP generates synaptic slots, which capture passively diffusing AMPARs. *For correspondence: [email protected] (JD-A); [email protected] (RAN) Introduction Long-term potentiation (LTP) requires the activity-dependent trafficking of AMPA receptors † Present address: Department (AMPARs) to the synapse (Collingridge et al., 2004; Malinow and Malenka, 2002; Nicoll, 2017). of Anatomy and Neurobiology, Most AMPARs in CA1 pyramidal cells are heterotetramers of either GluA1/GluA2 subunits or GluA2/ University of California, Irvine, GluA3 subunits, although other complexes can also occur (Zhao et al., 2019). The prevailing, recep- Irvine, United States tor centric, LTP model, posits that LTP-mediated covalent modification of the intracellular carboxy- Competing interests: The terminal domain (CTD, also referred to as C-tail) of GluA1 results in the capture of these modified authors declare that no GluA1 containing receptors by preexisting ‘slots’ in the postsynaptic density (PSD) (Hayashi et al., competing interests exist. -

Robles JTO Supplemental Digital Content 1
Supplementary Materials An Integrated Prognostic Classifier for Stage I Lung Adenocarcinoma based on mRNA, microRNA and DNA Methylation Biomarkers Ana I. Robles1, Eri Arai2, Ewy A. Mathé1, Hirokazu Okayama1, Aaron Schetter1, Derek Brown1, David Petersen3, Elise D. Bowman1, Rintaro Noro1, Judith A. Welsh1, Daniel C. Edelman3, Holly S. Stevenson3, Yonghong Wang3, Naoto Tsuchiya4, Takashi Kohno4, Vidar Skaug5, Steen Mollerup5, Aage Haugen5, Paul S. Meltzer3, Jun Yokota6, Yae Kanai2 and Curtis C. Harris1 Affiliations: 1Laboratory of Human Carcinogenesis, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 2Division of Molecular Pathology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 3Genetics Branch, NCI-CCR, National Institutes of Health, Bethesda, MD 20892, USA. 4Division of Genome Biology, National Cancer Center Research Institute, Tokyo 104-0045, Japan. 5Department of Chemical and Biological Working Environment, National Institute of Occupational Health, NO-0033 Oslo, Norway. 6Genomics and Epigenomics of Cancer Prediction Program, Institute of Predictive and Personalized Medicine of Cancer (IMPPC), 08916 Badalona (Barcelona), Spain. List of Supplementary Materials Supplementary Materials and Methods Fig. S1. Hierarchical clustering of based on CpG sites differentially-methylated in Stage I ADC compared to non-tumor adjacent tissues. Fig. S2. Confirmatory pyrosequencing analysis of DNA methylation at the HOXA9 locus in Stage I ADC from a subset of the NCI microarray cohort. 1 Fig. S3. Methylation Beta-values for HOXA9 probe cg26521404 in Stage I ADC samples from Japan. Fig. S4. Kaplan-Meier analysis of HOXA9 promoter methylation in a published cohort of Stage I lung ADC (J Clin Oncol 2013;31(32):4140-7). Fig. S5. Kaplan-Meier analysis of a combined prognostic biomarker in Stage I lung ADC. -

Altered Expression of Genes Encoding Neurotransmitter Receptors in Gnrh Neurons of Proestrous Mice
ORIGINAL RESEARCH published: 07 October 2016 doi: 10.3389/fncel.2016.00230 Altered Expression of Genes Encoding Neurotransmitter Receptors in GnRH Neurons of Proestrous Mice Csaba Vastagh 1*, Annie Rodolosse 2, Norbert Solymosi 3 and Zsolt Liposits 1, 4 1 Laboratory of Endocrine Neurobiology, Institute of Experimental Medicine, Hungarian Academy of Sciences, Budapest, Hungary, 2 Functional Genomics Core, Institute for Research in Biomedicine (IRB Barcelona), Barcelona, Spain, 3 Department of Animal Hygiene, Herd-Health and Veterinary Ethology, University of Veterinary Medicine, Budapest, Hungary, 4 Department of Neuroscience, Faculty of Information Technology and Bionics, Pázmány Péter Catholic University, Budapest, Hungary Gonadotropin-releasing hormone (GnRH) neurons play a key role in the central regulation of reproduction. In proestrous female mice, estradiol triggers the pre-ovulatory GnRH surge, however, its impact on the expression of neurotransmitter receptor genes in GnRH neurons has not been explored yet. We hypothesized that proestrus is accompanied by substantial changes in the expression profile of genes coding for neurotransmitter Edited by: receptors in GnRH neurons. We compared the transcriptome of GnRH neurons obtained Hansen Wang, from intact, proestrous, and metestrous female GnRH-GFP transgenic mice, respectively. University of Toronto, Canada About 1500 individual GnRH neurons were sampled from both groups and their Reviewed by: Pamela L. Mellon, transcriptome was analyzed using microarray hybridization and real-time -

Perkinelmer Genomics to Request the Saliva Swab Collection Kit for Patients That Cannot Provide a Blood Sample As Whole Blood Is the Preferred Sample
STAT Focused Autism and Intellectual Disability Panel Test Code D5130F Test Summary This test analyzes 264 genes that have been associated with Autism and Intellectual Disability. STAT means results are available in 7 - 10 days. Turn-Around-Time (TAT)* 7 - 10 days Acceptable Sample Types Whole Blood (EDTA) (Preferred sample type) DNA, Isolated Dried Blood Spots Saliva Acceptable Billing Types Self (patient) Payment Institutional Billing Indications for Testing This panel may be appropriate for individuals with a clinical suspicion of Lissencephaly and/or for indiviudals with a family history of Lissencephaly. Test Description This panel analyzes 264 genes that have been associated with Focused Autism and Intellectual Disability and/or disorders associated with Focused Autism and Intellectual Disability. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect the majority of deletions and duplications of three exons or greater in size. Smaller CNV events may also be detected and reported, but additional follow-up testing is recommended if a smaller CNV is suspected. All variants are classified according to ACMG guidelines. Condition Description Autism Spectrum Disorder (ASD) refers to a group of developmental disabilities that are typically associated with challenges of varying severity in the areas of social interaction, communication,